Induced Conformation vs. Induced Proximity: Unraveling the Molecular Mechanisms of Caspase-9 Activation
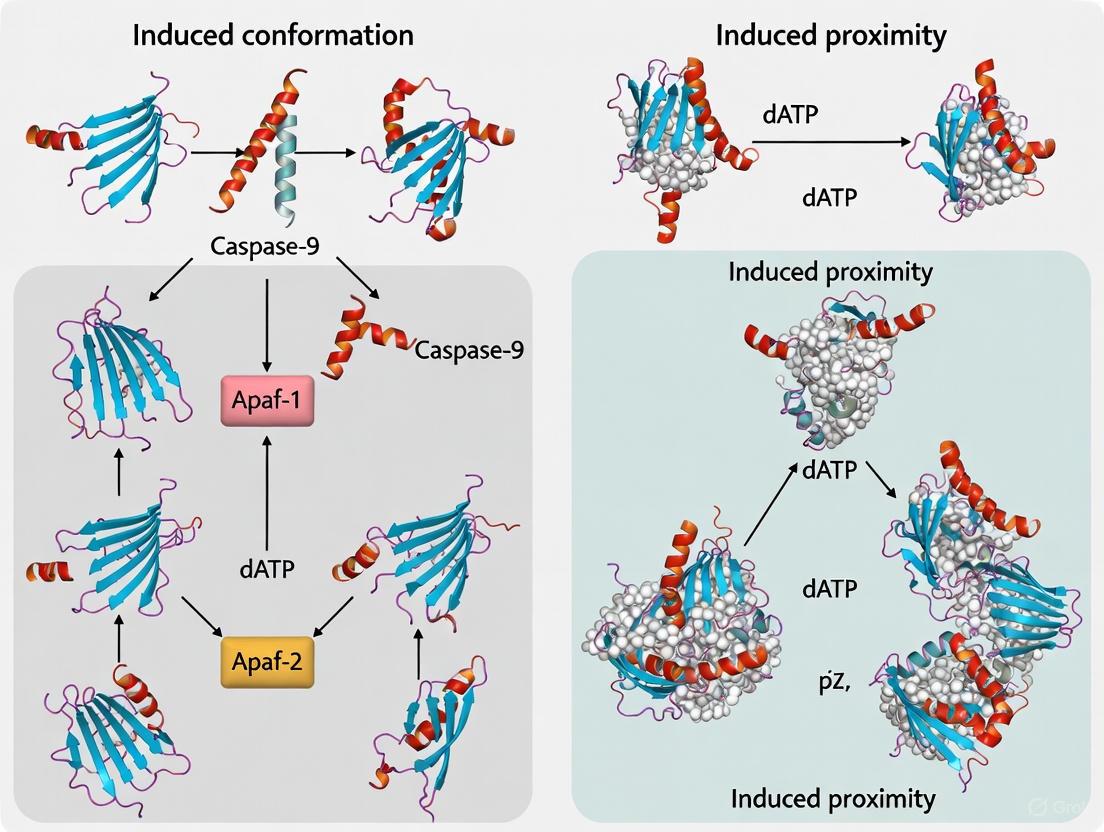
The activation of caspase-9, the initiator caspase of the intrinsic apoptosis pathway, is a pivotal event in programmed cell death with profound implications for cancer therapy and degenerative diseases.
Induced Conformation vs. Induced Proximity: Unraveling the Molecular Mechanisms of Caspase-9 Activation
Abstract
The activation of caspase-9, the initiator caspase of the intrinsic apoptosis pathway, is a pivotal event in programmed cell death with profound implications for cancer therapy and degenerative diseases. For decades, the 'induced proximity' model, which posits that the apoptosome merely serves as a platform to concentrate caspase-9 monomers for dimerization, was the prevailing paradigm. However, recent structural and biochemical studies have challenged this view, providing compelling evidence for an 'induced conformation' model where the apoptosome actively induces activating structural changes in caspase-9. This article synthesizes current research to compare these two models, exploring the foundational science, key experimental methodologies, unresolved controversies, and the direct impact of this fundamental biological mechanism on therapeutic discovery and clinical application for researchers and drug development professionals.
The Apoptotic Trigger: Foundational Principles of Caspase-9 in the Intrinsic Pathway
The Central Role of Caspase-9 in Intrinsic Apoptosis and Cellular Homeostasis
Caspase-9 functions as a critical initiator caspase in the intrinsic apoptotic pathway, serving as the molecular link between cellular stress signals and the execution phase of programmed cell death. Unlike effector caspases that directly dismantle cellular structures, caspase-9 acts as the apical protease that initiates the caspase cascade following mitochondrial outer membrane permeabilization [1] [2]. This pathway is essential for eliminating damaged or potentially dangerous cells, thus maintaining tissue homeostasis and preventing carcinogenesis [1] [3]. The activation mechanism of caspase-9 represents a fundamental biological process that has been the subject of intensive research, yielding two predominant but potentially complementary models: induced proximity and induced conformation [1] [3] [4]. Understanding these mechanisms provides crucial insights into both normal physiological processes and pathological conditions ranging from cancer to neurodegenerative disorders.
Molecular Mechanisms of Caspase-9 Activation
Structural Basis for Caspase-9 Function
Caspase-9 is a 416-residue enzyme comprising two primary functional domains: an N-terminal Caspase Activation and Recruitment Domain (CARD, residues 1-92) and a C-terminal Protease Domain (PD, residues 139-416) [5]. These domains are connected by a long, disordered linker (residues 93-138), which provides structural flexibility essential for activation [5]. The protease domain can be further subdivided into p20 (residues 139-289) and p10 (residues 341-416) subunits, connected by another disordered linker (residues 290-340) that contains critical autocleavage sites (E306, D315) [5].
Table 1: Structural Domains of Caspase-9 and Their Functions
| Domain | Residues | Key Features | Functional Role |
|---|---|---|---|
| CARD | 1-92 | Caspase Recruitment Domain | Homotypic interaction with Apaf-1 CARD for apoptosome recruitment |
| Linker | 93-138 | Disordered, flexible | Connects CARD and protease domain, allows conformational freedom |
| p20 Subunit | 139-289 | Large catalytic subunit | Contains elements of the catalytic site |
| p10 Linker | 290-340 | Disordered with cleavage sites (E306, D315) | Site for autocleavage and regulatory processing |
| p10 Subunit | 341-416 | Small catalytic subunit | Completes formation of the catalytic domain |
The CARD domain mediates specific protein-protein interactions, particularly with the CARD domain of Apaf-1, facilitating recruitment to the apoptosome complex [1] [5]. This interaction occurs through complementary interfaces that are indispensable for caspase-9 activation [1]. Recent structural investigations have revealed that this interaction involves multiple interfaces rather than a simple 1:1 binding [1].
The Apoptosome: Activation Platform for Caspase-9
The apoptosome is a massive 1.1-1.3 MDa heptameric complex that serves as the activation platform for caspase-9 [5]. This complex forms when cytochrome c is released from mitochondria and binds to Apaf-1, promoting ATP-dependent heptamerization [5]. Each Apaf-1 monomer consists of a CARD domain, an ATP-binding domain, and a regulatory domain that binds cytochrome c [5]. Cryo-EM structures have revealed that the apoptosome recruits caspase-9 through asymmetric CARD-CARD interactions with a 4:7 stoichiometry (caspase-9:Apaf-1) [5]. Interestingly, while the CARD interactions are well-resolved in structural studies, the caspase-9 protease domains remain flexibly tethered to the platform and are not visible in cryo-EM reconstructions, suggesting significant structural dynamics during activation [5].
Models of Caspase-9 Activation
Two primary models have been proposed to explain caspase-9 activation, each supported by distinct experimental evidence:
2.3.1 Induced Proximity Model This model posits that the apoptosome serves primarily as a platform to increase the local concentration of caspase-9 monomers, promoting homodimerization and subsequent autoactivation [3] [4]. According to this hypothesis, caspase-9 exists predominantly as a monomer in solution and gains catalytic activity through dimerization facilitated by proximity on the apoptosome scaffold [4]. Support for this model comes from observations that caspase-9 exhibits minimal catalytic activity as a monomer but shows enhanced activity when artificially dimerized or concentrated [4].
2.3.2 Induced Conformation Model This alternative model proposes that binding to the apoptosome induces specific conformational changes in caspase-9 that activate its catalytic function, rather than simply promoting dimerization [3] [4]. Evidence supporting this model includes the engineering of a constitutively dimeric caspase-9 that, while more active than wild-type monomers, displayed only a fraction of the activity of apoptosome-activated caspase-9 and could not be further stimulated by Apaf-1 [4]. This suggests that dimerization alone is insufficient to fully recapitulate the activation mediated by the apoptosome.
2.3.3 Hybrid Model and Recent Insights Recent research utilizing methyl-TROSY NMR spectroscopy has revealed that caspase-9 protease domains remain predominantly monomeric even when bound to the apoptosome and only undergo extensive dimerization upon substrate binding [5]. This suggests a revised model where apoptosome binding primes caspase-9 for activation by orienting the protease domains in a configuration that facilitates rapid dimerization specifically when substrate becomes available [5]. This substrate-induced dimerization represents an additional regulatory layer controlling caspase-9 activity.
Figure 1: Caspase-9 Activation Models. Three proposed mechanisms for caspase-9 activation on the apoptosome: Induced Proximity (dimerization-driven), Induced Conformation, and recent evidence supporting Substrate-Induced Dimerization.
Experimental Approaches and Key Findings
Engineered Dimeric Caspase-9 Studies
A pivotal experiment in evaluating the activation models involved engineering a constitutively dimeric caspase-9 by modifying residues at the dimer interface to relieve steric hindrance [4]. The crystal structure of this engineered dimer closely resembled wild-type caspase-9, confirming that the modifications did not induce major structural alterations [4].
Table 2: Comparison of Caspase-9 Activation States
| Parameter | Caspase-9 Monomer | Engineered Dimer | Apoptosome-Activated |
|---|---|---|---|
| Quaternary Structure | Monomer | Constitutive dimer | Monomeric PDs, primed for dimerization |
| Catalytic Activity | Low basal level | Moderately enhanced | Highly activated |
| Response to Apaf-1 | Activated | No significant enhancement | Dependent on apoptosome |
| Cell Death Induction | Baseline | More efficient than monomer | Highly efficient |
| Dimerization Constant | Very weak (mM range) | Stable dimer | Substrate-induced dimerization |
This engineered dimer exhibited higher catalytic activity in vitro and induced more efficient cell death compared to wild-type caspase-9 when expressed in cells [4]. However, its activity represented only a small fraction of that achieved through Apaf-1-mediated activation, and unlike wild-type caspase-9, could not be significantly enhanced by Apaf-1 [4]. These findings challenged the simple interpretation of the induced proximity model and suggested that dimerization alone is qualitatively different from apoptosome-mediated activation.
NMR Spectroscopy of Apoptosome-Bound Caspase-9
Recent advances in NMR spectroscopy have enabled detailed investigation of the caspase-9 activation mechanism within the massive 1.3 MDa apoptosome complex [5]. Using methyl-TROSY NMR with deuterated, methyl-labeled molecules, researchers examined the structural dynamics of caspase-9 protease domains tethered to the apoptosome [5].
Experimental Protocol:
- Isotope Labeling: Caspase-9 was produced with specific 13CH3-methyl labeling at isoleucine, leucine, and valine residues
- Apoptosome Reconstitution: The native apoptosome complex was reconstituted using insect cell expression systems
- NMR Measurements: Methyl-TROSY spectra were acquired for caspase-9 in various states: free in solution, bound to apoptosome, and with added substrate
- Dimerization Assessment: NMR signals were analyzed to determine the oligomeric state of caspase-9 protease domains in different conditions
- Activity Correlations: NMR findings were correlated with enzymatic activity assays using fluorogenic substrates
This approach revealed that caspase-9 protease domains remain predominantly monomeric when bound to the apoptosome in the absence of substrate, challenging both simple dimerization models [5]. Only upon substrate addition did significant dimerization occur, suggesting a model where apoptosome binding primes caspase-9 for substrate-induced dimerization [5].
Biochemical and Cellular Assays
Multiple complementary approaches have been employed to study caspase-9 function:
Activity Assays: Fluorogenic substrates containing the LEHD sequence are used to measure caspase-9 activity. The tetrapeptide Z-LEHD-fmk acts as an irreversible inhibitor by covalently modifying the active site cysteine [5].
Genetic Models: Caspase-9 knockout mice die perinatally with severe brain abnormalities due to impaired apoptosis during development [1]. Embryonic stem cells and fibroblasts lacking caspase-9 show resistance to apoptotic stimuli including UV irradiation, γ-irradiation, and dexamethasone [1].
Pharmacological Inhibition: Caspase-9 inhibitors include the broad-spectrum inhibitor Z-VAD-FMK and the more specific Z-LEHD-FMK [6] [7]. Q-VD-OPh offers improved cell permeability and reduced toxicity at high concentrations [6].
Research Reagent Solutions for Caspase-9 Studies
Table 3: Essential Research Tools for Caspase-9 Investigation
| Reagent/Category | Specific Examples | Key Applications | Technical Notes |
|---|---|---|---|
| Chemical Inhibitors | Z-LEHD-FMK, Q-VD-OPh, Emricasan (IDN-6556) | Inhibiting caspase-9 activity in cellular and in vivo models | Q-VD-OPh shows better permeability and lower toxicity than earlier inhibitors |
| Activity Assays | Fluorogenic substrates with LEHD sequence, Antibodies against cleaved caspase-9 (D315, D330 neoepitopes) | Measuring caspase-9 activation and activity | D315 and D330 neoepitopes indicate autocleavage vs. caspase-3 cleavage respectively |
| Genetic Tools | Caspase-9 knockout mice, siRNA/shRNA, CRISPR/Cas9 knockout constructs | Studying consequences of caspase-9 loss of function | Caspase-9 KO mice show perinatal lethality with brain malformations |
| Apoptosome Components | Recombinant Apaf-1, Cytochrome c, ATP | Reconstituting apoptosome system in vitro | Required for studying caspase-9 activation mechanisms |
| Structural Biology | Methyl-TROSY NMR with 13CH3-labeling, Cryo-EM analysis | Investigating caspase-9 structure and dynamics within apoptosome | NMR allows study of flexible regions invisible to cryo-EM |
| Cellular Models | Caspase-9 null embryonic stem cells, CTIAC11, MC38, MLE-12 cell lines | Assessing cell-type specific functions | Used in high-throughput drug screening and pathway analysis |
Caspase-9 in Disease and Therapeutic Targeting
Role in Disease Pathogenesis
Caspase-9 dysregulation contributes to various human diseases:
Cancer: Reduced caspase-9 activity represents a tumor escape mechanism that confers resistance to chemotherapy [1]. CASPASE-9 polymorphisms are associated with increased susceptibility to lung, bladder, pancreatic, colorectal, and gastric cancers [1] [8]. In head and neck squamous cell carcinoma, reduced caspase-9 activity and Apaf-1 expression contribute to cisplatin resistance [1].
Neurodegenerative Disorders: Caspase-9 activation occurs in end-stage Huntington's disease, suggesting apoptotic contribution to neuronal death [1]. Caspase-9 polymorphisms are linked to increased risk of discogenic low back pain through effects on disc degeneration [1].
Fibrotic Diseases: Recent evidence implicates caspase-9 in pulmonary fibrosis, where it promotes epithelial apoptosis and activates β-catenin signaling to drive fibrotic progression [7]. Caspase-9 inhibition in bleomycin-induced lung fibrosis models reduces collagen deposition and improves lung architecture [7].
Autoimmune and Inflammatory Conditions: CASPASE-9 gene polymorphisms are associated with multiple sclerosis susceptibility [1]. The CASPASE-9 (Ex5 + 32G/A) GG genotype correlates with higher disease risk [1].
Therapeutic Applications and Clinical Trials
Several therapeutic approaches targeting caspase-9 have been investigated:
Direct Caspase Inhibitors: Peptide-based inhibitors (Z-VAD-FMK), peptidomimetic compounds (emricasan/IDN-6556), and non-peptidic small molecules have been developed [6]. While showing promise in preclinical studies, many have faced challenges in clinical trials due to inadequate efficacy, poor target specificity, or adverse effects [6].
Immunomodulatory Approaches: Recent research reveals that caspase-9 inhibition can enhance antitumor immunity by promoting type I interferon production through the mtDNA/cGAS/STING pathway [9]. Combining caspase-9 inhibition with Hsp90 inhibitors triggers immunogenic cell death and synergizes with PD-L1 blockade to achieve complete tumor regression in mouse models [9].
Gene Therapy Applications: An inducible caspase-9 (iCasp9) system has been developed as a safety switch in cell therapies, allowing elimination of transplanted cells in case of adverse events [1].
The prevailing view of caspase-9 activation has evolved beyond a simple dichotomy between induced proximity and induced conformation models. Current evidence supports a hybrid model where apoptosome binding organizes caspase-9 protease domains in a primed state that remains predominantly monomeric until substrate binding induces productive dimerization and full catalytic activation [5]. This sophisticated regulatory mechanism ensures precise control over the initiation of apoptosis, preventing accidental cell death while allowing rapid activation when appropriate signals are received.
The multifaceted roles of caspase-9 extend beyond traditional apoptosis to include regulation of cellular differentiation, innate immunity, mitochondrial homeostasis, and autophagy [8]. These diverse functions, coupled with its central position in the intrinsic apoptotic pathway, establish caspase-9 as a critical mediator of cellular homeostasis and a promising therapeutic target for conditions ranging from cancer to fibrotic diseases. Future research will likely focus on developing more specific caspase-9 modulators and elucidating the full spectrum of its non-apoptotic functions in health and disease.
Figure 2: Integrated Caspase-9 Activation Pathway and Functional Consequences. The caspase-9 activation cascade from initial cellular stress to final physiological outcomes, highlighting how dysregulation leads to pathological conditions.
The apoptosome is a central signaling platform in the intrinsic pathway of apoptosis, responsible for the proteolytic activation of initiator caspase-9. This large protein complex assembles in response to cellular stress signals that trigger mitochondrial outer membrane permeabilization and cytochrome c release into the cytosol. The core components include apoptotic protease-activating factor 1 (Apaf-1) and cytochrome c, which together form a heptameric complex in the presence of ATP or dATP. This complex serves as an activation platform for procaspase-9, which then initiates a cascade of caspase activation leading to programmed cell death. The precise mechanism by which the apoptosome activates caspase-9 has been the subject of extensive research, primarily focusing on two competing models: the induced proximity model and the induced conformation model. Understanding the molecular architecture and function of the apoptosome provides critical insights for therapeutic interventions in diseases characterized by dysregulated apoptosis, including cancer, neurodegenerative disorders, and autoimmune conditions.
Structural Architecture of the Apoptosome
The apoptosome is a wheel-shaped complex with a central hub and seven spokes, formed through the oligomerization of seven Apaf-1 molecules, each bound to one cytochrome c molecule [10]. Cryo-electron microscopy studies have revealed the near-atomic structure of the human apoptosome at 3.8-4.1 Å resolution, providing unprecedented insights into its molecular organization [10] [11]. The complex measures approximately 270 Å in diameter and 75 Å in height, with a central hub that exhibits seven-fold rotational symmetry [11].
Each Apaf-1 protomer within the apoptosome contains multiple domains: an N-terminal caspase recruitment domain (CARD), a nucleotide-binding domain (NBD), a helical domain (HD1), a winged helix domain (WHD), a second helical domain (HD2), and 15 WD40 repeats that form two distinct β-propellers (WD1 and WD2) [10]. In the autoinhibited, monomeric state of Apaf-1, these domains are arranged to maintain the protein in an inactive conformation through extensive intramolecular interactions.
Table 1: Domain Organization of Apaf-1 in the Apoptosome
| Domain | Abbreviation | Primary Function | Structural Features |
|---|---|---|---|
| Caspase Recruitment Domain | CARD | Recruits procaspase-9 via CARD-CARD interactions | N-terminal domain, forms disk-like structure above central hub |
| Nucleotide-Binding Domain | NBD | Binds dATP/ATP and mediates oligomerization | Central hub component, AAA+ ATPase family |
| Helical Domain 1 | HD1 | Structural role in central hub formation | Part of the central hub with extensive α-helices |
| Winged Helix Domain | WHD | Connects central hub to HD2 | Key interface domain with charged residues |
| Helical Domain 2 | HD2 | Connects WHD to WD40 repeats | Extended arm structure |
| WD40 Repeats | WD1/WD2 | Binds cytochrome c; sensor domain | Forms two β-propellers, V-shaped sensor |
Conformational Changes During Activation
The transition from autoinhibited Apaf-1 monomer to active apoptosome involves dramatic conformational changes triggered by cytochrome c binding and nucleotide exchange. In the inactive state, Apaf-1 exists as an ADP-bound monomer with constrained domain arrangements that prevent oligomerization [10]. Cytochrome c binding to the WD40 repeats releases this autoinhibition by disrupting interdomain interactions, particularly those involving the HD2 and WHD domains [10].
The exchange of ADP for dATP or ATP triggers further conformational changes that enable oligomerization [10] [11]. Structural comparisons between autoinhibited Apaf-1 and the active apoptosome reveal significant domain rearrangements, with the NBD, HD1, and WHD forming the central hub, while the WD40 domains extend outward as spokes [10]. These conformational changes create binding surfaces that facilitate the heptameric assembly and formation of the caspase-9 activation platform.
Diagram 1: The apoptosome assembly pathway, showing key molecular events from cytochrome c release to caspase-9 activation.
Molecular Mechanism of Caspase-9 Activation
The CARD Disk and Caspase-9 Recruitment
The activation of caspase-9 on the apoptosome involves the formation of a unique substructure known as the CARD disk. This disk-like assembly sits atop the central hub of the apoptosome and consists of CARD domains from both Apaf-1 and procaspase-9 [11]. Structural studies reveal that this disk contains four Apaf-1/pc-9 CARD pairs arranged in a shallow spiral, with the fourth pc-9 CARD exhibiting lower occupancy [11]. This arrangement creates a structural mismatch with the seven-fold symmetry of the platform beneath.
The CARD-CARD interactions between Apaf-1 and procaspase-9 are critical for recruiting the zymogen to the activation platform. On average, the Apaf-1 CARDs recruit 3-5 procaspase-9 molecules to the apoptosome, with one catalytic domain potentially "parked" on the central hub when an odd number of zymogens are bound [11]. This suggests a stoichiometry of one or, at most, two pc-9 dimers per active apoptosome, challenging earlier assumptions about the activation mechanism.
Induced Proximity vs. Induced Conformation Models
The mechanism of caspase-9 activation on the apoptosome has been the subject of extensive debate, primarily between two competing models:
Induced Proximity Model: This model posits that the apoptosome serves as a platform to bring multiple caspase-9 zymogens into close proximity, facilitating their autocatalytic activation through trans-proteolysis [12]. According to this view, the primary role of the apoptosome is to increase the local concentration of caspase-9 molecules, enabling dimerization and subsequent autoactivation.
Induced Conformation Model: This alternative model suggests that binding to the apoptosome induces conformational changes in caspase-9 that directly activate the protease, independent of dimerization [3] [13]. Support for this model comes from experiments showing that engineered, dimeric caspase-9 exhibits only a fraction of the activity of Apaf-1-activated wild-type caspase-9 and is not further stimulated by Apaf-1 [3].
Current evidence suggests that both proximity and conformational changes contribute to caspase-9 activation, with the apoptosome functioning as an allosteric regulator that dramatically enhances the proteolytic activity of caspase-9 [14]. The resulting complex functions as a holoenzyme in which caspase-9 is the catalytic subunit and Apaf-1 serves as its allosteric regulator [14].
Table 2: Comparison of Caspase-9 Activation Models
| Feature | Induced Proximity Model | Induced Conformation Model | Current Understanding |
|---|---|---|---|
| Primary Mechanism | Dimerization through increased local concentration | Allosteric activation through conformational change | Combination of both mechanisms |
| Role of Apoptosome | Passive platform for concentration | Active allosteric regulator | Allosteric regulator that facilitates dimerization |
| Key Experimental Evidence | Caspase-9 dimerization enhances activity | Engineered dimers show limited activity without Apaf-1 | Apaf-1 increases catalytic efficiency by several orders of magnitude |
| Structural Basis | CARD disk facilitates caspase-9 proximity | Specific Apaf-1-caspase-9 interactions alter active site | CARD spiral creates unique activation environment |
| Catalytic Efficiency | Moderate enhancement | Potent activation | Holoenzyme with dramatically enhanced activity |
Diagram 2: Comparison of caspase-9 activation models, showing the distinct pathways proposed by induced proximity and induced conformation mechanisms.
Experimental Analysis of Apoptosome Function
Key Methodologies and Experimental Approaches
Structural and biochemical characterization of the apoptosome has relied on several sophisticated experimental techniques:
Cryo-Electron Microscopy (Cryo-EM): Recent advances in cryo-EM have enabled determination of the apoptosome structure at near-atomic resolution (3.8-4.1 Å) [10] [11]. Typical protocols involve expression and purification of full-length human Apaf-1 from baculovirus-infected insect cells, followed by in vitro assembly of the apoptosome with cytochrome c and dATP. The assembled complex is then imaged under cryo-conditions, often using a Titan Krios microscope operating at 300 kV. Single-particle analysis with 2D and 3D classification yields high-resolution structures that reveal molecular details of domain arrangements and interaction interfaces.
Biochemical Assays for Apoptosome Activity: Functional characterization of the apoptosome typically involves:
- Caspase activation assays: Measuring LEHDase (caspase-9) or DEVDase (caspase-3) activity using fluorogenic substrates.
- Stoichiometry analysis: Determining the number of caspase-9 molecules bound per apoptosome using quantitative Western blotting or fluorescence-based methods.
- Nucleotide exchange studies: Investigating the role of dATP/ATP binding and hydrolysis in apoptosome assembly using radioactive or fluorescent nucleotide analogs.
Site-directed Mutagenesis: Structure-guided mutagenesis of key residues in Apaf-1 has been instrumental in validating interaction interfaces and understanding the functional significance of specific domains. For example, mutations in the cytochrome c binding site on WD40 repeats or in the nucleotide-binding pocket have demonstrated the importance of these regions for apoptosome assembly and function [10].
Research Reagent Solutions
Table 3: Essential Research Reagents for Apoptosome Studies
| Reagent/Category | Specific Examples | Function/Application | Experimental Notes |
|---|---|---|---|
| Expression Systems | Baculovirus-infected insect cells | Production of full-length human Apaf-1 | Maintains proper folding and post-translational modifications |
| Assembly Components | Horse cytochrome c, dATP/ATP | In vitro apoptosome reconstitution | Excess cytochrome c and 1 mM dATP typically used |
| Protease Substrates | LEHD-AFC (7-amino-4-trifluoromethyl coumarin) | Caspase-9 activity measurement | Fluorogenic substrate for kinetic assays |
| Buffers & Solutions | Glycerol gradients, Gel filtration buffers | Complex purification and analysis | Linear glycerol gradients for complex separation |
| Structural Biology Tools | Cryo-EM grids (holey carbon), Vitrification devices | Sample preparation for cryo-EM | Titan Krios microscope with energy filter for high-resolution data |
Discussion and Research Implications
The structural and mechanistic insights into apoptosome function have significant implications for understanding and manipulating cell death pathways in human health and disease. The near-atomic resolution structures of the apoptosome represent a major advancement in the cell death field, providing a molecular framework for understanding how cytochrome c binding and nucleotide exchange trigger Apaf-1 activation and oligomerization [10] [11].
The debate between induced proximity and induced conformation models for caspase-9 activation has evolved toward a more integrated understanding, where the apoptosome functions as an allosteric regulator that dramatically enhances caspase-9 activity through a combination of proximity and conformational effects [3] [14]. The unique CARD disk architecture, with its spiral arrangement of CARD domains, creates a specialized environment for caspase-9 activation that differs from simple dimerization in solution [11].
From a therapeutic perspective, the apoptosome represents a potential target for modulating cell death in various pathological conditions. In cancer, where apoptosis is often suppressed, strategies to enhance apoptosome function could restore cell death in tumor cells. Conversely, in neurodegenerative diseases characterized by excessive apoptosis, inhibiting apoptosome assembly or function might protect vulnerable neurons. The structural insights gained from recent high-resolution studies provide a foundation for structure-based drug design targeting specific interfaces in the apoptosome.
Future research directions include elucidating the precise mechanism of caspase-9 activation on the CARD disk, understanding the regulation of apoptosome activity by endogenous inhibitors such as XIAP, and exploring the role of post-translational modifications in modulating apoptosome function. Additionally, the development of small molecule modulators of apoptosome activity will both provide useful research tools and potential therapeutic leads for diseases characterized by dysregulated apoptosis.
Procaspase-9, the initiator caspase of the intrinsic apoptotic pathway, exists predominantly as a monomeric zymogen in solution, with its Caspase Activation and Recruitment Domain (CARD) playing a critical role beyond simple apoptosome recruitment. Recent structural and biochemical evidence reveals intricate CARD:core domain interactions that influence catalytic function and regulation [15]. The long linker loop connecting the CARD to the catalytic core provides conformational flexibility, allowing the zymogen to adopt distinct states [1]. While the traditional induced proximity model posits that apoptosome-mediated dimerization drives activation, emerging research supports an induced conformation model where the apoptosome actively alters the structural architecture of procaspase-9 to generate catalytic competence [4]. This guide systematically compares these activation mechanisms through experimental data, structural insights, and methodological protocols to provide researchers with a comprehensive framework for understanding procaspase-9 regulation in health and disease.
Structural Architecture of Monomeric Procaspase-9
The procaspase-9 zymogen exhibits a unique domain organization that dictates its regulatory mechanisms and activation pathways.
Domain Organization and Key Features
- CARD Domain: An N-terminal six-helix bundle with protein-binding motifs that facilitate homotypic interactions with the Apaf-1 CARD in the apoptosome [1] [2]. This domain remains covalently attached to the catalytic core in the intracellular environment and participates in intramolecular regulation [15].
- Linker Region: A flexible connector between the CARD and catalytic domain that is susceptible to proteolytic processing but is not strictly required for activation [1]. Its length and flexibility allow for conformational rearrangements during activation.
- Catalytic Core: Comprised of large (p35) and small (p12) subunits connected by an extended intersubunit linker [1]. Unlike executioner caspases, procaspase-9 exhibits significant catalytic activity in its uncleaved form due to this elongated linker [1].
Table 1: Structural Domains of Monomeric Procaspase-9
| Domain | Structural Features | Functional Role | Regulatory Significance |
|---|---|---|---|
| CARD | Six-helix bundle, protein-binding motifs | Apoptosome recruitment via Apaf-1 CARD interaction | Intramolecular interaction with core domain modulates activity [15] |
| Linker Region | Flexible peptide connector | Connects CARD to catalytic core | Proteolytic processing regulates apoptosome dissociation [1] |
| Large Subunit (p35) | Contains critical active site residues | Catalytic activity | Longer intersubunit linker enables activity without cleavage [1] |
| Small Subunit (p12) | Stabilizes dimer interface | Structural integrity in active dimer | Contributes to catalytic efficiency in activated state |
| Intersubunit Linker | Extended, flexible loop | Connects large and small subunits | Cleavage acts as molecular timer for apoptosome activity [1] |
Distinctive Structural Characteristics
Procaspase-9 possesses several structural features that distinguish it from executioner caspases. It exists predominantly as a monomer in solution, whereas effector caspases are constitutive dimers [4]. The active site conformation in the monomeric zymogen exists in an unproductive state, requiring reorganization for full catalytic activity [12]. Notably, procaspase-9 maintains significant catalytic competence even before proteolytic processing at the intersubunit linker, a feature enabled by its extended linker length [1].
The CARD Domain: Beyond Apoptosome Recruitment
While traditionally viewed primarily as an apoptosome recruitment module, the CARD domain exhibits sophisticated regulatory functions that extend beyond simple tethering.
CARD:Core Domain Interactions
Biochemical evidence demonstrates that the CARD domain physically interacts with the catalytic core in the absence of the apoptosome [15]. This interaction requires a properly formed active site with ordered active-site loops. When these loops are disordered, the CARD and core domains behave as independent, loosely tethered units. However, with a properly ordered active-site loop bundle, these domains form a single folding unit that influences catalytic function [15]. This intramolecular interaction represents a previously unappreciated layer of caspase-9 regulation, suggesting the CARD may participate in substrate recruitment or recognition beyond its established role in apoptosome binding.
Functional Consequences of CARD Interactions
The presence of the CARD domain covalently linked to the catalytic core modulates enzymatic activity. Full-length caspase-9 with an intact CARD exhibits approximately 20% higher activity compared to ΔCARD versions where the CARD has been proteolytically removed [15]. This effect persists even in the absence of apoptosome binding, indicating the CARD plays an active role in regulating the catalytic core beyond simple localization. Furthermore, the isolated Apaf-1 CARD can enhance caspase-9 activity by approximately five-fold in vitro, suggesting allosteric effects beyond simple tethering [15].
Experimental Approaches to Studying Procaspase-9 Activation
Key Methodologies and Reagents
Table 2: Essential Research Reagents for Procaspase-9 Studies
| Reagent/Condition | Experimental Function | Application Example | Key Findings Enabled |
|---|---|---|---|
| Engineered Dimeric Caspase-9 | Forces constitutive dimerization via interface mutations | Comparison with apoptosome-activated caspase-9 [4] | Dimerization alone insufficient for full activation; Apaf-1 induces qualitative differences [4] |
| Thrombin-Cleavable Procaspase-9 (pc-9t) | Allows controlled separation of CARD from catalytic domain | Apoptosome binding studies without catalytic domain interference [16] | Confirms CARD-mediated binding is primary apoptosome interaction in low salt [16] |
| Caspase-9 Inhibitor Z-LEHD-FMK | Specific pharmacological inhibition of caspase-9 activity | Functional studies in cellular and animal models [17] | Establishes causal role in pulmonary fibrosis via β-catenin signaling [17] |
| Apoptosome Reconstitution | In vitro assembly with Apaf-1, cytochrome c, dATP | Direct measurement of activation kinetics [16] [18] | Demonstrates ~2000-fold activity enhancement upon apoptosome binding [15] |
| Site-Directed Mutagenesis | Targeted disruption of specific residues or domains | Mapping CARD:core interactions and phosphorylation sites [15] [1] | Identifies Thr125 as key regulatory phosphorylation site [1] |
Experimental Workflow for Activation Studies
Induced Proximity vs. Induced Conformation: Experimental Evidence
The mechanism of procaspase-9 activation has been the subject of extensive debate, with two predominant models emerging from experimental evidence.
The Induced Proximity Model
The original induced proximity model proposed that the apoptosome serves primarily to increase local concentration of procaspase-9 molecules, facilitating dimerization-driven activation [4] [12]. This model posits that caspase-9 zymogens are activated once brought into proximity with each other on the apoptosome platform, with dimerization via intrinsic interfaces being the central activation step [4]. Support for this model comes from observations that caspase-9 exists predominantly as monomers in solution and exhibits basal catalytic activity that increases upon concentration [4].
The Induced Conformation Model
The induced conformation model argues that the apoptosome actively induces structural changes in caspase-9 that generate catalytic competence beyond mere dimerization [4]. This model suggests that apoptosome binding stabilizes the active-site region, leading to a catalytically competent conformation that cannot be achieved through dimerization alone [4]. Recent high-resolution cryo-EM structures show caspase-9 bound to the apoptosome as monomers rather than dimers, supporting conformational activation independent of dimerization [15].
Critical Experimental Evidence
Table 3: Comparative Experimental Evidence for Activation Models
| Experimental Approach | Key Findings | Support for Induced Proximity | Support for Induced Conformation |
|---|---|---|---|
| Engineered Dimeric Caspase-9 | Dimeric form more active than wild-type monomer but significantly less active than apoptosome-activated caspase-9 [4] | Limited support (dimerization increases activity) | Strong support (dimerization insufficient for full activation) |
| Apoptosome-bound Caspase-9 Structure | Cryo-EM reveals monomeric caspase-9 bound to apoptosome with reconfigured active sites [16] [15] | Contradicts (shows monomers, not dimers) | Strong support (demonstrates conformational rearrangement) |
| CARD:Core Domain Interactions | CARD physically interacts with catalytic core, influencing activity independently of apoptosome [15] | Limited relevance | Supports complex intramolecular regulation |
| Activity Comparisons | Apoptosome binding enhances activity ~2000-fold; dimeric engineering provides much smaller increase [15] [4] | Partial support (some activation by dimerization) | Strong support (qualitative difference in activation mechanism) |
| Symmetry Considerations | Apoptosome has 7-fold symmetry while caspase-9 dimerization interface is 2-fold symmetric [4] | Problematic for model | Consistent with asymmetric activation |
Research Reagent Toolkit
Table 4: Essential Research Reagents for Procaspase-9 Studies
| Category | Specific Reagents | Research Applications | Key References |
|---|---|---|---|
| Expression Constructs | Human caspase-9 (1-416) in pET23b; C287A catalytic mutant; CARD-core linker variants | Recombinant protein production; structure-function studies | [15] [4] |
| Activity Assays | Ac-LEHD-AFC fluorogenic substrate; caspase-3 activation assays | Quantitative activity measurements; downstream signaling assessment | [16] [1] |
| Inhibitors/Activators | Z-LEHD-FMK (caspase-9 inhibitor); cytochrome c/dATP (apoptosome assembly) | Functional perturbation studies; pathway modulation | [17] [1] |
| Cell Culture Models | Caspase-9 null embryonic stem cells; MLE-12 alveolar epithelial cells | Physiological context studies; disease modeling | [17] [1] |
| Antibodies | Anti-caspase-9; anti-cleaved-caspase-9 (Asp315); Apaf-1 antibodies | Detection, localization, and quantification in complex systems | [17] |
Implications for Disease and Therapeutic Development
Understanding the precise mechanism of procaspase-9 activation has significant implications for therapeutic interventions in cancer, neurodegenerative disorders, and fibrotic diseases.
Dysregulated caspase-9 function contributes to pathological conditions including cancer development, where diminished caspase-9 activity enables tumor cell survival despite genotoxic stress [1]. Conversely, excessive caspase-9 activation appears to drive disease progression in pulmonary fibrosis, where caspase-9 inhibition attenuates collagen deposition and epithelial apoptosis [17]. The discovery of the caspase-9/β-catenin axis in pulmonary fibrosis reveals non-apoptotic signaling roles that may be therapeutically targeted [17].
The structural insights into CARD:core interactions provide new avenues for allosteric modulation of caspase-9 activity. Small molecules targeting these intramolecular interfaces could offer more precise control than conventional catalytic inhibitors, potentially overcoming the therapeutic challenges posed by the monomeric-dynamic nature of procaspase-9 regulation [15] [1]. As our understanding of procaspase-9 structure and activation mechanisms continues to evolve, so too will opportunities for targeting this critical mediator of programmed cell death in human disease.
The Induced Proximity Model represents a foundational concept in molecular biology, proposing that the activation of specific signaling molecules is driven primarily by their increased local concentration and subsequent dimerization within specialized cellular complexes. First formally proposed in the 1990s, this model emerged from elegant experiments demonstrating that artificially induced clustering of caspase zymogens could trigger their autoprocessing and activation [19]. The model has been particularly influential in explaining the activation mechanism of initiator caspases—proteases that orchestrate programmed cell death (apoptosis)—with caspase-9 serving as a key exemplar. According to this paradigm, the apoptosome, a heptameric complex comprising Apaf-1 and cytochrome c, functions primarily as a molecular platform to concentrate caspase-9 zymogens, facilitating their homodimerization via intrinsic dimerization interfaces and thereby triggering autoactivation [3] [19]. This framework has profoundly shaped understanding of apoptotic signaling and provided theoretical foundations for developing novel therapeutic strategies based on artificially induced proximity.
Competing Theories: Induced Proximity versus Induced Conformation
The Induced Proximity Model, while influential, does not stand unopposed. A competing theoretical framework, the Induced Conformation Model, has gained support from experimental evidence challenging the sufficiency of dimerization for full caspase activation.
Core Theoretical Differences
Induced Proximity Model: This model posits that the primary role of activation complexes like the apoptosome is to increase the local concentration of initiator caspase zymogens. This proximity facilitates homodimerization, which is itself sufficient to trigger caspase autoactivation. The model emphasizes stoichiometry and localization as the critical determinants of activation [19].
Induced Conformation Model: This alternative model suggests that mere dimerization is insufficient. Instead, binding to the activation complex induces specific allosteric changes in the caspase structure that reconfigure the active site into a catalytically competent state. The model emphasizes structural rearrangement as an essential component of activation [3] [12].
The Engineered Dimer Crucible
The critical test between these models came from a seminal experiment that engineered a constitutively dimeric form of caspase-9. Researchers rationally designed a caspase-9 variant by mutating key residues at the dimer interface (e.g., Phe404) that normally create steric hindrance and prevent stable dimerization of the wild-type protein [4]. The crystal structure of this engineered dimer confirmed that it closely resembled the wild-type protein, indicating the mutations did not cause major structural perturbations [3] [4]. When tested, this dimeric caspase-9 exhibited higher catalytic activity than the wild-type monomer, supporting the proximity model's prediction that dimerization enhances activity [4] [20]. However, this dimeric enzyme reached only a small fraction of the catalytic activity achieved by wild-type caspase-9 activated by the Apaf-1 apoptosome. Furthermore, the activity of the engineered dimer could not be significantly enhanced by Apaf-1, unlike the wild-type protein [3] [20]. This key finding suggested that the apoptosome provides more than just a platform for dimerization and posited that it induces a qualitative conformational change essential for full catalytic activation [4].
Experimental Analysis: Direct Comparison of Caspase-9 Activation Models
The debate between induced proximity and induced conformation has been driven by specific, quantifiable experimental data. The table below summarizes the critical quantitative findings from the study of engineered dimeric caspase-9 compared to other activation states.
Table 1: Quantitative Comparison of Caspase-9 Activity Under Different Activation Conditions
| Caspase-9 Form | Catalytic Activity | Apaf-1 Stimulation | Cell Death Induction | Primary Conclusion |
|---|---|---|---|---|
| Wild-Type Monomer | Basal level | Yes, significant enhancement | Lower | Requires apoptosome for full activation |
| Engineered Dimer | Higher than monomer, but low relative to Apaf-1-activated | No significant enhancement | Higher than monomer | Dimerization alone is insufficient for full activity |
| Apaf-1-Activated Wild-Type | Highest activity | Not Applicable | Highest | Apoptosome induces maximal activation beyond dimerization |
Experimental Protocol for Evaluating Caspase Activation
The methodology for generating and testing the engineered caspase-9 dimer involved a multi-step biochemical and cellular approach [4]:
Protein Engineering and Design: Based on structural analysis, residues on the β6 strand of the dimerization interface (e.g., Phe404 in caspase-9) were identified and mutated to relieve steric hindrance, creating a constitutively dimeric caspase-9 (e.g., F404A/C404S mutations).
Protein Expression and Purification: Both wild-type and engineered caspase-9 constructs were expressed in E. coli and purified using affinity and size-exclusion chromatography to homogeneity.
Biophysical Characterization: The oligomeric state of the engineered protein was confirmed using analytical ultracentrifugation or size-exclusion chromatography. The crystal structure was solved to verify no major structural deviations from the wild-type protein.
In Vitro Activity Assay: Catalytic activity was measured using fluorogenic tetrapeptide substrates (e.g., LEHD-AFC). Cleavage of the substrate releases a fluorescent product, allowing kinetic measurement of enzyme activity. Reactions contained purified caspase-9 (wild-type or engineered dimer) in the presence or absence of purified, reconstituted apoptosome (Apaf-1/cytochrome c).
Cellular Apoptosis Assay: Plasmids encoding wild-type or engineered dimeric caspase-9 were transfected into mammalian cells. Cell death was quantified using assays for apoptotic markers, such as DNA fragmentation, caspase-3 activation, or membrane integrity.
The following diagram illustrates the logical flow of the experimental workflow and the core hypotheses tested.
The Scientist's Toolkit: Key Reagents for Proximity Research
Research into caspase activation and induced proximity relies on a specific set of reagents and tools. The table below details several key resources essential for experiments in this field.
Table 2: Essential Research Reagents for Studying Caspase Activation and Induced Proximity
| Reagent / Tool | Function / Description | Application in Research |
|---|---|---|
| Engineered Caspase-9 Dimer | A constitutively dimeric caspase-9 variant with relieved steric hindrance at the dimer interface. | Directly tests the role of dimerization independent of apoptosome assembly [4]. |
| Reconstituted Apoptosome | A purified, in vitro assembled complex of Apaf-1, cytochrome c, and dATP/ATP. | Serves as a positive control for maximal caspase-9 activation in biochemical assays [3]. |
| Fluorogenic Caspase Substrates | Tetrapeptides (e.g., LEHD-AFC) linked to a fluorophore (e.g., AFC). | Quantifies caspase enzymatic activity; cleavage releases the fluorescent group [4]. |
| Cellular Death Assays | Kits and protocols for measuring apoptosis (e.g., TUNEL, Annexin V staining). | Evaluates the functional biological consequence of caspase activation in cells [4]. |
| Molecular Glues (e.g., Thalidomide) | Small molecules that induce proximity between a target protein and an E3 ubiquitin ligase. | Tool compounds for studying chemically induced proximity and targeted protein degradation [21]. |
| PROTACs (Proteolysis-Targeting Chimeras) | Bifunctional molecules with a target-binding ligand linked to an E3 ligase recruiter. | Modular tools for inducing targeted protein degradation via the ubiquitin-proteasome system [21]. |
Implications and Evolving Applications in Drug Discovery
The principles of induced proximity have transcended their original context in apoptosis research, evolving into a cornerstone of modern drug discovery. The recognition that artificially inducing proximity between proteins can powerfully modulate biological function has spawned several innovative therapeutic modalities [21].
The most advanced of these are PROTACs (Proteolysis-Targeting Chimeras) and molecular glue degraders, which function by recruiting a target protein to an E3 ubiquitin ligase, leading to the target's ubiquitination and degradation by the proteasome [21]. As of 2023, approximately 26 PROTAC degraders were in clinical trials, demonstrating the translational potential of this proximity-based approach [21]. The clinical success of molecular glues like thalidomide and its derivatives (lenalidomide, pomalidomide) for treating hematological malignancies provided an early, powerful validation of this concept, even before their mechanism was fully understood [21].
The field continues to expand beyond degradation. New modalities include RIPTACs (RIPTAC is a class of heterobifunctional molecules that aims to inhibit a protein's function by sequestering it in a non-productive complex), with Halda Therapeutics advancing the first RIPTAC, HLD-0915, into clinical trials for prostate cancer in 2025 [22]. These agents seek to provide tissue-selective target inhibition by forming a stable ternary complex that disrupts the target's normal function [22]. Other emerging approaches focus on inducing post-translational modifications like phosphorylation, acetylation, or glycosylation by bringing target proteins close to the relevant modifying enzymes [21].
The following diagram illustrates the signaling pathway of caspase activation in apoptosis, highlighting the central role of the apoptosome and the mechanistic debate.
The investigation into caspase-9 activation reveals a nuanced biological reality where the Induced Proximity Model and the Induced Conformation Model are not mutually exclusive but are likely complementary. Experimental evidence confirms that dimerization, driven by increased local concentration within the apoptosome, is a necessary step that enhances caspase-9 activity beyond its monomeric state [4]. However, the markedly superior activity achieved through Apaf-1 binding demonstrates that dimerization alone is insufficient [3] [20]. The apoptosome appears to function as an allosteric activator, inducing precise conformational changes that optimally configure the caspase-9 active site for efficient catalysis.
This refined understanding underscores the complexity of biological signaling mechanisms. The evolution of induced proximity from a fundamental biological concept to a generative principle for drug discovery highlights the translational power of basic scientific research. As the field advances, solving the high-resolution structure of the complete apoptosome and other activation complexes will be essential to fully elucidate the precise structural changes that drive initiator caspase activation [3]. This will not only resolve the long-standing mechanistic debate but also pave the way for the next generation of proximity-based therapeutics that can manipulate protein complexes and cellular fate with ever-greater precision.
For decades, the induced proximity model served as the prevailing explanation for initiator caspase activation. This model, formally proposed in the late 1990s, posited that initiator caspase zymogens possess low intrinsic activity and are activated primarily when oligomeric signaling complexes, such as the Apaf-1 apoptosome, bring them into close proximity, facilitating autoprocessing through homodimerization [19]. This framework dominated the field until experimental evidence began to reveal inconsistencies, particularly concerning the activation of caspase-9.
This guide objectively compares the induced conformation model against the induced proximity model for caspase-9 activation. We present supporting experimental data, detailed methodologies, and key research tools to provide scientists and drug development professionals with a clear, evidence-based resource on this fundamental apoptotic mechanism.
Model Comparison: Core Principles and Key Differentiators
The following table outlines the fundamental differences between the two competing models.
Table 1: Fundamental Comparison of the Induced Proximity and Induced Conformation Models
| Feature | Induced Proximity Model | Induced Conformation Model |
|---|---|---|
| Core Principle | Activation is driven by increased local concentration, promoting homodimerization [19] [23]. | Activation is driven by allosteric regulation via the apoptosome, inducing an active conformation [24] [4]. |
| Role of Apoptosome | Scaffold to increase local caspase-9 concentration for dimerization [25]. | Allosteric regulator that acts as a cofactor [26] [24]. |
| Active Caspase-9 State | Stable homodimer [23]. | Caspase-9/Apaf-1 holoenzyme; caspase-9 remains bound to the apoptosome [24]. |
| Predicted Symmetry | Not critical; any oligomeric state that promotes dimerization could suffice. | The heptameric symmetry of the apoptosome is likely critical for inducing the correct conformational change [26] [4]. |
Experimental Evidence: Quantitative Data Supporting Induced Conformation
Critical experiments have directly tested these models, generating quantitative data that challenge the simple induced proximity explanation.
Table 2: Key Experimental Evidence and Supporting Data for the Induced Conformation Model
| Experimental Approach | Key Finding | Quantitative Result | Interpretation |
|---|---|---|---|
| Engineered Dimeric Caspase-9 [4] | A constitutively dimeric caspase-9 mutant was created. Its activity was much lower than that of Apaf-1-activated caspase-9 and was not enhanced by Apaf-1. | The catalytic activity of the dimer was "only a small fraction" of the apoptosome-activated wild-type enzyme. | Dimerization alone is insufficient to recapitulate full activation, implying an additional regulatory role for Apaf-1 [4]. |
| Holoenzyme Activity Assay [24] | The proteolytic activity of caspase-9 was compared when free versus in a complex with APAF-1. | The caspase-9/APAF-1 holoenzyme was at least 1,000-fold more active than the free, processed caspase-9. | Caspase-9 is fully active only when bound to APAF-1, supporting an allosteric holoenzyme mechanism [24]. |
| Multimeric CARD Complex [26] | A previously unknown interface was identified as essential for forming a multimeric CARD assembly between Apaf-1 and caspase-9. | A 1:1 CARD complex was insufficient for activation; a higher-order oligomeric assembly was required. | The apoptosome activates caspase-9 through multiple specific interfaces, inducing a conformational change [26]. |
| Caspase-9 Homo-/Heterodimers [23] | The apoptosome was shown to mediate the formation of both caspase-9 homodimers and caspase-9/Apaf-1 heterodimers. | Uncleaved procaspase-9 had a higher affinity for homodimerization, while cleavage triggered a "molecular timer" for release. | Activation involves a complex interplay of both dimerization and allosteric interactions within the apoptosome [23]. |
Experimental Protocols: Methodologies for Key Studies
To enable critical evaluation and replication, this section details the methodologies behind pivotal experiments.
Engineering a Constitutive Caspase-9 Dimer
This experiment directly tested whether forced dimerization was sufficient for full activation [4].
- Rational Design: Based on structural analysis, researchers replaced five consecutive amino acids on the β6 strand of caspase-9 (Gly402-Cys-Phe-Asn-Phe406) with the corresponding residues from the strong dimerizer caspase-3 (Cys264-Ile-Val-Ser-Met268). This was designed to relieve steric hindrance at the dimer interface.
- Protein Expression & Purification: The engineered caspase-9 gene was expressed in E. coli, and the protein was purified using standard chromatography techniques (e.g., affinity and size-exclusion chromatography).
- Crystallography: The crystal structure of the engineered dimer was solved to confirm that the overall structure, including asymmetric monomer details, resembled the wild-type protein, ensuring the mutation did not cause unintended structural perturbations.
- Activity Assays:
- In Vitro: Catalytic activity of the engineered dimer was measured against peptide substrates and compared to wild-type caspase-9 activated in a reconstituted Apaf-1 apoptosome.
- In Cellulo: The engineered dimer and wild-type caspase-9 were expressed in mammalian cells, and their relative potency to induce apoptotic cell death was quantified.
Isolating and Comparing the Caspase-9 Holoenzyme
This approach demonstrated that the apoptosome-bound caspase-9 is orders of magnitude more active than the free enzyme [24].
- Cell-Free Apoptosis System: Extracts from 293 cells were activated with dATP/ATP to form the apoptosome and activate caspase-9.
- Immunoprecipitation (IP): Caspase-9 was immunoprecipitated from both active and inactive cell extracts.
- Sedimentation Analysis: Activated cell extracts were fractionated by density gradient sedimentation to separate the high-molecular-weight caspase-9/Apaf-1 holoenzyme from free caspase-9.
- Functional Rescue Assay: Extracts were depleted of endogenous caspase-9. The ability of added free caspase-9 (both precursor and processed forms) to restore caspase-3 activation was tested, with and without dATP to control for apoptosome formation.
Signaling Pathways and Experimental Workflows
The following diagrams illustrate the core concepts and experimental logic using the DOT language.
Diagram 1: Model Comparison of Caspase-9 Activation
Diagram 2: Logic Flow of the Engineered Dimer Experiment
The Scientist's Toolkit: Key Research Reagents
Table 3: Essential Reagents for Studying Caspase-9 Activation Mechanisms
| Research Reagent / Tool | Function & Application | Key Insight from Usage |
|---|---|---|
| Apaf-1-591 (Truncated Apaf-1) [26] | A constitutively active Apaf-1 variant (residues 1-591) that retains the ability to form a functional apoptosome without the regulatory WD40 domain. | Enabled simplified reconstitution of apoptosome complexes for biochemical and structural studies [26]. |
| "Frozen" Caspase Zymogen (e.g., C285A mutant) [19] | A catalytically inactive caspase mutant (active site cysteine mutated to alanine) that cannot undergo autoprocessing. | Allowed for the first direct measurement of the intrinsic catalytic activity of an unprocessed initiator caspase zymogen [19]. |
| "Miniapoptosome" (ApCARD–GroES fusion) [26] | A synthetic, stable heptameric scaffold created by fusing the Apaf-1 CARD domain (ApCARD) to the oligomeric protein GroES. | Demonstrated that an ApCARD-linked scaffold is sufficient to potently activate caspase-9, isolating the CARD-mediated activation step from the rest of the apoptosome [26]. |
| Z-LEHD-FMK [7] | A cell-permeable, irreversible caspase-9 specific inhibitor. | Used in pharmacological studies (in vitro and in vivo) to dissect the specific role of caspase-9 activity in apoptotic and non-apoptotic processes, such as pulmonary fibrosis [7]. |
| Kosmotropic Salts (e.g., Ammonium Citrate) [25] [4] | High concentrations of these salts can artificially induce dimerization and activation of initiator caspases like caspase-9 in solution. | Serves as a tool to enforce dimerization, allowing comparison between dimer-induced activity and apoptosome-induced activity [25] [4]. |
The induced proximity hypothesis, introduced by Salvesen and Dixit, proposed a foundational model for understanding the activation of initiator caspases, the proteases that orchestrate programmed cell death (apoptosis) [13] [27]. This model posited that initiator caspases, which exist as inactive monomers (zymogens) in healthy cells, undergo autoprocessing and activation simply by being brought into close proximity with one another on large oligomeric signaling platforms, such as the apoptosome [3] [4]. For caspase-9, the initiator of the intrinsic apoptotic pathway, the apoptosome—a heptameric complex of Apaf-1 and cytochrome c—was thought to function primarily as a concentrator, elevating the local concentration of caspase-9 to drive proximity-driven homodimerization and subsequent activation [3] [4]. This model served as the dominant paradigm for years, providing an elegant, generalized mechanism.
However, subsequent rigorous biochemical and structural investigations have challenged the simplicity of this model, leading to significant refinements. A pivotal re-evaluation, spearheaded by the work of Yigong Shi and colleagues, introduced the induced conformation model, suggesting that activation involves specific allosteric changes induced by the apoptosome, going beyond mere dimerization [3] [4]. This article traces the evolution of this concept, comparing the foundational induced proximity model against modern evidence for an induced conformational mechanism, providing researchers with a clear comparison of the supporting experimental data and protocols.
The Foundational Model: Core Tenets of Induced Proximity
The original induced proximity model was built upon several key observations and principles. The central premise was that the intrinsic dimerization of caspase-9 is energetically unfavorable, requiring an external catalyst [4]. The model asserted that the apoptosome's primary role was to act as a passive scaffolding structure, increasing the local concentration of caspase-9 monomers to a point where homodimerization becomes favorable [3]. This dimerization event was itself considered sufficient to trigger the catalytic activity of the enzyme. The model effectively generalized the activation mechanism for initiator caspases, framing it as a consequence of collective assembly rather than specific protein-protein interactions, with the symmetry of the oligomeric platform (e.g., the seven-fold symmetric apoptosome) serving to facilitate this clustering [4].
A Paradigm Challenged: Key Experimental Evidence for Induced Conformation
The induced proximity model began to face challenges when experimental data could not be fully explained by dimerization alone. The critical evidence came from a series of elegant experiments engineered to test the model's predictions directly.
The Engineered Dimeric Caspase-9 Experiment
A landmark study by Chao et al. (2005) directly tested the induced proximity model by creating a constitutively dimeric form of caspase-9, bypassing the need for the apoptosome [4].
- Experimental Rationale: If induced proximity (via dimerization) is the sole mechanism for activation, then a constitutively dimeric caspase-9 should display catalytic activity equivalent to that of the Apaf-1-activated wild-type caspase-9 [4].
- Engineering Methodology: The researchers designed a dimeric caspase-9 by mutating key residues at the dimer interface (based on the interface of the effector caspase-3) to relieve steric hindrance that naturally prevents stable dimerization in the wild-type protein. Crucially, X-ray crystallography confirmed that the engineered dimer's structure closely resembled the wild-type caspase-9, ensuring that the mutations did not cause unintended structural alterations [4].
- Key Findings and Quantitative Data: The engineered dimer exhibited higher activity than the monomeric wild-type but only a fraction of the activity achieved when the wild-type was activated by Apaf-1. Furthermore, the activity of the dimeric caspase-9 could not be significantly enhanced by Apaf-1, unlike the wild-type protein [3] [4].
Table 1: Key Findings from the Engineered Dimeric Caspase-9 Experiment
| Parameter | Wild-Type Monomeric Caspase-9 | Engineered Dimeric Caspase-9 | Apaf-1-Activated Wild-Type Caspase-9 |
|---|---|---|---|
| Basal Catalytic Activity | Low basal activity [4] | Higher than wild-type, but still low [3] [4] | Very High (1000-fold increase over monomer) [28] |
| Response to Apaf-1 | Activity dramatically stimulated [3] [4] | No significant enhancement [3] [4] | Not Applicable (Definition of full activation) |
| Cell Death Induction | Baseline level | More efficient than wild-type [4] | Highly efficient [4] |
The DNA Origami Apoptosome-Mimic
More recent research using cutting-edge DNA nanotechnology has provided further, nuanced support for a hybrid model. A 2020 study created a synthetic, DNA origami-based version of the apoptosome to precisely control the spatial organization of caspase-9 monomers [28].
- Experimental Rationale: To isolate the effect of proximity from other potential allosteric influences of the native apoptosome by using a programmable platform that mimics only its scaffolding function.
- Methodology: The CARD domain of caspase-9 was replaced with a synthetic DNA oligonucleotide. This allowed for the tethering of individual caspase-9 catalytic domains to a DNA origami scaffold at defined positions and stoichiometries, controlling inter-enzyme distance and orientation with nanometer accuracy [28].
- Key Findings: The study confirmed that proximity-induced dimerization is a key driver of activation, as bringing two monomers together on the scaffold enhanced activity. However, it also revealed that higher-order oligomerization (clusters of three or four enzymes) produced a further, significant boost in activity, suggesting a multivalent effect beyond simple dimerization. Furthermore, experiments with heterodimers of wild-type and catalytically dead mutants provided direct evidence for half-of-sites reactivity, an asymmetric mechanism where the two active sites in a dimer function differently [28].
Table 2: Insights from DNA Origami-Based Caspase-9 Activation Studies
| Experimental Condition | Observed Effect on Caspase-9 Activity | Implication for Activation Model |
|---|---|---|
| Monomeric (untethered) | Low basal activity [28] | Confirms innate inactivity of monomers. |
| Forced Dimerization | Significant activity increase [28] | Supports proximity-driven dimerization as a key step. |
| Trimer/Tetramer Clustering | Activity enhanced beyond dimer level [28] | Suggests a role for higher-order oligomerization (refinement to model). |
| Wild-type/Mutant Heterodimer | Activity similar to wild-type homodimer [28] | Supports asymmetric, induced conformation within the dimer. |
Comparative Analysis: Model Evolution at a Glance
The collective evidence has driven a refinement of the original hypothesis, leading to a more sophisticated understanding of caspase-9 activation.
Table 3: Comparison of the Induced Proximity and Induced Conformation Models
| Aspect | Original Induced Proximity Model | Modern Induced Conformation/Refined Model |
|---|---|---|
| Core Mechanism | Proximity-driven dimerization is necessary and sufficient for activation [4]. | Dimerization is necessary but not sufficient; activation requires an allosteric conformational change induced by the apoptosome [3] [4]. |
| Role of Apoptosome | Passive scaffold to increase local caspase-9 concentration [3]. | An active regulatory platform that may induce a specific, activating conformation in caspase-9 [3] [13]. |
| Nature of Active Caspase-9 | A symmetric homodimer. | Likely an asymmetric homodimer with evidence for half-of-sites reactivity [28]. |
| Predicted Activity of a Constitutive Dimer | Should be identical to apoptosome-activated caspase-9 [4]. | Should be less active than apoptosome-activated caspase-9, as it lacks the induced conformation [3] [4]. |
| Supporting Evidence | Generalization from experiments on other caspases; ability of oligomeric platforms to activate caspases [27]. | Engineered dimer experiment [4]; structural studies; DNA origami studies showing oligomerization effects [28]. |
The Scientist's Toolkit: Essential Reagents and Methods
Research in this field relies on a suite of specialized reagents and experimental protocols.
Table 4: Key Research Reagent Solutions for Studying Caspase-9 Activation
| Reagent / Assay | Function and Application | Experimental Utility |
|---|---|---|
| Caspase-9 Antibody | Detects and quantifies caspase-9 protein levels, cleavage, and localization via Western blot, immunohistochemistry, and flow cytometry [29]. | Essential for confirming expression, monitoring proteolytic processing, and cellular localization in response to apoptotic stimuli. |
| LEHD-AFC (Fluorogenic Substrate) | Synthetic peptide substrate (LEHD) conjugated to a fluorophore (AFC). Caspase-9 cleaves the peptide, releasing AFC, which emits fluorescence [28]. | Allows quantitative, real-time measurement of caspase-9 enzymatic activity in vitro (e.g., in purified systems with apoptosome or DNA origami). |
| Recombinant Apaf-1 & Cytochrome c | Purified proteins required for the in vitro reconstitution of the apoptosome complex [3] [4]. | The gold-standard biochemical system for studying direct caspase-9 activation in a controlled environment. |
| DNA Origami Scaffold | Programmable nanoscale DNA structure functionalized with oligonucleotide "handles" [28]. | Used as a reductionist synthetic platform to study proximity effects by positioning caspase-9-DNA conjugates with precise control over number, distance, and orientation. |
| Caspase-9 Enzyme-DNA Conjugate | Caspase-9 catalytic domain site-specifically conjugated to a DNA oligonucleotide via non-canonical amino acid incorporation and click chemistry [28]. | Enables the tethering of caspase-9 to DNA origami scaffolds, crucial for spatial organization experiments. |
Detailed Experimental Protocol: In Vitro Reconstitution of Apoptosome-Mediated Caspase-9 Activation
This classic biochemistry protocol is used to demonstrate and quantify the activation of caspase-9 by the apoptosome.
- Apoptosome Assembly: Incubate recombinant, purified Apaf-1 protein with cytochrome c and dATP in an appropriate buffer (e.g., containing KCl and HEPES pH 7.5) for 30-60 minutes at 30°C. This facilitates the oligomerization of Apaf-1 into the heptameric apoptosome complex [4].
- Caspase-9 Activation: Add purified, monomeric procaspase-9 zymogen to the assembled apoptosome and incubate further to allow recruitment and activation.
- Activity Measurement: Initiate the enzymatic reaction by adding the fluorogenic substrate LEHD-AFC to the mixture. The final reaction volume is typically 100 µL.
- Kinetic Analysis: Transfer the reaction mixture to a microplate reader and measure the fluorescence emission (e.g., at 505 nm upon excitation at 400 nm) continuously over 30-60 minutes.
- Data Calculation: Enzyme velocity is determined from the linear slope of the fluorescence increase over time. Activity can be reported as the rate of substrate cleavage (pmol/min) or normalized to enzyme concentration. Michaelis-Menten parameters (KM, Vmax) can be derived by varying the substrate concentration [28].
Visualizing the Molecular Pathways and Models
The following diagrams illustrate the key apoptotic pathway and the evolution of caspase-9 activation models.
The Intrinsic Apoptotic Pathway
Diagram 1: The intrinsic apoptotic pathway and the central role of the apoptosome.
Evolution of Caspase-9 Activation Models
Diagram 2: The evolution of caspase-9 activation models, from simple dimerization to a conformation-based mechanism.
The investigation into caspase-9 activation exemplifies the dynamic nature of scientific models. The journey from the original induced proximity hypothesis to the modern induced conformation model highlights how rigorous experimentation can refine our understanding of fundamental biological processes. The current evidence supports a synthesized model where the apoptosome first drives dimerization through proximity, but full catalytic potency is achieved only through an subsequent allosteric conformational change, potentially involving asymmetric active sites and enhanced by higher-order clustering [3] [4] [28]. This refined understanding has profound implications for drug discovery, particularly in cancer, where reactivating apoptosis in malignant cells is a key therapeutic goal. Targeting the specific interactions that facilitate the induced conformation on the apoptosome could offer new avenues for developing more precise and effective pro-apoptotic drugs.
Decoding Activation: Key Techniques and Models in Caspase-9 Research
In the intrinsic apoptotic pathway, caspase-9 functions as the initiator caspase, triggering a cascade that leads to programmed cell death. For decades, the central mechanistic question has been how this enzyme is activated within the multi-protein "apoptosome" complex. Two competing hypotheses have dominated this field: the induced proximity model, which posits that the apoptosome merely serves to bring caspase-9 monomers into close proximity, facilitating dimerization and consequent activation; and the induced conformation model, which proposes that binding to the apoptosome actively induces a conformational change essential for full catalytic activity [30] [31] [1]. To critically evaluate these models, researchers have employed protein engineering to create constitutively dimeric caspase-9 variants, bypassing the need for the apoptosome. These engineered dimers have become crucial tools for dissecting the precise molecular mechanism of caspase-9 activation.
Engineering Strategy and Structural Rationale
The Dimerization Interface as an Engineering Target
Wild-type caspase-9 exists predominantly as an inactive monomer in solution, with a very low propensity to dimerize [30] [32]. In contrast, effector caspases like caspase-3 are stable dimers. To create a constitutively dimeric caspase-9, researchers performed a detailed comparative analysis of the dimerization interfaces across different caspases [30] [4]. The interface is primarily mediated by two central β-strands (β6 and β6'), one from each monomer. Although the overall structure is conserved, the sequence of the β6 strand varies significantly.
A key finding was that five consecutive amino acids in the β6 strand of caspase-9 (Gly402-Cys-Phe-Asn-Phe406) differ substantially from the corresponding residues in caspase-3 (Cys264-Ile-Val-Ser-Met268) [30] [4]. Structural analysis revealed that the side chains of Phe404 and Phe404' in caspase-9 create steric hindrance, physically impeding stable dimer formation [30] [4]. This provided a clear engineering strategy: replace the caspase-9-specific β6 strand sequence with the corresponding sequence from caspase-3 to relieve this steric clash and promote constitutive dimerization.
Design and Validation of the Dimeric Variant
The engineered caspase-9 was created by substituting its five-residue β6 strand sequence (Gly402-Cys-Phe-Asn-Phe406) with the caspase-3 sequence (Cys264-Ile-Val-Ser-Met268) [30]. The oligomeric state of the wild-type and engineered proteins was analyzed using size exclusion chromatography and analytical ultracentrifugation [30].
Table 1: Oligomeric State Analysis of Wild-type vs. Engineered Caspase-9
| Protein | Size Exclusion Elution Volume | Calculated Molecular Mass | Analytical Ultracentrifugation |
|---|---|---|---|
| Wild-type Caspase-9 | ~60 kDa | ~50,550 Da (Monomer) [30] | |
| Engineered Caspase-9 | ~120 kDa | ~91,030 Da (Dimer) [30] |
The crystal structure of the engineered dimer confirmed that the overall structure, including the asymmetric nature of the two monomers, closely resembled that of the wild-type protein, indicating that the mutation specifically affected dimerization propensity without causing major structural perturbations [30] [4].
Functional Comparison: Engineered Dimer vs. Apoptosome-Activated Caspase-9
The critical test for the induced proximity model was to compare the catalytic activity of the engineered dimeric caspase-9 with that of wild-type caspase-9 activated by the Apaf-1 apoptosome.
Catalytic Activity and Apoptotic Potential
Biochemical and cellular assays revealed that the engineered dimer is functional but qualitatively different from the apoptosome-activated enzyme.
Table 2: Functional Comparison of Caspase-9 Activation States
| Parameter | Wild-type Monomer | Engineered Dimer | Apaf-1-Activated Wild-type |
|---|---|---|---|
| Catalytic Activity | Low basal activity [30] | Higher than wild-type, but only a small fraction of Apaf-1-activated activity [30] | Highest catalytic activity [30] |
| Apoptotic Potential | Low cell death induction [30] | More efficient cell death induction than wild-type [30] | Full apoptotic activation |
| Apaf-1 Enhancement | Activity significantly enhanced by Apaf-1 [30] | Activity not significantly enhanced by Apaf-1 [30] | N/A |
Implications for the Activation Mechanism
The data demonstrate that while dimerization enhances caspase-9 activity beyond its monomeric basal level, it is insufficient to recapitulate the full catalytic power conferred by the apoptosome. This key finding challenges a strict interpretation of the induced proximity model, which predicts that a stable dimer should be as active as the apoptosome-bound form [30]. The results instead suggest that the apoptosome does more than just concentrate caspase-9 monomers; it likely induces a specific, active conformation that the intrinsic dimerization interface alone cannot achieve, supporting an induced conformation model [30] [1].
Experimental Protocols for Key Assays
Protocol: Site-Directed Mutagenesis for Dimeric Caspase-9
This protocol outlines the creation of the constitutively dimeric caspase-9 variant [30] [4].
- Template Preparation: Use a plasmid containing the gene for human procaspase-9 (residues 1-416).
- Primer Design: Design mutagenic primers to replace the nucleotide sequence encoding the β6 strand residues (Gly402-Cys-Phe-Asn-Phe406) with the sequence encoding the caspase-3 residues (Cys-Ile-Val-Ser-Met).
- Mutagenesis Reaction: Perform a standard PCR-based site-directed mutagenesis protocol.
- Transformation and Selection: Transform the PCR product into competent E. coli cells, plate on selective antibiotic media, and incubate overnight.
- Sequence Verification: Pick several colonies, isolate plasmid DNA, and perform DNA sequencing across the mutated region to confirm the correct nucleotide substitution.
Protocol: Analyzing Oligomeric State by Size Exclusion Chromatography
This protocol is used to determine if the engineered caspase-9 is a monomer or dimer [30].
- Protein Purification: Overexpress and purify the wild-type and engineered caspase-9 proteins from bacteria to homogeneity.
- Column Equilibration: Equilibrate a size exclusion chromatography column (e.g., Superdex 200) with a suitable buffer (e.g., 20 mM Tris, 100 mM NaCl, pH 8.0).
- Sample Loading and Elution: Load the purified protein onto the column and elute isocratically at a constant flow rate (e.g., 1 ml/min), collecting fractions.
- Analysis: Monitor the elution profile by UV absorbance. Compare the elution volumes of the wild-type and engineered proteins to a calibration curve of standard proteins with known molecular masses to estimate the oligomeric state.
Protocol: Caspase-9 Catalytic Activity Assay
This protocol measures the enzymatic activity of caspase-9 variants using a fluorogenic substrate [32].
- Reaction Setup: Prepare an assay buffer (e.g., 10 mM Pipes, 0.1 M NaCl, 0.1 mM EDTA, 10 mM DTT, 10% sucrose, 0.1% CHAPS, pH 7.2).
- Enzyme Addition: Add the caspase-9 protein (wild-type, engineered dimer, or Apaf-1-activated) to the buffer.
- Substrate Addition: Initiate the reaction by adding the fluorogenic substrate Ac-LEHD-AFC (acetyl-Leu-Glu-His-Asp-7-amino-4-trifluoromethyl coumarin). The caspase cleaves the substrate, releasing the AFC fluorophore.
- Measurement: Monitor the increase in fluorescence (excitation ~405 nm, emission ~510 nm) over time using a plate reader or fluorometer at 37°C.
- Data Analysis: Calculate the initial velocity from the linear portion of the progress curve to determine catalytic activity.
Signaling Pathways and Experimental Workflows
The following diagram illustrates the core hypotheses and the experimental approach used to test them.
The Scientist's Toolkit: Key Research Reagents
Table 3: Essential Reagents for Caspase-9 Dimerization Research
| Reagent / Material | Function / Application | Specific Example |
|---|---|---|
| Caspase-9 Expression Constructs | Template for producing wild-type and mutant proteins in bacterial or mammalian systems. | Plasmid containing human procaspase-9 (residues 1-416) [30]. |
| Site-Directed Mutagenesis Kit | Method for introducing specific point mutations to alter the dimerization interface. | Kits for replacing caspase-9 β-strand residues with caspase-3 equivalents [30] [4]. |
| Size Exclusion Chromatography Column | Tool for separating proteins by size and analyzing oligomeric state. | Superdex 200 column for distinguishing monomeric vs. dimeric caspase-9 [30] [32]. |
| Fluorogenic Caspase Substrate | Sensitive probe for measuring caspase-9 enzymatic activity. | Ac-LEHD-AFC (cleavage releases fluorescent AFC moiety) [32]. |
| Apaf-1 Protein / Cytochrome c | Components required for in vitro reconstitution of the apoptosome complex. | Used to activate wild-type caspase-9 for comparative activity assays [30]. |
| Covalent Caspase Inhibitors | Tool for trapping and stabilizing transient active states, such as dimers. | Z-VAD-FMK (benzoxycarbonyl-Val-Ala-Asp-fluoromethyl ketone) [32]. |
Caspase-9 stands as a critical initiator caspase that regulates the intrinsic apoptotic pathway, a fundamental process for cellular homeostasis and elimination of damaged cells. The activation mechanism of caspase-9 has been a subject of extensive research, primarily centering on two competing hypotheses: "induced proximity" versus "induced conformation." The induced proximity model posits that caspase-9 activation occurs through increased local concentration and homodimerization when recruited to the apoptosome scaffold. In contrast, the induced conformation model suggests that the apoptosome directly allosterically activates caspase-9. Structural biology techniques, particularly X-ray crystallography and NMR spectroscopy, have provided indispensable but distinct insights into this debate, each offering unique advantages and limitations for studying caspase-9 complexes. This guide objectively compares the performance of these techniques in elucidating the structure and activation mechanism of caspase-9.
Technique Comparison: X-ray Crystallography vs. NMR for Caspase-9 Studies
Table 1: Direct Comparison of X-ray Crystallography and NMR for Caspase-9 Complex Research
| Feature | X-ray Crystallography | NMR Spectroscopy |
|---|---|---|
| Optimal System Size | Well-ordered, large complexes (>100 kDa) [5] | Smaller proteins, dynamic regions, flexibly-thered domains in large complexes (up to 1.3 MDa) [5] |
| Sample State | Crystalline solid state [33] | Solution state (near-physiological conditions) [5] |
| Key Caspase-9 Findings | Static structure of CARD domain and its interaction with Apaf-1; static view of protease domain dimer interface [5] | Apoptosome-bound protease domain remains monomeric until substrate binding; measures weak dimerization constants (K~d~ in mM range) [5] |
| Resolution & Atomic Detail | High (atomic level), precise atomic coordinates [33] | High for backbone and methyl groups, residue-specific information [5] [34] |
| Insight into Dynamics | Limited; time- and space-averaged structure [33] | Excellent; residue-specific dynamics and conformational changes in solution [5] [34] |
| Key Technical Limitation | Requires high-quality crystals; flexible regions often poorly resolved or missing [5] [33] | Sensitivity and resolution decrease with molecular size; requires isotope labeling [5] [34] |
Detailed Experimental Protocols
Methyl-TROSY NMR for Studying the Apoptosome-Caspase-9 Complex
This NMR approach has been pivotal in studying the massive, ~1.3 MDa apoptosome complex, revealing the dynamic behavior of caspase-9.
- Protein Expression and Isotope Labeling: Caspase-9 is expressed in E. coli grown in minimal media. For Methyl-TROSY, the protein is labeled with
[13CH3]on isoleucine, leucine, and valine methyl groups in a highly deuterated background. This labeling scheme dramatically simplifies spectra and enhances sensitivity for large complexes [5]. - Apoptosome Reconstitution: The native apoptosome is reconstituted in vitro by incubating Apaf-1 with cytochrome c and dATP in a suitable buffer. Isotope-labeled caspase-9 is then recruited to this complex to form the ~1.3 MDa holo-complex [5].
- NMR Data Collection: Spectra are acquired on high-field NMR spectrometers (e.g., 600-900 MHz) using specially optimized pulse sequences like
[1H,13C]-correlationspectra with TROSY (Transverse Relaxation-Optimized Spectroscopy). These sequences minimize line broadening, which is a major challenge for large molecules [5]. - Data Interpretation: The number, position, and intensity of peaks in the NMR spectrum report on the caspase-9 protease domain's conformational state (monomeric vs. dimeric) and dynamics. The addition of substrate mimics (e.g., Z-LEHD-fmk) allows for trapping and observing the active state [5].
X-ray Crystallography of Caspase-9 and Associated Domains
X-ray crystallography has provided high-resolution snapshots of caspase-9 domains and their interactions.
- Protein Purification and Crystallization: The caspase-9 protease domain or CARD domain is purified to homogeneity. A critical step involves screening thousands of conditions to find the right combination of precipitant, buffer, and temperature that yields well-ordered, diffraction-quality crystals. For the CARD-CARD interaction between Apaf-1 and caspase-9, a heterodimeric complex was crystallized [35].
- Crystal Freezing (Cryo-cooling): Crystals are flash-frozen in liquid nitrogen, typically with a cryo-protectant (e.g., glycerol) to prevent ice formation [35].
- X-ray Data Collection and Processing: A crystal is exposed to a high-intensity X-ray beam, and the resulting diffraction pattern is collected on a detector. The data is processed to determine the crystal's symmetry (space group) and the intensity of diffraction spots [33].
- Structure Solution and Refinement: Molecular replacement, using a known similar structure (e.g., another CARD domain), is often used to phase the diffraction data. The initial model is then iteratively refined against the diffraction data to build and optimize the final atomic structure [35].
Visualizing Caspase-9 Activation and Structural Analysis
Caspase-9 Activation Pathway
Structural Techniques Workflow
The Scientist's Toolkit: Key Research Reagents and Materials
Table 2: Essential Research Reagents for Caspase-9 Structural Studies
| Reagent/Material | Function in Research | Example Use Case |
|---|---|---|
| Isotope-labeled Nutrients (15N-NH4Cl, 13C-Glucose, D2O) | Incorporation of NMR-active isotopes into proteins for detection. | Producing 2H/13CH3-labeled caspase-9 for Methyl-TROSY studies of the apoptosome [5] [34]. |
| Apoptosome Components (Recombinant Apaf-1, Cytochrome c) | In vitro reconstitution of the physiological activation platform for caspase-9. | Studying caspase-9 recruitment and activation mechanism in a native-like complex [5]. |
| Substrate Mimics / Inhibitors (Z-LEHD-fmk) | Irreversibly traps caspase-9 in an active conformation by covalently modifying the catalytic cysteine. | Stabilizing the active dimeric state for both NMR and X-ray studies [5]. |
| Crystallization Screening Kits | Pre-formulated solutions to identify initial conditions for protein crystallization. | Finding conditions to grow diffraction-quality crystals of caspase-9 domains or complexes [35]. |
| Engineered Apoptosome Mimics (Heptameric Proteasome Scaffold) | Provides a simpler, more stable scaffold to study caspase-9 crowding and activation. | Validating that activation is due to crowding rather than specific Apaf-1 interactions [5]. |
The combined application of X-ray crystallography and NMR spectroscopy has been instrumental in advancing our understanding of caspase-9 activation, providing compelling evidence that reconciles the induced proximity and induced conformation models. X-ray crystallography offered the foundational, high-resolution structures of caspase-9 domains and critical interfaces. However, it was the unique capability of NMR spectroscopy to probe dynamics and weak interactions in solution that revealed a more nuanced mechanism: the apoptosome scaffold primes caspase-9 for activation by organizing its protease domains (a form of induced proximity), but extensive dimerization and full activation are triggered only upon substrate binding (an induced conformational change) [5]. This synergistic finding, enabled by the complementary strengths of both techniques, highlights that caspase-9 activation is not a simple binary switch but a tightly regulated, multi-step process. For drug development professionals targeting caspase-9 in diseases like cancer or pulmonary fibrosis [17], this refined mechanism opens avenues for therapeutic intervention at specific stages of the activation pathway.
The long-standing debate regarding the activation mechanism of initiator caspases, oscillating between the induced proximity and induced conformation models, has been a central focus in apoptosis research. This review employs a product comparison framework to evaluate a DNA origami-based synthetic apoptosome against traditional biological and biochemical methods for studying caspase-9 activation. We objectively compare the performance of this bottom-up nanotechnology platform in controlling protein dimerization, elucidating activation mechanisms, and enabling precise spatial organization. Supporting experimental data demonstrates that DNA origami not only recapitulates fundamental biological phenomena but also introduces unique capabilities for engineering and analyzing signaling complexes with an unprecedented level of control.
The apoptosome is a quintessential signaling platform in the intrinsic apoptosis pathway, responsible for activating caspase-9 following cellular stress. For decades, the central mechanistic question has revolved around how this multi-protein complex regulates the initiator caspase. The induced proximity model posits that activation occurs through dimerization of caspase-9 monomers when brought into close proximity by the apoptosome scaffold [23]. Competing evidence has alternatively supported an induced conformation or holoenzyme model, suggesting that the apoptosome induces allosteric changes within monomeric caspase-9 to enhance its activity [23].
Resolving this debate requires precise control over the spatial organization of caspase-9 molecules to systematically probe the relationship between proximity and activity. Traditional approaches, including protein-based scaffolds and enforced dimerization through kosmotropic salts or chimeric proteins, have provided valuable insights but face limitations in precision, scalability, and programmability [23]. These limitations have motivated the development of engineered platforms that offer greater control over the molecular arrangement of signaling components.
Platform Comparison: DNA Origami vs. Traditional Methodologies
The search for an ideal platform to study proximity-induced activation has led to the development of various technological approaches. The table below provides a systematic comparison of the DNA origami-based synthetic apoptosome with conventional methods.
Table 1: Performance comparison of platforms for studying caspase-9 activation
| Platform Feature | DNA Origami-Based Apoptosome | Native Apoptosome | Protein-Based Scaffolds | Enforced Dimerization (e.g., Kosmotropic Salts) |
|---|---|---|---|---|
| Spatial Control | Nanometer-precision over enzyme placement [28] | Defined by protein structure; limited modifiability | Moderate control; limited by protein design | No control; induces random dimerization |
| Programmability | High; fully programmable handle positions [28] | None; fixed native structure | Low; requires extensive protein engineering | None |
| Dimerization Control | Precise control over inter-enzyme distances and orientations [28] | Recruits up to four monomers with flexible tethering [28] | Designed for bivalent dimerization | Non-specific, solution-wide dimerization |
| Mechanistic Insight | Isolates proximity effects from allostery; tests oligomer configurations [36] | Contains native complexity; hard to isolate variables | Tests simple dimerization | Confirms dimerization capability without physiological context |
| Scalability & Throughput | Modular design allows for high-throughput testing of multiple configurations | Low; relies on native assembly | Moderate; each scaffold requires separate design | High for simple confirmation, low for detailed study |
| Key Experimental Finding | Proximity-driven dimerization induces activity with multivalent enhancement in 3-4 enzyme oligomers [36] | Mediates formation of both homo- and heterodimers with distinct activities [23] | Confirms capacity for proximity-induced activation | Activates caspase-9 but yields less robust activity than apoptosome [23] |
Experimental Protocol: Recapitulating Proximity-Induced Activation
The fundamental workflow for employing DNA origami to study caspase-9 activation involves key steps in platform design, protein preparation, and functional assessment.
DNA Origami Nanostructure Design and Assembly
- Scaffold Preparation: A twist-corrected rectangular DNA origami nanostructure (∼75 × 100 nm) is self-assembled from a long single-stranded DNA scaffold (typically M13mp18) and hundreds of complementary staple strands [28].
- Handle Functionalization: Selected staple strands are extended to include unique 15-nucleotide single-stranded DNA "handles" that protrude from the origami surface. These serve as attachment points and allow for programmable positioning of proteins with ∼6 nm resolution across the entire platform [28].
- Purification and Validation: Assembled nanostructures are purified via agarose gel electrophoresis or PEG precipitation. Structural integrity and homogeneity are confirmed using atomic force microscopy (AFM) and transmission electron microscopy (TEM) [28].
Caspase-9 Functionalization and Immobilization
- Site-Specific Conjugation: The non-canonical amino acid p-azidophenylalanine is incorporated at the N-terminus of the caspase-9 catalytic domain using amber codon suppression. This enables site-specific conjugation to a bicyclononyne-functionalized oligonucleotide (BCN-DNA) via strain-promoted azide-alkyne cycloaddition (SPAAC) [28].
- Oligonucleotide Design: The conjugate oligonucleotide includes a 10-nt single-stranded linker separating the enzyme from a 15-nt "anti-handle" sequence complementary to the handles on the DNA origami [28].
- Quality Control: Successful conjugation and removal of unreacted components are confirmed through polyacrylamide gel electrophoresis (PAGE) and subsequent purification [28].
Activity Assay and Kinetic Analysis
- Proteolytic Assay: Enzymatic activity is measured by monitoring the proteolytic cleavage of the synthetic caspase-9 substrate LEHD-AFC (Leu-Glu-His-Asp-7-amino-4-trifluoromethylcoumarin) [28].
- Kinetic Measurements: Fluorescence emission (AFC release) is tracked over time. Michaelis-Menten kinetics are determined by varying substrate concentrations, allowing for the calculation of ( Km ) and ( V{max} ) values [28].
- Comparative Analysis: Activity is compared across different enzyme configurations (monomers, dimers, oligomers) on the origami platform and against traditional activation methods.
Table 2: Key experimental findings from DNA origami-based caspase-9 studies
| Experimental Manipulation | Measured Outcome | Key Quantitative Result | Interpretation |
|---|---|---|---|
| Dimerization on DNA Template | Enzymatic activity (LEHD-AFC cleavage) | Sharp activity increase only with template present [28] | Validation that DNA hybridization can induce proximity-driven activation |
| Michaelis Constant (KM) Determination | KM of the ternary complex | 1.1 ± 0.1 mM [28] | Similar to native apoptosome activity, confirming physiological relevance |
| Inter-Enzyme Distance Variation | Activity at different spacings | Activity is highly sensitive to sub-10nm distances [28] | Demonstrates critical role of precise spatial organization in activation |
| Wildtype/Mutant Heterodimer | Enzymatic activity of heterodimer vs. homodimer | Similar activity to wildtype homodimer [28] | Supports asymmetric activation mechanism with half-of-sites reactivity |
| Three- and Four-Enzyme Oligomers | Activity enhancement factor | Multivalent activity enhancement beyond simple dimer [36] | Reveals higher-order clustering effects not accessible with traditional scaffolds |
The Scientist's Toolkit: Essential Research Reagents
The following table catalogues the essential materials and reagents required to implement the DNA origami-based synthetic apoptosome system.
Table 3: Key research reagent solutions for constructing a synthetic apoptosome
| Reagent / Material | Function / Application | Specific Example / Notes |
|---|---|---|
| Caspase-9 Catalytic Domain | The signaling protein under study; requires functionalization | N-terminal azide modification for bioorthogonal conjugation [28] |
| DNA Scaffold (e.g., M13mp18) | Structural backbone of the origami | ∼7000-8000 nucleotides; provides the framework for the nanostructure [28] |
| Staple Strands with Handles | Define origami shape and protein attachment points | 15-nt single-stranded handles provide hybridization sites [28] |
| BCN-DNA Conjugate | Linker for protein-DNA conjugation | Bicyclononyne-functionalized oligonucleotide for SPAAC with protein azide [28] |
| LEHD-AFC | Fluorogenic caspase-9 substrate | Cleavage releases fluorescent AFC; enables kinetic measurements [28] |
| Ammonium Citrate | Kosmotropic salt for enforced dimerization (control experiment) | Traditional method to induce caspase-9 dimerization independent of scaffolds [23] |
Signaling Pathways and Experimental Workflows
The diagrams below illustrate the core biological concept and the corresponding experimental approach using DNA origami.
Native Apoptosis Pathway. The intrinsic apoptosis pathway, culminating in caspase-9 activation via the apoptosome.
Synthetic Apoptosome Construction. The experimental workflow for creating and testing the DNA origami-based synthetic apoptosome.
Discussion and Future Perspectives
The experimental data derived from the DNA origami platform provides compelling evidence for the induced proximity model of caspase-9 activation. The platform's capacity to demonstrate that enzymatic activity is induced by proximity-driven dimerization, and to further reveal a multivalent activity enhancement in oligomers of three and four enzymes, offers a resolution to the longstanding debate [36]. These findings were enabled by the platform's unique capability to systematically control variables that are intrinsically linked in the native apoptosome.
Looking forward, the programmability of DNA origami suggests broader applications beyond caspase-9 studies. This bottom-up approach serves as a blueprint for engineering synthetic signaling complexes for other multi-enzyme systems involved in inflammation, innate immunity, and programmed cell death [36]. Recent advances demonstrate the potential for even greater complexity, such as modular DNA origami nanocompartments that couple protein unfolding with proteolytic degradation, creating functional analogs of natural modular enzymes like the proteasome [37]. As the technology for rigid, multipoint attachment of proteins to DNA scaffolds improves [38], so too will the precision and resolution of the mechanistic insights we can derive. This positions DNA origami not merely as a tool for recapitulating biology, but as a platform for creating entirely new functional architectures with engineered properties.
The apoptosome, a multi-subunit caspase-activating complex, functions as the central proteolysis machine in the intrinsic apoptosis pathway. Its core component, Apaf-1, oligomerizes into a wheel-like structure upon binding cytochrome c and dATP/ATP, creating a platform for procaspase-9 activation. This activation mechanism has been the subject of intense debate, primarily between the induced proximity model (proximity-driven homodimerization) and the induced conformation model (allosteric activation upon binding). This guide objectively compares the experimental data and kinetic parameters derived from key in vitro reconstitution studies that have shaped this debate, providing researchers with a consolidated resource for understanding apoptosome function.
The apoptosome is a multi-protein complex that activates initiator caspases in response to intrinsic cell death signals. In mammals, the core components include Apaf-1, cytochrome c, and caspase-9 [13] [39]. Upon mitochondrial outer membrane permeabilization, cytochrome c is released into the cytosol where it binds to monomeric, auto-inhibited Apaf-1. In the presence of dATP/ATP, this binding triggers extensive conformational changes, leading to Apaf-1 oligomerization into a heptameric platform that recruits and activates procaspase-9 [40] [39].
The precise molecular mechanism by which the apoptosome activates procaspase-9 remains a fundamental question in apoptosis research. Two primary models have emerged:
- Induced Proximity/Dimerization Model: This model posits that the apoptosome serves as a scaffold to increase the local concentration of procaspase-9 monomers, facilitating their homodimerization and subsequent autoactivation [3] [23].
- Induced Conformation/Allosteric Model: This model suggests that binding to the apoptosome induces allosteric conformational changes in procaspase-9 that directly activate the protease, independent of homodimerization [41] [42].
Recent evidence suggests these mechanisms are not mutually exclusive, and a hybrid model may more accurately represent the activation process [23] [39].
Comparative Analysis of Apoptosome Activity and Kinetics
Table 1: Comparative kinetic parameters of apoptosome activity under different experimental conditions
| Experimental Condition | Caspase-9 Activity | Caspase-3 Activation | Key Findings | Study |
|---|---|---|---|---|
| Native apoptosomes (immunopurified) | Robust LEHDase activity | Efficient DEVDase activity | Contains Apaf-1, caspase-9, caspase-3, XIAP; cytochrome c not stably associated | [43] |
| Mathematical simulation (dimerization model) | Delayed, inefficient | Failed to replicate experimental data | Poor fit to XIAP threshold and molecular timer function | [41] [42] |
| Mathematical simulation (allosteric model) | Rapid, efficient | Accurately reproduced experimental data | Matched XIAP threshold, procaspase-9 processing half-times | [41] [42] |
| Engineered caspase-9 dimer | Moderately enhanced | Reduced efficiency | Activity not stimulated by Apaf-1; less robust than apoptosome-activated wild-type | [3] |
| Procaspase-9-TM (non-cleavable) | Higher V~max~ | More robust activation | Higher affinity for apoptosome than processed caspase-9-p35/p12 | [23] |
| Caspase-9-p35/p12 (processed) | Lower V~max~ | Less robust activation | Lower affinity for apoptosome; initiates molecular timer | [23] |
| PHAPI addition | Enhanced | Direct caspase-3 enhancement | Acts as co-factor for caspase-3; enhances apoptosome activity | [43] |
| Smac/DIABLO addition | Enhanced (IAP antagonism) | Indirect enhancement | Neutralizes XIAP inhibition | [43] |
Table 2: Key molecular interactions in the apoptosome
| Complex Component | Binding Partner | Interaction Domain | Functional Consequence | Structural Data Source |
|---|---|---|---|---|
| Apaf-1 | Cytochrome c | WD40 β-propeller | Relieves autoinhibition; triggers oligomerization | Cryo-EM [39] |
| Apaf-1 CARD | Procaspase-9 CARD | CARD-CARD disk | Recruits procaspase-9 to apoptosome | Cryo-EM [40] [39] |
| Procaspase-9 | Procaspase-9 (homodimer) | Small subunit (GCFNF motif) | Enhances avidity for apoptosome; selective intramolecular cleavage | Site-specific crosslinking [23] |
| Procaspase-9 | Apaf-1 NOD domain | Small subunit | Forms heterodimer; efficient caspase-3 activation | Biochemical assays [23] |
| Caspase-9-p35/p12 | Apaf-1 | Small subunit (weakened) | Reduced affinity; initiates molecular timer | SEC-MALS [23] |
| XIAP | Caspase-9 | BIR3 domain | Inhibits caspase-9 activity; recruited to native apoptosomes | Co-immunoprecipitation [43] |
Key Experimental Protocols in Apoptosome Research
One-Step Immunopurification of Native Apoptosomes
This protocol enables the isolation of catalytically active apoptosomes from cell extracts for compositional and functional analysis [43].
- Cell Extract Preparation: Prepare cytosolic extracts from Jurkat T-cells in low-salt buffer to maintain apoptosome stability.
- Apoptosome Assembly: Initiate assembly by adding cytochrome c (10 µM) and dATP (1 mM) to cell extracts. Incubate at 30°C for 10-30 minutes.
- Immunoprecipitation: Use monoclonal anti-caspase-9 antibody for differential co-immunoprecipitation in the presence versus absence of cytochrome c/dATP.
- Washing and Elution: Wash immunocomplexes extensively with buffer containing 150mM NaCl to remove unbound proteins. Elute complexes with low-pH buffer or competitive peptides.
- Activity Assay: Assess catalytic activity using fluorogenic tetrapeptide substrates (LEHD-afc for caspase-9, DEVD-afc for caspase-3). Measure hydrolysis kinetics spectrofluorometrically.
- Composition Analysis: Identify complex constituents by analytical and preparative 2D-PAGE coupled with MALDI mass spectrometry.
In Vitro Reconstitution with Recombinant Components
This approach allows controlled assembly of apoptosomes from purified proteins for mechanistic studies [23].
- Protein Purification: Express and purify recombinant Apaf-1, procaspase-9, and caspase-3 from E. coli or insect cells.
- Apoptosome Assembly: Mix Apaf-1 (50-100 nM) with cytochrome c (10 µM) and dATP (1 mM) in assembly buffer. Incubate at 30°C for 30-60 minutes.
- Caspase Recruitment: Add procaspase-9 (50-100 nM) to pre-assembled apoptosomes. Incubate for additional 15-30 minutes.
- Activity Measurement:
- Peptide Cleavage: Monitor LEHD-afc or DEVD-afc hydrolysis.
- Procaspase-3 Processing: Use catalytically inactive procaspase-3 (C163A) as substrate. Analyze cleavage by immunoblotting.
- Crosslinking Studies: Incorporate site-specific crosslinkers to stabilize and detect caspase-9 homo- and heterodimers within the complex.
Systems Biology Modeling of Apoptosome Formation
Mathematical simulations test different activation mechanisms against experimental data [41] [42].
- Model Implementation: Implement ordinary differential equations representing molecular interactions during apoptosome formation.
- Parameterization: Use published kinetic constants (K~d~ for procaspase-9 binding to Apaf-1 = 0.7 µM) and cellular concentrations (HeLa cells).
- Scenario Testing:
- Dimerization Scenario: Implement procaspase-9 homodimerization as prerequisite for activation.
- Allosteric Scenario: Implement procaspase-9 activation upon binding to apoptosome backbone.
- Validation Metrics: Compare simulation results to experimental data including XIAP threshold concentrations, procaspase-9 processing half-times, and molecular timer function.
Mechanistic Insights and Signaling Pathways
Caspase-9 Activation Pathway in Apoptosome Assembly
The diagram illustrates the sequential process of apoptosome-mediated caspase activation, highlighting key regulatory nodes. Mitochondrial stress triggers cytochrome c release, which binds to monomeric Apaf-1 and promotes nucleotide exchange (ADP to ATP/dATP). This induces conformational changes that enable Apaf-1 oligomerization into the heptameric apoptosome platform [39]. Procaspase-9 is then recruited through CARD-CARD interactions with Apaf-1. The activation of caspase-9 occurs through a combination of induced conformation and dimerization mechanisms [23]. Active caspase-9 then processes executioner caspase-3, leading to apoptosis. Regulatory proteins including XIAP (inhibitor), Smac/DIABLO (XIAP antagonist), and PHAPI (caspase-3 co-factor) fine-tune this process [43].
Mechanistic Models of Caspase-9 Activation
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key reagents for apoptosome reconstitution and activity studies
| Reagent/Solution | Function in Experiment | Example Application | Key Characteristics |
|---|---|---|---|
| Recombinant Apaf-1 | Core apoptosome scaffold | In vitro reconstitution studies | Full-length (Apaf-1XL) with 15 WD40 repeats; autoinhibited in ADP-bound state [39] |
| Cytochrome c (equine/heart) | Apoptosome assembly trigger | Initiate Apaf-1 oligomerization | Binds WD40 domain of Apaf-1; not stably associated in active complex [43] |
| dATP/ATP | Cofactor for assembly | Nucleotide exchange and oligomerization | dATP/ATP binding induces conformational change; hydrolysis not required [41] |
| Caspase-9 Antibodies | Immunopurification | Isolation of native complexes | Monoclonal antibodies for differential co-IP; epitope accessibility changes upon oligomerization [43] |
| Fluorogenic Substrates | Activity measurement | Kinetic assays (LEHD-afc, DEVD-afc) | Tetrapeptide sequences with amc or afc reporters; measure hydrolysis spectrofluorometrically [23] |
| Site-specific Crosslinkers | Stabilize protein interactions | Detect caspase-9 dimers in complex | Chemical crosslinkers to stabilize transient homo- and heterodimers for biochemical analysis [23] |
| XIAP, Smac/DIABLO | Regulation studies | Modulation of apoptosome activity | XIAP inhibits caspase-9; Smac/DIABLO neutralizes XIAP [43] |
| PHAPI | Activity enhancement | Caspase co-factor studies | Tumor suppressor that enhances caspase-3 activity and apoptosome function [43] |
The in vitro reconstitution of the apoptosome has been instrumental in deciphering the complex mechanisms governing caspase-9 activation. Quantitative data from biochemical, structural, and systems biology approaches increasingly support a hybrid model that incorporates elements of both induced proximity and induced conformation mechanisms [23] [39]. The emerging consensus suggests that caspase-9 can form both homodimers and heterodimers with Apaf-1 within the apoptosome, with each configuration potentially contributing distinct functional properties [23]. The regulatory landscape, fine-tuned by proteins such as XIAP, Smac/DIABLO, and PHAPI, adds further complexity to this activation machinery [43]. Future research using increasingly sophisticated reconstitution approaches will continue to refine our understanding of this essential cell death complex, with significant implications for therapeutic targeting in cancer and other diseases characterized by apoptotic dysregulation.
Caspase-9 is a fundamental initiator caspase in the intrinsic apoptotic pathway, responsible for executing programmed cell death in response to various cellular stresses, including DNA damage and developmental cues [1] [2]. Its activation occurs within a multi-protein complex called the apoptosome, where it dimerizes and subsequently activates downstream effector caspases, such as caspase-3 and -7, leading to controlled cellular disassembly [1] [2]. Given its pivotal role in determining cell fate, caspase-9 is crucial for normal embryonic development, particularly in the central nervous system, and for suppressing tumorigenesis by eliminating potentially cancerous cells [1] [44]. The study of Caspase-9 knockout (KO) mouse models has been instrumental in unraveling its physiological functions, the consequences of its disruption, and the molecular intricacies of its activation mechanism, which is a subject of debate between the induced proximity and induced conformation models [44] [4].
Phenotypic Analysis of Caspase-9 Knockout Models
Core Developmental and Cellular Phenotypes
Genetic ablation of Caspase-9 in mice results in severe and often lethal developmental defects, unequivocally establishing its non-redundant role in apoptosis during embryogenesis.
Table 1: Primary Phenotypes of Caspase-9 Knockout Mice
| Phenotype Category | Specific Manifestations | Underlying Cause/Mechanism | Citations |
|---|---|---|---|
| Viability | Perinatal lethality | Failure to execute essential developmental apoptosis | [44] |
| Brain Development | Malformations of the cerebrum; supernumerary neurons; ectopic cell masses; blockage of brain aqueducts | Profound reduction in apoptotic cell death in the central nervous system during development | [1] [44] |
| Cellular Apoptosis | Resistance to apoptotic stimuli like UV irradiation, γ-irradiation, and dexamethasone | Disruption of the intrinsic apoptotic pathway; inability to activate effector caspases | [1] |
| Cancer Susceptibility | Associated with increased risk (inhibition prevents elimination of damaged cells) | Failure to activate cell death in response to internal damage signals; can be a tumor escape mechanism | [1] |
The most striking abnormality in Caspase-9 KO mice is the profound disruption of brain development. These mice exhibit a marked increase in neuronal cellularity, leading to a grossly malformed cerebrum and ectopic cell populations that can obstruct vital structures like the brain aqueduct [44]. This phenotype stems directly from the near-complete absence of apoptosis in the developing nervous system, demonstrating that caspase-9 is the principal initiator caspase for developmental cell death in the brain [1] [44]. At the cellular level, fibroblasts and embryonic stem cells derived from these null mice are highly resistant to apoptosis induced by various intrinsic pathway stimuli, such as UV radiation and chemotherapeutic drugs, confirming the central role of caspase-9 in stress-induced cell death [1].
Implications for Disease and Therapy
The resistance to apoptosis observed in Caspase-9 null cells has significant implications for human diseases, particularly cancer. Many cancer cells exhibit defective caspase-9 activation or expression, which serves as a mechanism to evade cell death and increase resistance to chemotherapy [1]. For instance, reduced caspase-9 activity and low levels of its activator, Apaf-1, have been linked to acquired resistance to cisplatin in head and neck squamous cell carcinoma [1]. Furthermore, specific polymorphisms in the human CASPASE-9 gene are associated with altered susceptibility to various cancers, including lung, bladder, and gastric cancers [1]. Beyond oncology, caspase-9 has been implicated in degenerative diseases. Activated caspase-9 is detected in the end stages of Huntington's disease, suggesting a contribution to neuronal loss, and certain caspase-9 polymorphisms are associated with a higher risk of discogenic low back pain [1].
The Central Debate: Induced Proximity vs. Induced Conformation
A fundamental question in cell biology concerns the precise mechanism by which initiator caspases like caspase-9 are activated on their respective signaling platforms. The investigation of Caspase-9 KO models and complementary in vitro studies has been central to a key theoretical debate between two main models.
The Induced Proximity Model
The original induced proximity model proposed that the apoptosome primarily serves as a platform to concentrate caspase-9 zymogens, bringing them into close proximity. This increased local concentration was thought to drive homodimerization, which in turn facilitates autocatalytic activation and the subsequent cleavage and activation of downstream effector caspases [4] [12]. In this model, dimerization via the intrinsic dimerization interface of caspase-9 is the central and sufficient step for its activation.
The Induced Conformation Model
Contrasting this is the induced conformation model, which posits that the apoptosome does more than just cluster caspase-9 monomers. Through specific, multimeric interactions between the CARD domains of Apaf-1 and caspase-9, the apoptosome induces a critical conformational change in caspase-9 that is essential for its full catalytic activity [3] [1] [4]. This model suggests that dimerization alone is qualitatively different and insufficient for full activation.
Key Experimental Evidence
A critical experiment engineered a constitutively dimeric form of caspase-9 to test the induced proximity model [4]. While this dimeric caspase-9 showed higher activity than the wild-type monomer in vitro and induced more cell death when overexpressed, its activity was only a small fraction of that achieved by Apaf-1-activated wild-type caspase-9 [4]. Furthermore, the activity of this engineered dimer could not be significantly enhanced by Apaf-1, unlike the wild-type protein [4]. The crystal structure of the dimeric caspase-9 closely resembled the wild-type protein, ruling out major structural alterations as the cause of its low activity [4]. These findings strongly suggest that the apoptosome activates caspase-9 through a mechanism that goes beyond simple dimerization, supporting a model where an induced conformational change is required for full enzymatic activity [3] [4]. The two models are not necessarily mutually exclusive, and a refined understanding may incorporate elements of both proximity-driven dimerization and activation through specific conformational adjustment [3].
Diagram 1: Contrasting Models of Caspase-9 Activation. The induced proximity model emphasizes dimerization driven by local concentration, while the induced conformation model highlights an essential conformational change induced by specific interactions with the apoptosome. Key experimental evidence from an engineered dimer supports the latter model.
Detailed Experimental Workflow for Caspase-9 Phenotype Analysis
The definitive characterization of Caspase-9 knockout phenotypes relies on a multi-faceted experimental approach, combining genetic engineering, histological analysis, and molecular biochemistry.
Table 2: Key Methodologies in Caspase-9 Knockout Research
| Method | Key Application | Specific Protocols & Reagents | Outcome Measured |
|---|---|---|---|
| Gene Targeting | Generation of global Caspase-9 null mice. | Homologous recombination in embryonic stem cells to disrupt the Casp9 gene. | Germline transmission of mutated allele; confirmation by Southern blotting/PCR. |
| Histological Analysis | Assessment of developmental brain defects. | Tissue fixation, paraffin embedding, sectioning; Toluidine blue or H&E staining to identify pyknotic cells (apoptotic). | Brain structure malformations; presence/absence of apoptotic clusters in embryonic CNS. |
| Cell Death Assays | Quantifying apoptosis resistance in derived cells. | Treatment with UV, γ-irradiation, or dexamethasone; analysis by TUNEL assay, Annexin V staining, or caspase activity assays. | Percentage of cell death; caspase-3/-7 activation; phosphatidylserine externalization. |
| Biochemical Analysis | Studying caspase-9 activation mechanisms. | In vitro reconstitution with purified Apaf-1, cytochrome c, caspase-9; use of engineered dimeric caspase-9 [4]. | Catalytic activity of caspase-9 (using fluorogenic substrates like LEHD-AFC); cleavage of downstream caspases. |
Diagram 2: A Simplified Workflow for Characterizing Caspase-9 KO Mice. The approach integrates observations from whole-organism viability down to molecular mechanisms to build a conclusive picture of gene function.
Essential Research Reagents and Tools
Advancing the understanding of caspase-9 biology and its translational application depends on a specific toolkit of reagents and model systems.
Table 3: The Scientist's Toolkit for Caspase-9 Research
| Reagent / Model | Function/Application | Key Characteristics & Examples |
|---|---|---|
| Caspase-9 KO Mice | In vivo analysis of physiological function and developmental phenotypes. | Global KO is perinatal lethal with brain defects; conditional KO mice allow tissue-specific study. |
| Pharmacological Inhibitors | Acute inhibition of caspase-9 activity in cells or animals. | e.g., Z-LEHD-FMK (cell-permeable, irreversible inhibitor used in vivo at 10 mg/kg) [17]. |
| siRNA/shRNA | Transient or stable knockdown of caspase-9 expression in cell lines. | Used in in vitro models (e.g., MLE-12 alveolar cells) to study function in fibrosis [17]. |
| Engineered Dimeric Caspase-9 | Mechanistic studies to dissect activation models. | A constitutively dimeric mutant used to test induced proximity vs. induced conformation [4]. |
| Activity Assays | Quantifying caspase-9 enzymatic activity. | Fluorogenic substrates containing the LEHD sequence (e.g., LEHD-AFC); cleavage releases a fluorescent signal. |
| Specific Antibodies | Detecting expression, cleavage, and localization. | Antibodies against full-length caspase-9 and cleaved-caspase-9 (e.g., at Asp315) for IHC/Western blot [17]. |
Caspase-9 knockout models have provided indispensable insights, firmly establishing this protease as a non-redundant initiator of the intrinsic apoptotic pathway during development and in disease. The phenotypic analysis of these models highlights the severe consequences of its loss, particularly for the central nervous system. Furthermore, the study of caspase-9 has driven a critical theoretical debate, with experimental evidence increasingly favoring an induced conformation model of activation, where the apoptosome acts as a specialized allosteric regulator rather than a mere clustering platform. Future research will likely focus on further elucidating the fine details of this conformational switch, exploring the non-apoptotic functions of caspase-9 in processes like differentiation, and leveraging this knowledge for therapeutic ends. Targeting the caspase-9/β-catenin axis, for instance, is emerging as a promising strategy against pulmonary fibrosis [17], showcasing the translational potential of fundamental research into this critical cellular regulator.
Caspase-9 serves as a critical initiator caspase that regulates the intrinsic apoptotic pathway, playing an essential role in both physiological cell death and pathological tissue degeneration [8]. Unlike effector caspases that are activated through proteolytic cleavage, initiator caspases like caspase-9 undergo activation through complex molecular mechanisms that remain actively debated in scientific literature [3]. The induced proximity model posits that caspase-9 activation occurs primarily through dimerization when brought into close contact within the apoptosome complex [12]. In contrast, the induced conformation model suggests that binding to Apaf-1 in the apoptosome induces structural changes that drive activation beyond mere dimerization [3]. Research by Shi and colleagues demonstrated that engineered, dimeric caspase-9 exhibited only a fraction of the activity of Apaf-1-activated wild-type caspase-9, suggesting that dimerization alone may not be the major activation mechanism [3]. These competing models provide the theoretical framework for understanding caspase-9 substrate specificity and cleavage activity toward sequences such as LEHD, a recognized caspase-9 cleavage motif.
The significance of caspase-9 extends beyond its apoptotic functions to include roles in cellular differentiation, mitochondrial homeostasis, autophagy, and innate immunity [8]. This multifunctional nature, combined with its association with various human diseases including cancer, neurodegenerative disorders, and cardiovascular conditions, makes accurate measurement of its activity particularly valuable for both basic research and drug discovery efforts [8]. This guide provides an objective comparison of methodologies for assessing caspase-9 activity, with particular focus on cleavage assays employing the LEHD recognition sequence.
Caspase-9 Specificity and Biological Function
Substrate Recognition and the LEHD Motif
Caspase-9 is a cysteine-aspartic protease that cleaves target proteins following specific aspartic acid residues, with preference determined by adjacent amino acids that create unique recognition motifs [8]. The LEHD sequence represents one such motif optimized for caspase-9 recognition and cleavage. This tetrapeptide motif corresponds to the P4-P1 positions in substrate proteins, with cleavage occurring after the aspartic acid (D) residue [8].
The biological relevance of caspase-9 extends beyond its role in apoptosis initiation through activation of effector caspases-3, -6, and -7 [8]. Caspase-9-mediated cleavage of caspase-6 contributes to axonal degeneration in ischemic stroke, while its cleavage of vimentin facilitates dismantling of intermediate filaments during apoptosis [8]. Additional physiological substrates include RNF2, MVP, and Sema7a, each mediating distinct cellular processes from polycomb protein function to axonal projection during olfactory development [8].
Caspase-9 in Apoptotic and Non-Apoptotic Pathways
Table 1: Caspase-9 Functions in Cellular Processes
| Function Type | Specific Role | Key Substrates/Effectors | Biological Outcome |
|---|---|---|---|
| Apoptotic | Initiator caspase | Caspases-3, -6, -7 | Execution of apoptosis |
| Intermediate filament disruption | Vimentin | Cytoskeletal dismantling | |
| Polycomb regulation | RNF2 | Epigenetic alterations | |
| Non-Apoptotic | Neuronal development | Sema7a | Olfactory axon projection |
| Mitochondrial homeostasis | Unknown | ROS regulation, autophagy | |
| Endosomal sorting | IGFR2 | Retrograde transport (non-catalytic) | |
| Immune response | Unknown | Viral infection defense |
The diversity of caspase-9 functions necessitates precise measurement tools that can distinguish between its various activities in different cellular contexts. This is particularly important given that caspase-9 can drive neurovascular injury through non-apoptotic endothelial cell dysfunction, demonstrating that its activation does not always lead to immediate cell death [8].
Comparative Analysis of Caspase Activity Assays
Direct Biochemical Assays
Biochemical assays utilizing recombinant caspases and synthetic substrates represent the most direct approach for measuring enzymatic activity and kinetics. These typically employ fluorogenic or chromogenic substrates containing the LEHD motif to quantify caspase-9 activity in purified systems [45].
In practice, these assays are conducted in multi-well plates with reaction volumes optimized for high-throughput screening (e.g., 12µL in 384-well plates) [45]. The standard protocol involves pre-incubating enzyme with potential inhibitors for 15 minutes, followed by substrate addition and 40-minute reaction at room temperature before measurement [45]. Detection employs excitation/emission wavelengths of 485/535nm for rhodamine-based substrates like (Ac-LEHD)₂-Rh110 [45].
Table 2: Comparison of Caspase Activity Measurement Platforms
| Method Type | Detection Principle | Key Reagents | Advantages | Limitations |
|---|---|---|---|---|
| Biochemical (Direct) | Fluorogenic substrate cleavage | Recombinant caspase, (Ac-LEHD)₂-Rh110 | High specificity, suitable for inhibitor screening | Non-physiological context |
| Whole-Cell ELISA | Endogenous substrate cleavage | Antibodies against cleaved lamin A/C | Physiological relevance, caspase-6 specific | Indirect caspase-9 measurement |
| HTPS Profiling | Mass spectrometry | Native lysates, 96FASP filters | Unbiased substrate discovery | Technically demanding |
| LDH Release | Membrane integrity | LDH antibody, colorimetric substrate | Measures cell death endpoint | Indirect, not caspase-specific |
Whole-Cell Caspase Activity Assays
While biochemical assays provide controlled conditions, whole-cell assays preserve physiological context that significantly influences caspase activation and substrate specificity. A whole-cell chemiluminescent ELISA approach has been developed for caspase-6 that monitors proteolytic degradation of endogenously expressed lamin A/C during early apoptosis [45]. This methodology could be adapted for caspase-9 by targeting specific endogenous substrates.
The key advantage of whole-cell assays is maintenance of cellular compartmentalization and native protein complexes that regulate caspase activity. In the caspase-6 assay, plasma membrane integrity remained intact despite apoptosis induction, as assessed by lactate dehydrogenase (LDH) retention and exclusion of cell-impermeable peptide inhibitors [45]. This preservation of cellular architecture provides more physiologically relevant activity data, though with potentially reduced specificity compared to biochemical assays.
High-Throughput Protease Screening (HTPS)
A recently developed High-Throughput Protease Screen (HTPS) method enables comprehensive profiling of protease substrates and cleavage specificity under near-native conditions [46]. This approach uses native cell lysates with endogenous proteases blocked by inhibitors, followed by proteolysis with the protease of interest in 96FASP filter plates with 10 kDa molecular weight cut-off [46].
The HTPS workflow involves:
- Preparation of native lysate with inhibitor cocktail
- Protease digestion at 1:50 enzyme-to-substrate ratio
- Peptide separation using 96FASP filter plates
- Direct analysis by data-dependent acquisition mass spectrometry
- Computational identification of cleavage products using unspecific database search parameters [46]
This method preserves native substrate folding and disulfide bridges, which significantly impact substrate accessibility and cleavage efficiency. For caspase-9, this could provide unbiased identification of physiological substrates beyond the LEHD motif, revealing potential novel recognition patterns influenced by tertiary structure rather than linear sequences alone [46].
Experimental Protocols for Caspase Activity Measurement
Recombinant Caspase-9 Biochemical Assay
Materials:
- Recombinant caspase-9 (expressed in E. coli and purified)
- Fluorogenic substrate (Ac-LEHD)₂-Rh110
- Assay buffer: 50 mM HEPES pH 7.2, 25 mM MgSO₄, 0.5 mM EGTA, 5 mM glutathione, 0.01% Triton X-100 with 0.1% BGG
- Low-volume 384-well assay plates
- Plate reader capable of fluorescence detection (excitation/emission: 485/535nm)
Procedure:
- Prepare inhibitor dilutions in DMSO (final concentration ≤1.33%)
- Pre-incubate caspase-9 with inhibitors in assay buffer for 15 minutes
- Add substrate to initiate reaction (final volume: 12µL)
- Incubate for 40 minutes at room temperature
- Measure fluorescence intensity
- Calculate IC₅₀ values for inhibitors from dose-response curves [45]
Whole-Cell Caspase Activity Assessment
Materials:
- Appropriate cell line (e.g., SKNAS cells for caspase-6)
- Apoptosis inducer (e.g., staurosporine)
- Lysis buffer
- Primary antibodies against caspase-9 neoepitopes (D315 or D330)
- Secondary antibodies (HRP-conjugated or fluorescently labeled)
- Chemiluminescent or fluorescent detection reagents
Procedure:
- Culture cells under standard conditions (37°C, 5% CO₂, 90% humidity)
- Induce apoptosis with appropriate stimulus
- Harvest cells at specific time points
- Lyse cells and quantify protein content
- Perform SDS-PAGE and transfer to nitrocellulose
- Block membrane and incubate with primary antibodies (1:1000 dilution)
- Incubate with secondary antibodies (1:2500-1:10000 dilution)
- Detect using appropriate scanning instrument [45]
High-Throughput Protease Specificity Profiling
Materials:
- Native cell lysate
- Low molecular weight protease inhibitors
- 96FASP filter plates (10 kDa MWCO)
- Recombinant caspase-9
- Mass spectrometry-compatible buffers
Procedure:
- Prepare native lysate, inhibiting endogenous proteases
- Remove inhibitors and background proteolysis products using 10 kDa filters
- Aliquot 50µg lysate per well in 96FASP plate
- Add caspase-9 at 1:50 enzyme-to-substrate ratio
- Incubate to allow proteolysis
- Collect flow-through containing cleavage products
- Analyze by LC-MS/MS with data-dependent acquisition
- Process data using unspecific database search parameters [46]
Caspase-9 Activation Pathway and Experimental Framework
Caspase-9 Activation and Measurement Pathway
Research Reagent Solutions
Table 3: Essential Research Reagents for Caspase-9 Studies
| Reagent Category | Specific Examples | Application/Function | Detection Method |
|---|---|---|---|
| Fluorogenic Substrates | (Ac-LEHD)₂-Rh110 | Direct activity measurement | Fluorescence (485/535nm) |
| Inhibitors | XIAP Bir3 domain, dominant-negative caspase-9 mutants | Selective caspase-9 inhibition | IC₅₀ determination |
| Activity-Based Probes | LEHD-based covalent probes | Active enzyme labeling and enrichment | Fluorescence/MS |
| Antibodies | Anti-D315 cl-caspase-9, Anti-D330 cl-caspase-9 | Specific neoepitope detection | Western blot, ELISA |
| Caspase Sources | Recombinant caspase-9 (E. coli expressed) | Biochemical assays | Activity validation |
| Cell Death Inducers | Staurosporine, other apoptotic stimuli | Inducing intrinsic pathway | Model establishment |
The selection of appropriate assay methodologies for measuring caspase-9 activity against LEHD and other substrates depends heavily on the specific research questions being addressed. Biochemical assays using recombinant enzymes provide the highest specificity and are ideal for inhibitor screening and kinetic studies [45]. Whole-cell assays offer superior physiological context but may compromise direct attribution to caspase-9 specifically [45]. Emerging technologies like HTPS enable unbiased discovery of novel substrates and cleavage motifs under near-native conditions, potentially revealing new aspects of caspase-9 biology [46].
The ongoing debate between induced proximity and induced conformation models of caspase-9 activation underscores the importance of methodological selection in experimental design [3] [12]. Researchers must consider whether their assay conditions preserve the potential conformational changes that may be essential for full caspase-9 activation, particularly when screening for potential therapeutic compounds targeting this important regulator of cell death and disease.
Resolving the Mechanism: Critical Evaluation of Conflicting Evidence and Technical Challenges
The activation of caspase-9, the initiator caspase of the intrinsic apoptotic pathway, represents a critical control point in programmed cell death. For years, the induced proximity model has served as the dominant framework for understanding this process, proposing that the apoptosome primarily serves as a platform to concentrate caspase-9 molecules, driving their dimerization and subsequent activation [3] [12]. However, a puzzling contradiction emerged when researchers engineered constitutively dimeric forms of caspase-9: these dimers demonstrated only partial activity compared to their apoptosome-activated counterparts [4]. This dimerization paradox challenges fundamental assumptions about caspase activation mechanisms and suggests our understanding requires refinement. This article examines the experimental evidence behind this paradox and explores its implications for drug development targeting apoptotic pathways.
Competing Models of Caspase-9 Activation
The Induced Proximity Model
The induced proximity model posits that initiator caspase zymogens are activated when brought into close proximity by upstream activating complexes [12]. For caspase-9, this occurs through recruitment to the apoptosome, a heptameric complex of Apaf-1 and cytochrome c [3]. This model was reinterpreted to emphasize dimerization-driven activation, where the apoptosome's primary function is to facilitate dimerization of caspase-9 through its intrinsic dimerization interface [4]. Under this framework, caspase-9 exists predominantly as an inactive monomer at physiological concentrations, and apoptosome assembly creates a high local concentration that promotes dimer formation and consequent activation [32] [47].
The Induced Conformation Model
In contrast, the induced conformation model proposes that binding to the apoptosome induces specific structural changes in caspase-9 that drive its activation [1] [3]. This model suggests that the apoptosome does more than merely concentrate caspase-9 molecules—it actively reconfigures their catalytic sites into competent conformations [1]. Support for this model comes from structural studies showing that apoptosome-bound caspase-9 undergoes conformational rearrangements distinct from those achieved through dimerization alone [4].
Experimental Evidence: Direct Comparison of Caspase-9 Activity
Engineering a Constitutive Caspase-9 Dimer
To test the induced proximity model, researchers engineered a constitutively dimeric caspase-9 by modifying its dimerization interface [4]. The design rationale stemmed from comparative analysis of caspase structures, which revealed that caspase-9 contains phenylalanine residues (Phe404) at the dimer interface that create steric hindrance, unlike effector caspases like caspase-3 that dimerize readily [4]. By relieving this steric hindrance, researchers created a stable caspase-9 homodimer that forms independently of the apoptosome.
Quantitative Activity Assessment
The catalytic activity of this engineered dimer was systematically compared to wild-type caspase-9 activated by Apaf-1, with revealing results:
Table 1: Comparative Activity of Caspase-9 Forms
| Caspase-9 Form | Relative Catalytic Activity | Apaf-1 Responsiveness | Cell Death Induction |
|---|---|---|---|
| Monomeric (wild-type) | Low basal activity | Yes - significantly enhanced | Moderate when overexpressed |
| Engineered dimer | Higher than monomer but significantly lower than apoptosome-activated | No - not significantly enhanced | More efficient than wild-type monomer |
| Apoptosome-activated wild-type | Highest activity | N/A | Potent |
Despite sharing nearly identical structure with wild-type caspase-9—including the characteristic asymmetry between monomers—the engineered dimer exhibited only a fraction of the activity achieved through Apaf-1 activation [4]. Crucially, unlike wild-type caspase-9, the engineered dimer's activity could not be significantly enhanced by Apaf-1, suggesting its activation mechanism was qualitatively different [4].
Methodological Framework: Key Experimental Approaches
Protein Engineering and Structural Analysis
The engineering of dimeric caspase-9 involved rational design based on crystal structures of caspase-3 and caspase-9 [4]. Researchers replaced specific residues in the β6 strand of caspase-9's dimer interface to resemble those in caspase-3, creating a caspase-9 variant (C287S) with enhanced dimerization propensity [4]. The structural integrity of the engineered dimer was confirmed through X-ray crystallography, which revealed that its overall structure closely matched wild-type caspase-9, with root mean square deviation (RMSD) values indicating minimal structural perturbation [48] [4].
Activity Assays and Binding Studies
Catalytic activity was measured using fluorogenic substrates (Ac-LEHD-AFC), allowing quantitative comparison of enzyme kinetics across different caspase-9 forms [32]. Researchers employed gel filtration and chemical crosslinking to assess the oligomeric status of caspase-9 variants under various conditions [32]. To evaluate physiological relevance, cell death induction was measured by expressing engineered caspase-9 forms in mammalian cells and assessing apoptotic markers [4].
Table 2: Key Experimental Methods in Caspase-9 Dimerization Studies
| Method | Application | Key Findings |
|---|---|---|
| Site-directed mutagenesis | Engineering dimer interface | Creation of constitutive caspase-9 dimer through steric hindrance relief |
| X-ray crystallography | Structural characterization | Engineered dimer closely resembles wild-type structure with preserved asymmetry |
| Gel filtration chromatography | Oligomeric status assessment | Wild-type caspase-9 primarily monomeric at physiological concentrations |
| Fluorogenic substrate assays | Catalytic activity measurement | Engineered dimers show intermediate activity between monomer and apoptosome-bound forms |
| Co-immunoprecipitation | Protein interaction studies | Differential Apaf-1 binding capacity between caspase-9 forms |
Resolution of the Paradox: Beyond Simple Dimerization
The dimerization paradox finds resolution in the recognition that caspase-9 activation requires more than mere dimerization. While dimerization is necessary, it appears insufficient for full catalytic competence [4]. The current evidence suggests that the apoptosome provides allosteric regulation beyond concentration effects, inducing conformational changes that optimize the catalytic site for substrate binding and cleavage [1] [3].
Structural studies reveal that in the engineered dimers, the active sites may not achieve the same optimized configuration as in apoptosome-bound caspase-9 [4]. Specifically, the asymmetric conformation observed in caspase-9 dimers—where the two active sites display different geometries—suggests that proper orientation of catalytic residues requires specific interactions with the apoptosome platform [32]. This explains why artificial dimerization produces partially active enzymes, while apoptosome recruitment generates fully active caspase-9.
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Research Reagents for Caspase-9 Activation Studies
| Reagent | Function/Application | Experimental Utility |
|---|---|---|
| Ac-LEHD-AFC | Fluorogenic caspase-9 substrate | Quantitative measurement of caspase-9 enzymatic activity |
| Z-VAD-FMK | Pan-caspase irreversible inhibitor | Determination of active enzyme concentration through titration |
| Apaf-1-deficient cell lines | Tools for apoptosome function studies | Dissection of Apaf-1-dependent and independent activation pathways |
| Engineered caspase-9 dimer | Constitutively dimeric caspase-9 variant | Testing induced proximity model and dimerization sufficiency |
| XIAP Bir3 domain | Endogenous caspase-9 inhibitor | Studying regulation of caspase-9 activity and neoepitope specificity |
| Anti-caspase-9 neoepitope antibodies | Detection of specific caspase-9 cleavage forms | Distinguishing autocleavage (D315) from caspase-3-mediated cleavage (D330) |
The dimerization paradox in caspase-9 activation has fundamentally advanced our understanding of apoptotic initiation. The evidence clearly demonstrates that while dimerization is a necessary component of caspase-9 activation, the apoptosome provides essential allosteric regulation beyond mere concentration effects [1] [4]. This refined understanding has significant implications for drug development targeting apoptotic pathways, particularly in cancer therapeutics where overcoming apoptosis resistance is a key goal.
Future research should focus on obtaining high-resolution structures of the complete apoptosome-caspase-9 complex to visualize the precise conformational changes induced by Apaf-1 binding. Additionally, the development of small molecules that can mimic the apoptosome's allosteric effects could provide new therapeutic avenues for modulating caspase-9 activity in disease contexts. As our understanding of caspase-9 activation continues to evolve, so too will opportunities for innovative interventions in the many pathological conditions involving dysregulated apoptosis.
Caspase-9 stands as the critical initiator caspase in the intrinsic apoptotic pathway, triggering a proteolytic cascade that leads to programmed cell death. Its activation on the multiprotein platform known as the apoptosome represents a pivotal commitment step to cellular destruction. However, the precise functional role of caspase-9 autocleavage has remained a subject of intense scientific debate, primarily centered on two competing hypotheses: whether autocleavage serves as the primary activation trigger for the enzyme or functions as a molecular timer to control the duration of apoptotic signaling [1] [49]. This debate is intrinsically linked to the broader mechanistic frameworks of induced proximity versus induced conformation that explain how initiator caspases become activated [1] [49]. Resolving this dichotomy is not merely academic; it carries profound implications for understanding normal development, disease pathogenesis, and developing therapeutic strategies for cancer, neurodegenerative disorders, and other conditions where apoptotic regulation is disrupted.
Molecular Mechanisms: Two Models of Caspase-9 Activation
The Induced Proximity/Dimerization Model
The induced proximity model posits that the apoptosome primarily serves as a platform to bring procaspase-9 zymogens into close proximity. This dramatically increases the local concentration of caspase-9, facilitating homodimerization which drives its activation [1] [2]. In this model, autocleavage is an integral part of the activation process, stabilizing the active site and generating the fully mature caspase-9 (p35/p12) heterotetramer [1]. The key premise is that dimerization of caspase-9 leads to rapid autocatalytic cleavage, which is necessary for achieving full catalytic competence [1]. This perspective views autocleavage as a fundamental activation trigger in the apoptotic cascade.
The Induced Conformation Model
Contrasting with the proximity model, the induced conformation model suggests that binding to the apoptosome itself induces a specific conformational change in caspase-9 that activates its proteolytic function [1] [49]. Supporting evidence comes from engineered caspase-9 that exists as a constitutive dimer yet shows significantly lower activity than apoptosome-bound wild-type caspase-9, and its activity is not further stimulated by Apaf-1 [49]. This indicates that dimerization alone may be insufficient for full activation and that a qualitative change in structure induced by Apaf-1 binding is essential. Within this model, autocleavage may play a more regulatory, rather than activating, role.
Table 1: Core Principles of Competing Activation Models
| Feature | Induced Proximity/Dimerization Model | Induced Conformation Model |
|---|---|---|
| Core Mechanism | Apoptosome increases local caspase-9 concentration, promoting dimerization | Apaf-1 binding induces conformational change activating caspase-9 |
| Role of Autocleavage | Integral part of activation; stabilizes active site | Secondary event; may regulate activity duration |
| Primary Evidence | Dimerization correlates with increased activity [1] | Engineered dimers show low activity vs. apoptosome-bound caspase-9 [49] |
| View of Autocleavage | Activation Trigger | Molecular Timer |
Experimental Evidence: Weighing the Data
The scientific investigation into caspase-9 autocleavage has yielded evidence supporting both perspectives, requiring careful analysis of experimental data.
Evidence Supporting the Activation Trigger View
Several lines of evidence suggest autocleavage is a key activating event:
- Cleavage and Activity Correlation: Dimerization of caspase-9 leads to rapid autocatalytic cleavage, producing the active caspase-9 (p35/p12) form, and this cleavage is associated with proteolytic activity [1].
- Altered Apoptosome Affinity: Procaspase-9 demonstrates higher affinity for the apoptosome compared to its cleaved form, suggesting cleavage induces a structural change that affects complex interactions [1].
- Regulatory Phosphorylation: Phosphorylation at Thr125, a site near the N-terminus of the large subunit, inhibits caspase-9 processing and activity, indicating the importance of the cleavage region for function [1].
Evidence Supporting the Molecular Timer View
Conversely, significant findings position autocleavage as a regulatory feedback mechanism:
- Timer Function: Research indicates procaspase-9 autoprocessing is "not designed to activate caspase-9, but to turn on a molecular timer that stimulates the duration of apoptosome activity" [1]. In this view, cleavage does not initiate activity but rather controls its lifespan.
- Activity Without Cleavage: Caspase-9 demonstrates complete activity in its uncleaved form, implying the proteolytic function is gained primarily through dimerization/conformation, not cleavage itself [1].
- Differential Regulation: The cleaved and uncleaved forms of caspase-9 are differentially regulated by inhibitors like XIAP, which selectively inhibits caspase-9 cleaved at the D315 autocleavage site [8]. This allows for nuanced control of caspase-9 signaling downstream of the initial activation event.
Table 2: Key Experimental Findings on Caspase-9 Autocleavage
| Experimental Finding | Interpretation as Activation Trigger | Interpretation as Molecular Timer |
|---|---|---|
| Uncleaved caspase-9 is active | Activity is suboptimal without cleavage | Full activation is achieved prior to cleavage |
| Cleaved caspase-9 has lower apoptosome affinity | A consequence of activation and release | A mechanism to limit signaling duration |
| Autocleavage occurs after dimerization | A necessary step in the activation sequence | A feedback mechanism to time activity |
| XIAP inhibits D315-cleaved caspase-9 | Regulation of the active enzyme | A key component of the timer mechanism |
Research Methodologies for Investigating Autocleavage
Protein Engineering and Mutagenesis
A pivotal approach involves generating caspase-9 variants to test specific hypotheses.
- Engineered Constitutive Dimers: Researchers have created caspase-9 mutants that exist as stable dimers independently of the apoptosome. A critical finding was that these dimers, while more active than wild-type monomers, showed significantly lower activity than Apaf-1-activated wild-type caspase-9 and were not further stimulated by Apaf-1 [49]. This challenges the sufficiency of dimerization and supports the induced conformation model.
- Cleavage-Site Mutants: Mutating aspartic acid cleavage sites (e.g., D315) prevents autocleavage. Comparing the activity and apoptotic potential of cleavage-resistant mutants versus wild-type caspase-9 allows researchers to isolate the functional contribution of the cleavage event itself.
Structural Biology and Biophysical Analysis
- Crystallography: Determining the high-resolution crystal structures of caspase-9 alone, in a constitutive dimer, and in complex with Apaf-1 CARD domains reveals critical differences in active site conformation [1] [49] [50]. These structures can show whether binding to Apaf-1 induces a conformational change that re-arranges the catalytic machinery.
- Binding Affinity Studies: Using techniques like surface plasmon resonance (SPR) or isothermal titration calorimetry (ITC) to measure the binding kinetics of uncleaved (zymogen) versus cleaved caspase-9 to the apoptosome. The finding that procaspase-9 has higher affinity for the apoptosome than the cleaved form is a cornerstone for the molecular timer hypothesis [1].
Cellular and In Vitro Functional Assays
- Reconstituted Apoptosis Systems: Using cell-free extracts or purified components (Apaf-1, cytochrome c, caspase-9) to reconstitute apoptosome formation. This allows for precise manipulation of components (e.g., adding cleavage-resistant caspase-9) and measurement of downstream caspase-3/7 activation over time, directly testing the timer model [1].
- Cell-Based Death Assays: Transfecting cells with wild-type versus cleavage-site mutant caspase-9 and measuring apoptosis sensitivity under stress (e.g., UV, chemotherapeutic agents). If the mutant induces apoptosis as effectively as wild-type, it suggests cleavage is not the trigger.
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagents for Caspase-9 Autocleavage Research
| Reagent / Tool | Key Function / Utility | Experimental Application |
|---|---|---|
| Caspase-9 Inhibitor (Z-LEHD-FMK) | Cell-permeable inhibitor that irreversibly binds the active site of caspase-9. | Validating the specific role of caspase-9 activity in apoptotic models [17]. |
| Anti-Caspase-9 Antibodies (neo-epitope specific) | Antibodies specifically recognizing cleaved forms (e.g., at D315 or D330). | Discriminating between initiator (autocleavage) and executioner (caspase-3)-mediated cleavage in western blot, ICC [17] [8]. |
| Recombinant Engineered Caspase-9 (e.g., iCasp9) | A caspase-9 fusion protein (with FKBP) activated by a small molecule dimerizer. | A controlled system to induce caspase-9 dimerization/activation independent of the apoptosome to study downstream events [50]. |
| Apaf-1 Deficient Cell Lysates | Cell extracts incapable of forming a functional apoptosome. | Reconstitution experiments to test the necessity of the apoptosome for caspase-9 activation and autocleavage [1]. |
| Site-Directed Mutagenesis Kits | For generating cleavage-site (D315A) or phosphorylation-site (T125A) mutants. | Creating caspase-9 variants to test the functional necessity of specific post-translational modifications [1] [49]. |
Integrated Signaling and the Path Forward
The role of caspase-9 autocleavage is best understood not as strictly one function or the other, but as a potentially integrated process where both triggering and timing aspects are important. The initial activation is likely driven by a combination of proximity-induced dimerization and apoptosome-induced conformational changes. Subsequent autocleavage then fine-tunes this activity, potentially acting as a molecular timer that regulates signal duration by altering affinity for the apoptosome and creating a defined window for downstream caspase activation [1]. This model reconciles conflicting data and underscores the sophisticated multi-layer regulation of apoptosis.
Future research directions should focus on obtaining high-resolution structures of the full-length caspase-9 within the intact apoptosome, which remains a technical challenge but would provide unprecedented insight [49]. Furthermore, developing more sophisticated cellular and animal models expressing cleavage-resistant or phospho-mimetic caspase-9 mutants will be crucial for understanding the physiological relevance of these regulatory mechanisms in development, tissue homeostasis, and disease. Finally, elucidating the crosstalk between caspase-9 and other regulated cell death pathways, such as pyroptosis and necroptosis, will provide a more holistic view of its role in cellular fate decisions [2] [51].
The Role of the Linker Loop and CARD Domain in Regulation
Caspase-9 is a critical initiator caspase in the intrinsic apoptotic pathway, serving as the essential link between mitochondrial stress and cellular demolition. Its activation represents a commitment to cell death, and understanding its regulatory mechanism is paramount for developing therapies for cancer and degenerative diseases. The scientific community has long been divided between two principal models explaining caspase-9 activation: the induced proximity model, which posits that the apoptosome serves primarily as a platform to concentrate caspase-9 molecules, driving their dimerization and activation; and the induced conformation model, which argues that binding to the apoptosome actively induces a conformational change that creates a catalytically competent enzyme [1] [31]. The Caspase Activation and Recruitment Domain (CARD) and the flexible linker loop connecting it to the catalytic core are at the heart of this debate, serving as critical regulatory modules that control the transition from a latent zymogen to an active protease. This guide objectively compares the experimental evidence for how these domains regulate caspase-9, providing a structured overview of the key data and methodologies shaping current understanding.
Structural Architecture of Caspase-9
Caspase-9 is synthesized as an inactive single-chain zymogen (procaspase-9). Its domain structure consists of [1] [50] [31]:
- CARD Domain (Residues ~1-92): An N-terminal prodomain that mediates homotypic protein-protein interactions.
- Linker Loop: A flexible segment connecting the CARD to the catalytic core.
- Catalytic Core: Comprising a large (p35) and a small (p12) subunit. The active site features a unique QACGG motif, differing from the QACRG motif conserved in other caspases [50].
Unlike executioner caspases, caspase-9 is predominantly monomeric in solution and can exhibit activity even prior to proteolytic cleavage at its intersubunit linker, a property attributed to its longer linker region [15] [1]. The following diagram illustrates the domain organization and the key structural elements involved in its activation.
Diagram 1: Domain structure of procaspase-9, highlighting the CARD, linker loop, and catalytic core.
Quantitative Comparison of Caspase-9 Activation and Regulation
The following tables summarize key experimental data and regulatory mechanisms pertaining to caspase-9 activation, with a focus on the roles of the CARD domain and linker loop.
Table 1: Experimental Activity Data of Caspase-9 Constructs and Complexes
| Caspase-9 Construct / Complex | Catalytic Activity | Experimental Context | Key Finding |
|---|---|---|---|
| Full-length Caspase-9 (Monomer) | Low basal activity [15] [4] | In vitro enzymatic assay | Serves as the inactive ground state. |
| Caspase-9 ΔCARD (Core Domain) | ~20% lower than full-length [15] | In vitro enzymatic assay | CARD is required for full basal activity, suggesting a direct role in core domain regulation. |
| Apoptosome-Bound Caspase-9 | ~2,000-fold increase vs. monomer [15] | In vitro reconstitution with Apaf-1/cytochrome c | Demonstrates the profound activating power of the apoptosome. |
| Engineered Constitutive Dimer | Higher than WT monomer, but only a small fraction of apoptosome-activated activity [4] | Recombinant protein assay | Dimerization alone is insufficient for full activation; supports induced conformation. |
| CARD:Core complex | Dependent on ordered active-site loops [15] | Biophysical analysis | Direct physical interaction between CARD and core domains is a key regulatory step. |
Table 2: Key Regulatory Mechanisms Targeting the Linker and CARD Domain
| Regulatory Mechanism | Target Site | Effect on Caspase-9 | Biological Implication |
|---|---|---|---|
| Phosphorylation (e.g., by Akt/ERK) | Thr125 (near N-terminus of large subunit) [1] [50] | Inhibits activation and protease activity; affects dimerization. [50] | A major survival signal; suppresses apoptosis in cancer. |
| Alternative Splicing (Caspase-9β) | Missing exons 3-6 (part of CARD/linker) [50] | Acts as a dominant-negative inhibitor of full-length caspase-9. | Promotes cell survival; linked to chemoresistance. |
| CARD:Apaf-1 CARD Interaction | N-terminal CARD Domain [15] [1] [16] | Recruits caspase-9 to the apoptosome; essential for activation. | The primary event in intrinsic pathway activation. |
Detailed Experimental Protocols and Workflows
Probing CARD:Core Domain Interactions
This methodology, derived from [15], aims to uncover intramolecular interactions within caspase-9 independent of the apoptosome.
- 1. Protein Engineering: Generate a series of caspase-9 mutants using site-directed mutagenesis on the human caspase-9 full-length gene (amino acids 1-416) in an expression vector. Key mutants include:
- Active Site Mutant (C287A): Inactivates catalytic cysteine to decouple conformation from activity.
- Phosphomimetic/Charge Mutants (e.g., T125E, S183E): To perturb specific loop conformations and domain interactions.
- 2. Protein Expression and Purification: Transform constructs into E. coli (e.g., BL21(DE3)). Induce soluble protein expression with IPTG. Purify via immobilized metal affinity chromatography (IMAC) using a Ni-affinity column, followed by ion-exchange chromatography (e.g., Macro-Prep High Q column) to achieve high purity.
- 3. Biophysical Analysis:
- Size-Exclusion Chromatography (SEC): Analyze the hydrodynamic volume of wild-type and mutant proteins. When active-site loops are disordered, the CARD and core behave as "loosely tethered beads." A properly ordered active-site loop bundle results in a more compact structure, indicating CARD:core interaction.
- Electrospray Ionization Mass Spectrometry (ESI-MS): Confirm protein mass and composition.
- Activity Assays: Measure enzymatic activity against synthetic substrates (e.g., Ac-LEHD-AFC) to correlate conformational states with function.
Diagram 2: Experimental workflow for analyzing CARD and core domain interactions.
Testing the Induced Proximity Model via Engineered Dimerization
This pivotal experiment [4] directly tested the induced proximity hypothesis by creating a stable, constitutively dimeric caspase-9.
- 1. Rational Design:
- Compare the dimerization interface (specifically the β6 strand) of caspase-9 with that of constitutively dimeric caspase-3.
- Identify steric clashes in caspase-9 (e.g., from Phe404) that impede stable dimerization.
- 2. Protein Engineering:
- Mutate the caspase-9 β6 strand sequence to mimic the corresponding sequence in caspase-3, thereby relieving steric hindrance and creating a stabilized homodimer (e.g., caspase-9-F404A).
- 3. Structural and Functional Validation:
- Crystallography: Determine the high-resolution structure of the engineered dimer to confirm it resembles the wild-type dimer interface and maintains asymmetric active sites.
- In Vitro Activity Assay: Compare the catalytic activity of the engineered dimer against wild-type monomeric caspase-9 and apoptosome-activated caspase-9 using fluorogenic substrates.
- Cell Death Assay: Transfect mammalian cells with vectors expressing wild-type or dimeric caspase-9 and measure the efficiency of apoptosis induction (e.g., via Annexin V staining).
The Scientist's Toolkit: Key Research Reagents
Table 3: Essential Reagents for Caspase-9 Domain-Function Studies
| Reagent / Tool | Function in Research | Key Application / Finding |
|---|---|---|
| Caspase-9 Full-Length Construct (1-416) | Baseline for studying full-length, monomeric caspase-9 activity and regulation. [15] | Measuring basal activity and CARD:core interactions. |
| Caspase-9 ΔCARD Construct | Isolates the function of the catalytic core from the CARD domain. [15] | Demonstrated that CARD removal decreases catalytic efficiency. |
| Site-Directed Mutagenesis Kits | For generating active-site (C287A), phosphomimetic (T125E), and interface mutants. [15] | Probing the role of specific residues in activity, regulation, and dimerization. |
| Engineered Dimeric Caspase-9 (e.g., F404A) | A constitutive dimer to test the induced proximity model. [4] | Showed dimerization is insufficient for full apoptosome-level activity. |
| Reconstituted Apoptosome (Apaf-1, Cytochrome c, dATP) | The physiological activation platform for caspase-9. [15] [16] | The gold standard for generating fully active caspase-9 (~2000-fold activation). |
| Fluorogenic Caspase-9 Substrate (e.g., Ac-LEHD-AFC) | A sensitive reporter for measuring caspase-9 enzymatic activity. [50] [16] | Quantifying activation levels in different experimental conditions. |
Integrated Discussion: Reconciling the Models
The accumulated evidence reveals a sophisticated, multi-layered regulatory system where the CARD and linker loop integrate signals from multiple sources to control caspase-9 activity. The following diagram synthesizes this integrated regulatory mechanism.
Diagram 3: An integrated model of caspase-9 activation, incorporating conformational change and dimerization.
The data decisively shows that the old dichotomy between "induced proximity" and "induced conformation" is an oversimplification. A more accurate model integrates both concepts [15] [4] [16]:
- Ground State Regulation: In the absence of the apoptosome, the CARD domain can interact with the catalytic core, and this interaction is modulated by the conformation of the active-site loops and phosphorylation, fine-tuning the enzyme's basal activity and readiness for activation [15].
- Induced Conformation is Key: Recruitment to the apoptosome via CARD:CARD interactions is only the first step. The apoptosome actively induces a conformational change in the caspase-9 catalytic core that is essential for creating a fully competent active site. This is supported by the fact that an engineered dimer, which mimics pure induced proximity, has only a fraction of the activity of the apoptosome-bound enzyme [4].
- Dimerization within the Platform: The apoptosome also serves to concentrate caspase-9, facilitating dimerization. However, this dimerization appears to be a secondary event that further stabilizes the enzyme on the platform, rather than the primary activating trigger [15] [16].
In conclusion, the CARD domain and linker loop are not simple tethers. They are dynamic regulatory modules that mediate intramolecular interactions, respond to post-translational modifications, and ultimately transduce the activation signal from the apoptosome into the profound conformational change that unleashes caspase-9's proteolytic power. For drug development, this implies that targeting the CARD:core interface or the apoptosome-induced conformational switch may offer more specific therapeutic opportunities than targeting the conserved catalytic site itself.
Post-translational modifications (PTMs) represent a crucial regulatory layer that expands protein functionality beyond genetic encoding. Among the more than 650 known PTMs, phosphorylation is one of the most well-studied, involving the addition of a phosphate group from ATP to amino acid side chains by kinases, a process reversible by phosphatases [52]. This review examines the specific role of phosphorylation as an endogenous inhibitory mechanism, focusing on its function within the central debate surrounding caspase-9 activation—the induced proximity versus the induced conformation model [3] [12]. Caspase-9, a key initiator of the intrinsic apoptotic pathway, is activated through formation of the apoptosome complex (Apaf-1, cytochrome c). The induced proximity model posits that activation is driven by dimerization, while the induced conformation model argues for activation via conformational changes [3]. Phosphorylation serves as a critical control point in this system, with specific sites acting to directly inhibit caspase-9 activity and prevent unintended apoptosis. Understanding these mechanisms provides essential insights for drug development targeting apoptotic pathways in diseases like cancer and neurodegeneration.
Phosphorylation as an Endogenous Inhibitor – Key Evidence
The following table summarizes experimental data demonstrating phosphorylation-mediated inhibition of caspase-9 and related apoptotic components.
Table 1: Experimental Evidence for Phosphorylation-Mediated Inhibition in Apoptosis
| Protein Target | Phosphorylation Site | Kinase Responsible | Experimental System | Observed Effect on Apoptosis | Key Experimental Readout |
|---|---|---|---|---|---|
| Caspase-9 | Sourced from literature | AKT (PKB) | In vitro kinase assays, cell culture models (e.g., MLE-12 cells) [17] | Inhibition of caspase-9 proteolytic activity; reduced cleavage of downstream effector caspases [2] | Decreased caspase-3/7 activity; reduced PARP cleavage; increased cell viability |
| Other Apoptotic Components* | Multiple | CDK1, PKA, PKC | Recombinant protein assays, overexpression/knockdown studies [52] [53] | Context-dependent inhibition or activation; alters signal transduction flow | Changes in phospho-specific antibody signal (Western Blot/Immuno-BRET); altered kinase reporter activity |
| Note: The regulatory landscape is complex. For instance, phosphorylation continuously activates proteins in pathways like MAPK, JAK/STAT, and PI3K/AKT in cancer, while inhibiting others like caspase-9 [52]. |
The mechanism of inhibition often involves steric hindrance, where the phosphate group physically blocks substrate access to the active site or induces conformational changes that inactivate the protein. In the case of caspase-9, phosphorylation can stabilize its inactive monomeric form, thereby interfering with the dimerization proposed by the induced proximity model and potentially promoting an inactive conformation [3].
Detailed Experimental Protocols for Key Findings
To enable replication and critical evaluation, this section details core methodologies used to generate key data in this field.
Protocol 1: Monitoring Phosphorylation Dynamics via Immuno-BRET
This homogeneous immunoassay leverages CRISPR/Cas9 genome editing and bioluminescence resonance energy transfer (BRET) to monitor PTMs on endogenous proteins with high sensitivity [54].
Cell Line Generation:
- CRISPR-mediated Knock-in: Design a guide RNA (gRNA) to target the C-terminus of the gene of interest (e.g.,
STAT3: CCCATGTGAGGAGCTGAGAA). Co-electroporate cells with the gRNA:Cas9 ribonucleoprotein (RNP) complex and a single-stranded oligodeoxynucleotide (ssODN) donor template containing the 1.3 kDa HiBiT peptide sequence. - Clone Isolation: After 24-48 hours, use FACS to dispense single edited cells into 96-well plates. Screen growing colonies for bioluminescence after complementation with LgBiT protein to identify positive clones [54].
- CRISPR-mediated Knock-in: Design a guide RNA (gRNA) to target the C-terminus of the gene of interest (e.g.,
Immuno-BRET Assay Execution:
- Seed HiBiT-tagged cells in white 96-well plates and treat with stimuli (e.g., growth factors, kinase inhibitors).
- Lyse cells using digitonin-containing buffer supplemented with Nano-Glo HiBiT Lytic Substrate and LgBiT protein. This reconstitutes the active NanoLuc luciferase, generating a donor bioluminescent signal (~465 nm).
- Simultaneously, incubate the lysate with a fluorescently labeled (e.g., Alexa Fluor 594) antibody specific to the PTM of interest (e.g., phospho-tyrosine).
- If the antibody binds to the HiBiT-tagged protein, BRET occurs, and light emission at the acceptor's fluorescence wavelength (~594 nm) is detected.
- The BRET ratio (acceptor emission / donor emission) reflects the degree of phosphorylation, normalized to the total protein abundance [54].
Protocol 2: Functional Validation in Disease Models
This protocol assesses the functional consequence of phosphorylation inhibition in a pathophysiological context, such as pulmonary fibrosis [17].
In Vivo Modeling:
- Induce pulmonary fibrosis in C57BL/6 mice via intratracheal bleomycin (5 mg/kg) instillation.
- Systemically administer a caspase-9 inhibitor (e.g., Z-LEHD-FMK, 10 mg/kg, i.p.) or vehicle control over 21 days.
Ex Vivo Analysis:
- Collect lung tissues for histological (H&E, Masson's trichrome for collagen) and molecular analysis.
- Perform Western blotting using antibodies against cleaved-caspase-9, collagen I, α-SMA, and other fibrotic markers.
- Assess apoptosis in lung sections via TUNEL assay [17].
In Vitro Mechanistic Studies:
- Treat alveolar epithelial cells (e.g., MLE-12) with TGF-β1 (10 ng/mL, 48 hrs) to model fibrotic stimulation.
- Transfert cells with plasmids for caspase-9 overexpression (oe-Caspase-9) or siRNA for knockdown (Caspase-9-siRNA).
- Measure apoptosis (TUNEL, caspase-3/7 activity) and fibrotic marker expression (qPCR, Western blot) to establish the caspase-9/β-catenin axis [17].
Visualizing the Regulatory Network
The following diagram synthesizes the core concepts and relationships, illustrating how phosphorylation integrates into the regulatory network controlling caspase-9 activation.
Inhibitory Phosphorylation of Caspase-9: This diagram integrates the induced proximity and induced conformation models for caspase-9 activation. A survival signal activates a kinase (e.g., AKT), which phosphorylates caspase-9. The phosphorylated form competes with the inactive monomer, preventing its apoptosome-mediated activation and subsequent apoptosis execution.
The Scientist's Toolkit: Essential Research Reagents
Table 2: Key Reagents for Investigating Inhibitory Phosphorylation
| Reagent / Tool | Function / Application | Example Use Case |
|---|---|---|
| Z-LEHD-FMK | Irreversible, cell-permeable caspase-9 inhibitor [17] | Functional validation of caspase-9's role in apoptotic and non-apoptotic pathways (e.g., pulmonary fibrosis). |
| CRISPR/Cas9 & HiBiT System | Precise tagging of endogenous proteins with a small (1.3 kDa) luminescent peptide for study under native regulation [54]. | Monitoring PTM dynamics on endogenous targets via Immuno-BRET, avoiding overexpression artifacts. |
| Phospho-specific Antibodies | Detect specific phosphorylation events in proteins (e.g., anti p-STAT3 (Y705)) [54]. | Western blot, Immuno-BRET, and immunohistochemistry to map and quantify phosphorylation. |
| Kinase Inhibitors | Selective chemical tools to inhibit specific kinase activity (e.g., SB202190 for p38 MAPK) [53]. | Probing kinase-substrate relationships and validating upstream regulators of phosphorylation. |
| PTMNavigator | Bioinformatics tool for projecting PTM datasets onto canonical signaling pathways for visualization and enrichment analysis [55]. | Contextualizing phosphoproteomics data within pathways like apoptosis to identify key regulatory nodes. |
Caspase-9 activation represents a pivotal control point in the intrinsic apoptotic pathway, yet its precise molecular mechanism continues to generate conflicting data and interpretations across experimental models. The long-standing debate primarily centers on two competing hypotheses: the induced proximity model and the induced conformation model. The former posits that caspase-9 activation occurs primarily through dimerization when brought into close proximity by the apoptosome complex, while the latter suggests that binding to the apoptosome induces essential conformational changes that drive activation [3] [12]. This fundamental discrepancy permeates the field, creating challenges for researchers and drug development professionals seeking to target this pathway therapeutically. This guide objectively compares experimental data from key studies that support each model, provides detailed methodologies for critical experiments, and presents visualizations of the competing mechanisms to facilitate reconciliation of these conflicting datasets.
Molecular Models of Caspase-9 Activation
Competing Theoretical Frameworks
The induced proximity model originated from observations that initiator caspases fused to heterologous dimerization domains underwent autoactivation and induced cell death [56]. This model was subsequently refined into a proximity-induced dimerization framework, suggesting that the apoptosome's primary function is to promote caspase-9 homo-dimerization, which is sufficient for catalytic activation [12] [56].
In contrast, the induced conformation model proposes that apoptosome binding induces essential allosteric changes in caspase-9 that cannot be achieved through dimerization alone. Supporters of this model argue that artificial dimerization produces caspase-9 with altered enzymatic properties compared to apoptosome-activated enzyme, particularly regarding substrate specificity [3] [56].
Table 1: Core Principles of Competing Activation Models
| Feature | Induced Proximity Model | Induced Conformation Model |
|---|---|---|
| Primary Mechanism | Dimerization driven by increased local concentration | Allosteric conformational change upon apoptosome binding |
| Role of Apoptosome | Caspase-9 recruitment and concentration platform | Allosteric regulator inducing active conformation |
| Key Evidence | Artificial dimerization activates caspase-9 | Apoptosome-bound caspase-9 has unique catalytic properties |
| Catalytic Outcome | Enhanced activity through dimer interface formation | Altered active site configuration and substrate specificity |
Caspase-9 Activation Pathways
The following diagram illustrates the key steps in caspase-9 activation within the intrinsic apoptosis pathway, highlighting where the competing models propose different mechanisms:
Caspase-9 Activation in Intrinsic Apoptosis
Comparative Experimental Data
Key Studies Supporting Each Model
Research from different laboratories has yielded conflicting data on caspase-9 activation, often depending on the experimental system and specific constructs employed. The table below summarizes quantitative findings from pivotal studies that support each model:
Table 2: Comparative Experimental Data from Key Caspase-9 Studies
| Experimental System | Key Findings | Supporting Model | Reference |
|---|---|---|---|
| LZ-C9 (Leucine-zipper dimer) | 2-3 fold higher activity than C9Holo for LEHD-AFC peptide substrate | Induced Proximity | [56] |
| C9Holo (Apoptosome-bound) | ~1000x greater activity for procaspase-3 than monomeric caspase-9 | Induced Conformation | [56] |
| Engineered caspase-9 dimer | Lower activity than C9Holo despite dimerization | Induced Conformation | [3] |
| CARD-deleted caspase-9 | Structural studies show dimerization interface formation | Both Models | [56] |
| Apoptosome-bound caspase-9 | Km for procaspase-3 significantly lower than LZ-C9 | Induced Conformation | [56] |
Substrate-Specific Activation Patterns
A critical reconciliation point emerges from examining substrate-specific differences in caspase-9 activity. While artificial dimerization generates caspase-9 with enhanced activity against synthetic peptide substrates, the same dimers show markedly reduced efficiency toward physiological substrates like procaspase-3:
Table 3: Substrate-Specific Activity Profiles of Different Caspase-9 Forms
| Caspase-9 Form | LEHD-AFC (Peptide) Activity | Procaspase-3 Processing | Km for Procaspase-3 |
|---|---|---|---|
| Monomeric | Low | Minimal | High (>1 µM) |
| LZ-C9 (Dimeric) | High | Reduced | ~230 nM |
| C9Holo (Apoptosome-bound) | Moderate | Very High | ~25 nM |
Detailed Experimental Protocols
Caspase-9 Dimerization Experiments
Objective: To test whether artificial dimerization is sufficient for caspase-9 activation.
Methodology:
- Construct Engineering: Replace caspase-9's CARD domain with GCN4 leucine zipper dimerization domain using molecular cloning techniques [56]
- Protein Expression: Transform plasmid into E. coli BL21(DE3) strain, grow to OD600 ~0.6-1.0, induce with 1 mM IPTG at 25-30°C for 4 hours [57]
- Protein Purification: Ni2+-affinity chromatography followed by anion exchange chromatography [57]
- Gel Filtration: Analyze oligomeric state using Superdex 200 HR 10/300 column to confirm dimer formation [56]
- Enzymatic Assays: Measure activity against fluorogenic substrate LEHD-AFC and physiological substrate procaspase-3 [56]
Key Controls:
- Wild-type caspase-9 as reference
- CARD-deleted caspase-9 without dimerization domain
- Apoptosome-activated caspase-9 (C9Holo) as gold standard
Apoptosome Reconstitution assays
Objective: To compare the activity of apoptosome-bound caspase-9 with artificially dimerized forms.
Methodology:
- Apoptosome Assembly: Incubate Apaf-1 with cytochrome c and dATP/ATP in appropriate buffer system [56]
- Caspase-9 Recruitment: Add procaspase-9 to pre-formed apoptosome complex
- Kinetic Analysis: Measure enzyme kinetics using varying concentrations of procaspase-3 substrate
- Km Determination: Calculate Michaelis constants for different caspase-9 forms using nonlinear regression [56]
Critical Parameters:
- Strict control of ionic strength and divalent cation concentrations
- Measurement of initial reaction rates to avoid product inhibition
- Parallel assays with dimeric caspase-9 constructs under identical conditions
Signaling Pathways and Experimental Workflows
Caspase-9 Experimental Decision Pathway
The following workflow illustrates how researchers can navigate the conflicting data when designing experiments related to caspase-9 activation:
Experimental Decision Pathway for Caspase-9 Research
The Scientist's Toolkit: Essential Research Reagents
Table 4: Key Research Reagents for Caspase-9 Activation Studies
| Reagent/Category | Specific Examples | Function/Application |
|---|---|---|
| Expression Plasmids | pET23b-Casp3-His, pET23b-Casp9-His, pET11a-Casp6-His [57] | Recombinant protein expression in E. coli |
| Caspase-9 Constructs | LZ-C9 (leucine zipper fusion), C9ΔCARD (CARD-deleted) [56] | Testing dimerization sufficiency |
| Chemical Inhibitors | Z-LEHD-FMK (caspase-9 inhibitor) [17] | Pharmacological inhibition in cellular models |
| Activity Assays | LEHD-AFC fluorogenic substrate [56] | In vitro enzymatic activity measurement |
| Antibodies | Anti-cleaved caspase-9 (Asp315, Asp330) [8] | Detection of specific caspase-9 activation neoepitopes |
| Apoptosis Inducers | Bleomycin, TGF-β1 [17] | Inducing intrinsic apoptosis in cellular models |
| Animal Models | Caspase-9 knockout mice [58] | In vivo functional validation |
Reconciliation of Conflicting Data
The apparent contradictions between the induced proximity and induced conformation models can be largely reconciled by recognizing that both mechanisms contribute to caspase-9 activation, but their relative importance depends on context:
Substrate-Specific Mechanism: Artificial dimerization may be sufficient for activity against simple peptide substrates, but physiological substrate processing requires apoptosome-induced conformational changes that enhance substrate affinity [56].
Context-Dependent Function: The cellular environment, including competition with endogenous inhibitors like XIAP, may favor the apoptosome activation mechanism in physiological settings [8].
Integrated Model: A hybrid model emerges where the apoptosome both dimerizes caspase-9 and induces conformational changes that optimize its active site for specific physiological substrates, particularly procaspase-3 [3] [56].
This reconciliation explains why both models retain experimental support while highlighting the importance of using physiologically relevant substrates and conditions when evaluating caspase-9 activation for drug discovery applications.
Technical Limitations of Current Methodologies and Opportunities for Innovation
Caspase-9 stands as the pivotal initiator caspase of the intrinsic apoptosis pathway, and its precise activation mechanism carries profound implications for understanding cell death, developing cancer therapeutics, and treating degenerative diseases. The central theoretical framework for understanding initiator caspase activation has long been the induced proximity model, which posits that activation occurs primarily through dimerization when caspase-9 molecules are brought into close contact by activation platforms like the apoptosome [3] [4]. However, emerging research challenges the sufficiency of this model, providing compelling evidence for an induced conformation model, where the apoptosome actively induces structural changes in caspase-9 that drive its activation [3] [4]. This ongoing debate is critically informed by the methodologies used to probe caspase-9 function, each with its own set of technical limitations and capacities for innovation.
Methodological Comparison in Caspase-9 Research
Researchers employ diverse methodologies to study caspase-9 activation, each providing unique insights and facing distinct limitations. The table below summarizes the core approaches, their applications, and key constraints.
| Methodology | Key Application | Technical Limitations | Supporting Data/Evidence |
|---|---|---|---|
| Engineered Dimer Studies [4] | Testing the induced proximity model by creating constitutively dimeric caspase-9. | Engineered dimers may not fully replicate native apoptosome-induced activation; activity is only a fraction of Apaf-1-activated caspase-9 [4]. | Dimeric caspase-9 showed higher activity than wild-type monomers but had only ~5-10% the activity of apoptosome-activated wild-type enzyme [4]. |
| Proteolytic Mutant Analysis [59] | Determining the role of proteolytic processing in caspase-9 activation using cleavage-site mutants. | Triple mutant (E306A/D315A/D330A) cripples activity, but cannot distinguish between conformation and proximity effects [59]. | Uncleavable caspase-9 triple mutant (Casp9-TM) exhibited "drastically reduced" protease activity and impaired homodimer formation in the presence of apoptosome [59]. |
| FRET-Based Live-Cell Imaging [60] | Visualizing real-time caspase-9 activation kinetics in live cells. | Requires specialized probe design and equipment; signal interpretation can be complex due to cellular heterogeneity [60]. | Custom bioprobes enabled simultaneous observation of caspase-9 and -3 dynamics, revealing that cumulative caspase-9 activity regulates caspase-3 execution timing [60]. |
| High-Content Screening (HCS) [61] | Automated, multiparameter screening of caspase activation in live cells for drug discovery. | Fluorogenic substrates (e.g., DEVD-NucView488) may have toxicity over long periods and require optimization for multiple measurements over time [61]. | HCS allowed quantification of caspase activation in the same cell population over several days, identifying differential responses to chemotherapeutics [61]. |
| Caspase-9 Antibody-Based Detection [29] | Detecting and quantifying Caspase-9 protein levels and cleavage in research and diagnostics. | Subject to batch-to-batch variability; may not distinguish between functional states without additional functional assays [29]. | Used for Western blot, immunohistochemistry, and flow cytometry; evolving toward higher precision and automation by 2025 [29]. |
Detailed Experimental Protocols and Data
Engineered Dimeric Caspase-9 Assay
This protocol is designed to test the hypothesis that dimerization alone is sufficient for full caspase-9 activation, a cornerstone of the induced proximity model.
- Rationale: Wild-type caspase-9 exists predominantly as a monomer in solution. To create a constitutively dimeric form, researchers engineered the protein by mutating residues on the β6 strand (e.g., Phe404) that cause steric hindrance at the dimer interface, based on structural comparisons with caspase-3 [4].
- Expression and Purification: The engineered caspase-9 gene is cloned into an expression vector (e.g., pET-29b) with a C-terminal hexahistidine tag and expressed in E. coli. The protein is then purified using affinity chromatography (e.g., nickel-nitrilotriacetic acid columns) followed by gel filtration to confirm dimeric state [4].
- Activity Assay: Catalytic activity is measured in vitro using fluorogenic substrates (e.g., Ac-LEHD-AFC). The key experiment involves comparing the activity of the dimeric caspase-9 with that of wild-type caspase-9 activated by the Apaf-1 apoptosome [4].
- Data Interpretation: The finding that the engineered dimer has significantly lower activity than the apoptosome-activated wild-type enzyme (~5-10%) challenges the induced proximity model and suggests that the apoptosome provides more than just a platform for dimerization, potentially inducing critical conformational changes [3] [4].
Analysis of Proteolytic Processing Role
This methodology investigates the necessity of intrachain cleavage for caspase-9 activation, a process previously thought to be dispensable.
- Mutant Generation: An uncleavable caspase-9 zymogen is generated via site-directed mutagenesis of three aspartic acid cleavage sites (E306A/D315A/D330A), known as Casp9-TM [59].
- Apoptosome Reconstitution: The apoptosome is assembled in vitro by incubating full-length Apaf-1 with cytochrome c and dATP [59].
- Functional Assessment: The activity of wild-type versus Casp9-TM caspase-9 is measured in the presence of the apoptosome using two methods: (1) cleavage of a fluorogenic substrate (Ac-LEHD-AFC), and (2) the ability to activate its physiological substrate, procaspase-3, monitored by SDS-PAGE [59].
- Key Result: The Casp9-TM mutant showed a "drastically reduced" ability to be activated by the apoptosome and a "markedly reduced ability" to form homodimers, indicating that proteolytic processing is essential for full activation, likely by stabilizing an active conformation [59].
Live-Cell Imaging with FRET-Based Bioprobes
This advanced protocol allows for multiplexed, real-time observation of caspase signaling flow in live cells, capturing population heterogeneity.
- Bioprobe Design: A chimeric molecular sensor is constructed with a donor fluorescent protein (e.g., GFP), a caspase-9 recognition sequence (LEHD), and an acceptor fluorescent dye (e.g., Alexa Fluor 532). In the intact probe, FRET occurs; upon cleavage by active caspase-9, FRET is disrupted, leading to a measurable change in the donor-to-acceptor fluorescence ratio [60].
- Cell Preparation and Imaging: Cells (e.g., HeLa) are loaded with the bioprobe and treated with apoptotic inducers (e.g., TNF-α/cycloheximide). Fluorescence is tracked over time using live-cell microscopy [60].
- Data Analysis: Single-cell FRET ratio trajectories are analyzed to determine the timing and magnitude of caspase-9 activation. Statistical analysis across the cell population can reveal correlations, such as the finding that cumulative caspase-9 activity, not its rate, inversely regulates the time to caspase-3 activation [60].
Caspase-9 Activation Pathway Models
The following diagrams illustrate the key mechanistic models for caspase-9 activation and the experimental workflow used to differentiate between them.
Caspase-9 Activation Mechanisms
Experimental Workflow for Model Differentiation
The Scientist's Toolkit: Key Research Reagents and Materials
A successful research program in caspase-9 activation relies on a suite of specialized reagents and tools.
| Reagent / Material | Function / Application | Key Characteristics & Considerations |
|---|---|---|
| Caspase-9 Antibodies [29] [17] | Detection of Caspase-9 expression and cleavage (e.g., via Western Blot, IHC). | Select antibodies specific for full-length vs. cleaved forms (e.g., Asp315); be mindful of batch-to-batch variability. |
| Fluorogenic Caspase Substrates [61] [60] | Quantifying caspase activity in vitro and in live cells. | Ac-LEHD-AFC/-AMC for caspase-9; DEVD-based substrates for effector caspases. DEVD-NucView488 is cell-permeable for live-cell imaging. |
| Caspase-9 Inhibitors [17] | Functional studies to probe caspase-9 role in biological processes. | Z-LEHD-FMK is a selective, cell-permeable caspase-9 inhibitor used in vitro and in vivo. |
| Recombinant Proteins [59] [4] | In vitro reconstitution of apoptosome and activity assays. | Require high-purity recombinant Apaf-1, caspase-9 (wild-type and mutants), and cytochrome c. |
| FRET-Based Bioprobes [60] | Real-time, multiplexed imaging of caspase activity in live cells. | Customizable components (donor FP, recognition sequence, acceptor dye) allow optimization for specific imaging systems. |
| Expression Vectors [59] [4] | Overexpression of caspase-9 variants and Apaf-1 in cell lines. | Vectors with tags (e.g., His-tag) facilitate purification for biochemical studies. |
The debate between the induced conformation and induced proximity models of caspase-9 activation is far from settled, driven forward by the continuous evolution of research methodologies. While traditional biochemical and structural approaches have revealed the insufficiency of simple dimerization, newer live-cell imaging and high-content screening technologies are uncovering the dynamic and heterogeneous nature of caspase signaling in physiological contexts. The primary limitations of current tools—their inability to fully replicate the native activation environment, their temporal resolution, and their dependency on engineered probes—directly map onto the frontiers of innovation. The future of this field lies in the development of even more sensitive, non-perturbative biosensors, the integration of single-cell 'omics' data with functional imaging, and the application of structural biology to visualize the apoptosome-caspase-9 holoenzyme in unprecedented detail. Overcoming these technical hurdles will not only resolve a fundamental mechanistic question in cell biology but also accelerate the discovery of novel therapeutics that modulate cell death by targeting the caspase-9 activation pathway.
Weighing the Evidence: A Comparative Analysis of Induced Proximity vs. Induced Conformation
The mechanism of caspase-9 activation is a pivotal question in apoptosis research, centering on two principal models: the induced proximity model, which posits that the apoptosome serves as a platform for caspase-9 dimerization, and the induced conformation model, which suggests the apoptosome imposes allosteric changes that activate monomeric caspase-9. This comparison guide objectively evaluates the catalytic activity of engineered dimeric caspase-9 against apoptosome-bound caspase-9. Synthesizing key experimental data, we demonstrate that while induced dimerization enhances basal activity, it is insufficient to replicate the robust catalytic efficiency conferred by integration into the apoptosome complex. The emerging paradigm supports a hybrid model where the apoptosome facilitates both dimerization and unique activating conformational changes, with distinct functional consequences for homo- and heterodimeric caspase-9 complexes.
In the intrinsic apoptotic pathway, caspase-9 functions as the initiator caspase, activated through its incorporation into a multi-protein complex known as the apoptosome, composed of Apaf-1 and cytochrome c [1] [62]. The precise mechanism of its activation, however, has been the subject of intense scientific debate, forming a critical thesis in cell death research.
- Induced Proximity/Dimerization Model: This model asserts that initiator caspases are monomers in solution and are activated when their local concentration is dramatically increased on a signaling platform, such as the apoptosome. This proximity drives homodimerization, which is the primary event leading to catalytic activation [12]. The apoptosome's role, in this view, is largely passive [3].
- Induced Conformation/Holoenzyme Model: This competing hypothesis proposes that the apoptosome plays an active role by inducing specific allosteric changes in caspase-9. The oligomerized Apaf-1 is thought to generate a unique holoenzyme that conformationally activates bound caspase-9 monomers, a process qualitatively different from simple dimerization [3] [4].
This guide provides a direct comparative analysis of the activity of dimeric caspase-9 versus apoptosome-bound caspase-9, presenting key experimental data and methodologies that have shaped this ongoing scientific discussion.
Quantitative Comparison of Caspase-9 Activity
The catalytic proficiency of caspase-9 in its different states is best illustrated through direct quantitative comparison of key enzymatic parameters, as summarized in the table below.
Table 1: Direct comparison of catalytic activity between dimeric and apoptosome-bound caspase-9
| Feature | Engineered Dimeric Caspase-9 | Apoptosome-Bound Caspase-9 | Experimental Context & Citation |
|---|---|---|---|
| Catalytic Activity | "Only a small fraction" of apoptosome-activated activity; enhanced over wild-type monomer but not equivalent to full activation [4]. | Significantly higher catalytic activity than the engineered dimer; represents the fully activated state [4]. | In vitro cleavage assays comparing the engineered dimer to caspase-9 activated by the reconstituted apoptosome [4]. |
| Activation by Apaf-1 | Activity "can no longer be significantly enhanced in an Apaf-1-dependent manner" [4]. | Absolutely dependent on Apaf-1 apoptosome assembly for full activity [23] [63]. | Assays measuring caspase-9 activity with and without addition of Apaf-1/cytochrome c [4]. |
| Cell Death Induction | Induces "more efficient cell death when expressed" compared to wild-type monomeric caspase-9 [4]. | The physiological, high-efficiency trigger for intrinsic apoptosis [1]. | Cell-based assays measuring apoptosis upon expression of engineered proteins [4]. |
| Dimerization State | Constitutive homodimer stabilized by engineered mutations at the dimer interface (e.g., F404D) [23] [4]. | Can form both caspase-9 homodimers and caspase-9/Apaf-1 heterodimers on the apoptosome platform [23] [63]. | Site-specific crosslinking and biophysical analysis within the reconstituted apoptosome [23] [63]. |
| Substrate Cleavage Preference | Primarily mediates intramolecular autoprocessing at Asp-315 [23]. | Heterodimers with Apaf-1 more efficiently activate the downstream substrate procaspase-3 [23]. | Substrate cleavage assays with specific caspase-9 constructs and substrates [23]. |
The data clearly indicate that the activity of a simple caspase-9 homodimer, while elevated, is qualitatively different from and quantitatively inferior to the activity displayed when caspase-9 is incorporated into the apoptosome.
Detailed Experimental Protocols for Key Studies
Engineering a Constitutive Dimeric Caspase-9
This experiment was designed to test the core premise of the induced proximity model by creating a stable, dimeric caspase-9 independent of the apoptosome [4].
- Objective: To determine if enforced dimerization is sufficient to fully activate caspase-9 to a level equivalent to apoptosome-mediated activation.
- Methodology:
- Rational Design: Based on comparative structural analysis with effector caspases like caspase-3, researchers identified steric clashes at the dimer interface, specifically involving Phe404, that destabilize caspase-9 dimerization.
- Protein Engineering: Mutations were introduced at the dimer interface (e.g., F404D) to relieve steric hindrance and create a constitutively dimeric caspase-9 protein.
- Biophysical Validation: The engineered protein was confirmed to be a stable dimer in solution using techniques like size-exclusion chromatography, in contrast to the wild-type protein, which is predominantly monomeric.
- Activity Assays: The catalytic activity of the engineered dimer was quantitatively measured using fluorogenic peptide substrates (e.g., LEHD-amc) and compared directly to the activity of wild-type caspase-9 activated by a reconstituted Apaf-1 apoptosome.
- Key Result: The engineered dimer showed increased activity over the wild-type monomer but reached only a small fraction of the activity achieved by the apoptosome-bound caspase-9. Furthermore, the dimer's activity could not be further stimulated by Apaf-1 [4].
Demonstrating Dimerization within the Apoptosome
This study provided the first direct biochemical evidence for caspase-9 homodimerization within the apoptosome, while also revealing a more complex picture [23].
- Objective: To directly probe the oligomeric state of caspase-9 during its activation on the apoptosome.
- Methodology:
- Site-Specific Crosslinking: A novel technique employing an unnatural amino acid (L-DOPA) was used to incorporate a chemical crosslinker at the dimer interface (Phe406) of caspase-9.
- Apoptosome Reconstitution: The modified procaspase-9 was incorporated into a functional apoptosome complex in vitro.
- Crosslinking and Analysis: Oxidation-induced crosslinking successfully trapped caspase-9 homodimers, proving their existence in the complex. Surprisingly, the same method also captured crosslinks between caspase-9 and Apaf-1, indicating the formation of caspase-9/Apaf-1 heterodimers.
- Functional Characterization: Saturation experiments revealed that even monomeric caspase-9 bound to the apoptosome hub (via Apaf-1 NOD domains) could activate procaspase-3, a hallmark of heterodimer activity [23].
- Key Result: The apoptosome supports the formation of both caspase-9 homodimers and caspase-9/Apaf-1 heterodimers, with the latter being more efficient at activating downstream effector caspases [23].
Visualization of Caspase-9 Activation Pathways
The following diagram synthesizes the experimental findings to illustrate the pathway of caspase-9 activation on the apoptosome, integrating both homo- and heterodimerization events.
The Scientist's Toolkit: Key Research Reagents
The following table catalogues essential reagents and their applications for studying caspase-9 activation, as evidenced in the cited research.
Table 2: Key research reagents for investigating caspase-9 activation mechanisms
| Reagent / Method | Function in Research | Key Experimental Insight |
|---|---|---|
| Recombinant Apaf-1 & Cytochrome c | To reconstitute the functional apoptosome complex in a cell-free system [23]. | Allows for the biochemical dissection of caspase-9 activation independent of cellular upstream signals. |
| Non-Cleavable Caspase-9 Mutant (e.g., ProC9-TM) | A caspase-9 mutant that cannot undergo autoprocessing, used to decouple dimerization from cleavage events [23]. | Revealed that cleavage is not required for activity but regulates dissociation from the apoptosome (molecular timer) [23]. |
| Dimerization-Defective Mutant (e.g., F404D) | A caspase-9 mutant with a disrupted dimer interface, used to test the necessity of dimerization [23]. | Demonstrated that dimerization is essential for stable recruitment and full activity of caspase-9 on the apoptosome [23]. |
| Kosmotropic Salts (e.g., Ammonium Citrate) | Chemicals that enhance protein-protein interactions, used to artificially induce caspase dimerization in solution [64] [23]. | Provides supporting evidence that dimerization can increase basal caspase-9 activity, but this activity is less robust than apoptosome-driven activation [64]. |
| Site-Specific Crosslinking (e.g., L-DOPA incorporation) | A sophisticated biochemical technique to trap and identify transient protein-protein interactions directly within a complex [23]. | Provided the first direct proof of both caspase-9 homodimerization and caspase-9/Apaf-1 heterodimerization within the apoptosome [23]. |
The direct comparative analysis leads to a synthesized conclusion that moves beyond a simple dichotomy:
- Dimerization is Necessary but Not Sufficient: The experimental data refute the idea that induced proximity-driven homodimerization is the sole mechanism for caspase-9 activation. While dimerization is a key event that enhances the avidity of caspase-9 for the apoptosome and promotes its initial activation, an engineered dimer cannot replicate the full catalytic power of the apoptosome-bound enzyme [4].
- The Apoptosome is an Allosteric Regulator: The apoptosome platform provides more than a passive tether; it actively participates in catalysis. The formation of caspase-9/Apaf-1 heterodimers demonstrates a direct allosteric role for Apaf-1, likely inducing conformational changes that optimize the caspase-9 active site for cleaving downstream substrates like procaspase-3 [23] [63].
- A Hybrid Model for Activation: The current paradigm integrates both models. The apoptosome increases local concentration to drive proximity-induced homodimerization, which is critical for autocleavage and stable binding. Concurrently, specific interactions with the oligomerized Apaf-1 hub induce conformational changes that are essential for achieving maximum catalytic competence against effector caspases [3] [23] [63]. This hybrid mechanism ensures the precise and powerful activation of the intrinsic apoptotic pathway.
The apoptosome, a central signaling platform in the intrinsic apoptotic pathway, has long been recognized for its essential role in activating caspase-9 and initiating programmed cell death. For decades, the scientific community has debated the precise mechanism by which this multiprotein complex regulates caspase-9 activation, with two primary models emerging: the induced proximity model (dimerization-driven) and the induced conformation model (allosteric activation). Historically, the induced proximity model dominated the literature, proposing that the apoptosome serves primarily to bring caspase-9 molecules into close proximity, facilitating homodimerization and subsequent activation [3]. However, emerging structural and biochemical evidence now reveals a more complex picture, demonstrating that the apoptosome facilitates multivalent interactions that extend beyond simple dimerization. This paradigm shift recognizes that caspase-9 activation involves a sophisticated network of homo- and heterodimeric interactions, allosteric regulation, and precise stoichiometric relationships that collectively determine the cell's fate [23] [39].
This comparison guide examines the evolving understanding of apoptosome structure and function by critically evaluating key experimental approaches that have illuminated these multivalent interactions. We objectively analyze data from structural studies, biochemical assays, and cellular experiments to provide researchers and drug development professionals with a comprehensive resource for understanding the molecular intricacies of apoptosome-mediated caspase activation.
Structural Insights into Multivalent Apoptosome Architecture
CARD Disk Organization and Stoichiometry
High-resolution structural studies have revolutionized our understanding of apoptosome organization, particularly regarding the CARD (Caspase Recruitment Domain) disk that forms atop the central hub. Cryo-EM analyses reveal that the human apoptosome forms an accentric, spiral-shaped disk comprising CARDs from both Apaf-1 and procaspase-9, creating a structural arrangement that fundamentally enables multivalent interactions.
Table 1: Comparative Structural Features of Apoptosomes Across Species
| Organism | Adapter Protein | Oligomeric State | Caspase Recruited | CARD Disk Organization | Key Structural Features |
|---|---|---|---|---|---|
| C. elegans | CED-4 | Tetramer (8 subunits) | CED-3 | Two CED-4 CARD tetrameric ring layers | 8:2 CED-4 to CED-3 stoichiometry; activation via L2' loop and "hutch" interactions |
| D. melanogaster | Dark | Octamer | Dronc | 8 Dark CARDs bind 8 Dronc CARDs | Strict 8:8 stoichiometry; β-propellers stabilize assembly but don't bind cytochrome c |
| H. sapiens | Apaf-1 | Heptamer | Procaspase-9 | Spiral with 3-4 pc-9 CARDs bound to Apaf-1 CARDs | Acentric disk with mismatch to platform symmetry; facilitates multiple interaction modes |
The human apoptosome exhibits a stoichiometric mismatch between its heptameric platform and the CARD disk, which typically contains only three to four procaspase-9 molecules despite having seven potential binding sites [65] [39]. This arrangement creates a structural environment where a single procaspase-9 catalytic domain may be "parked" on the central hub while others engage in dimerization, enabling functional specialization within the complex [65]. This sophisticated architecture contrasts with the simpler, symmetric organizations observed in C. elegans and D. melanogaster, highlighting evolutionary specialization in the human apoptosome that supports multivalent interactions.
Visualizing the Multivalent Apoptosome
The following diagram illustrates the key structural relationships and multivalent interactions within the human apoptosome, based on recent near-atomic resolution structures:
Diagram 1: Multivalent Interactions in the Human Apoptosome. This schematic illustrates the key structural relationships and interaction modes within the active apoptosome, highlighting the acentric CARD disk and multiple dimerization possibilities.
Experimental Approaches for Studying Multivalent Interactions
Structural Biology Techniques
Cryo-Electron Microscopy (Cryo-EM) has been instrumental in revealing the architectural principles governing apoptosome function. The near-atomic structure of the human apoptosome (∼4.1 Å resolution) provided unprecedented insights into cytochrome c binding, nucleotide exchange, and conformational changes driving assembly [65]. Technical advances including energy filtering to remove inelastically scattered electrons and movie frame corrections for global movements during imaging have been critical for achieving this resolution [65]. The resulting models demonstrate how an acentric disk containing four Apaf-1/procaspase-9 CARD pairs arranged in a shallow spiral creates the structural foundation for multivalent interactions, with the fourth procaspase-9 CARD exhibiting lower occupancy, suggesting dynamic binding [65].
X-ray Crystallography has complemented these findings, particularly in elucidating death domain superfamily interactions. Docking models of key complex components based on crystal structures of Pelle/Tube and Apaf-1/procaspase-9 revealed that death domain superfamily members have potential for multivalent interactions, enabling novel regulatory mechanisms for apoptotic signaling [66]. These structural approaches collectively demonstrate that the apoptosome is not a simple symmetric platform but rather a sophisticated machine with specialized functional regions that facilitate multiple interaction modes.
Biochemical and Biophysical Methods
Site-Specific Crosslinking techniques have provided direct evidence for procaspase-9 homodimerization within the apoptosome, demonstrating markedly increased avidity for the complex and inducing selective intramolecular cleavage at Asp-315 [23]. This novel biochemical approach confirmed that the apoptosome mediates the formation of both caspase-9 homo- and heterodimers, both impacted by cleavage and contributing to overall function.
Size-Exclusion Chromatography Coupled to Multi-Angle Light Scattering (SEC-MALS) has been employed to assess the oligomeric states of caspase-9 variants. These experiments revealed that at high concentrations (40 μM), recombinant non-cleavable procaspase-9 (ProC9-TM) forms homodimers in solution, while the fully processed caspase-9 (C9-p35/p12) does not, indicating that autocatalytic cleavage regulates dimerization propensity [23].
Engineered Caspase Dimerization studies have further illuminated the relationship between dimerization and activation. When caspase-9 was engineered to exist as a constitutive homodimer, it exhibited increased activity in vitro and induced more cell death when expressed in cells compared to wild-type enzyme [3]. However, this activity remained only a fraction of that achieved by Apaf-1-activated wild-type caspase-9, suggesting that dimerization alone is insufficient for full activation and supporting a role for additional regulatory mechanisms [3].
Table 2: Key Experimental Approaches for Studying Multivalent Interactions
| Methodology | Experimental Application | Key Findings | Technical Considerations |
|---|---|---|---|
| Cryo-EM | Near-atomic structure determination of active human apoptosome | Revealed acentric CARD disk with 3-4 pc-9 molecules; stoichiometric mismatch with platform | Resolution not isotropic (3-4 Å central hub, 4.5-10 Å peripheral domains) |
| Site-Specific Crosslinking | Direct detection of caspase-9 homo- and heterodimers | First direct evidence of procaspase-9 homodimerization within apoptosome | Requires introduction of specific crosslinkable residues without disrupting native interactions |
| SEC-MALS | Characterization of caspase-9 oligomeric states | ProC9-TM forms homodimers at high concentration; C9-p35/p12 remains monomeric | High protein concentrations required may not reflect physiological conditions |
| Engineered Dimers | Artificial enforcement of caspase-9 dimerization | Dimeric caspase-9 more active than wild-type but less than apoptosome-bound form | Confirms dimerization contributes to but isn't sufficient for full activation |
| Fluorescence Assays | Monitoring LEHDase activity of apoptosome-bound caspase-9 | Provides functional readout of caspase activation state | Measures activity but not specific oligomeric state |
The following diagram outlines a generalized experimental workflow for investigating multivalent interactions in the apoptosome, integrating multiple methodological approaches:
Diagram 2: Experimental Workflow for Apoptosome Studies. This flowchart outlines the integrated methodological approach for investigating multivalent interactions in the apoptosome, from sample preparation to data integration.
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Research Reagents for Apoptosome Studies
| Reagent / Tool | Function / Application | Experimental Utility | Key Findings Enabled |
|---|---|---|---|
| Recombinant Apaf-1 | Core apoptosome scaffold protein | In vitro reconstitution of functional apoptosomes | Enabled biochemical characterization of assembly requirements and caspase activation mechanisms |
| Non-cleavable Procaspase-9 (ProC9-TM) | Caspase-9 mutant resistant to autocleavage | Distinguishes roles of recruitment vs. cleavage in activation | Demonstrated that cleavage regulates caspase-9 affinity for apoptosome and initiates "molecular timer" |
| Z-LEHD-FMK TFA | Caspase-9 inhibitor | Pharmacological inhibition in cellular and animal models | Confirmed caspase-9 role in pulmonary fibrosis; established caspase-9/β-catenin axis in disease [17] |
| Catalytically Inactive Procaspase-3 (ProC3*) | Caspase-9 substrate proxy | Measures caspase-9 activation without feedback complications | Revealed differential activities of caspase-9 variants and dimerization mutants |
| F404D Caspase-9 Mutant | Disrupted GCFNF dimerization motif | Tests necessity of homodimerization for activation | Confirmed dimerization interface essential for caspase-9 activity in apoptosome context [23] |
| Ammonium Citrate | Kosmotropic salt | Artificial induction of caspase dimerization | Demonforced dimerization can activate caspase-9 independently of apoptosome [23] |
Integrated Model of Caspase-9 Activation
The emerging paradigm of apoptosome function integrates elements from both the induced proximity and induced conformation models, recognizing that caspase-9 activation involves a sophisticated coordination of multiple interaction types. The current evidence supports a model where the apoptosome serves as a dynamic platform that facilitates both caspase-9 homodimerization through proximity effects and allosteric activation through heterodimeric interactions with Apaf-1 [23] [39].
This integrated model functions as a "molecular timer" that regulates the duration of apoptosome activity through a carefully orchestrated sequence of events [23] [67]. Procaspase-9 initially binds to the apoptosome with high affinity, primarily through CARD-CARD interactions. The increased local concentration promotes homodimerization via the GCFNF motif in the small subunits, significantly enhancing avidity for the complex through multivalent interactions [23]. Following activation, autocatalytic cleavage at Asp-315 occurs, producing caspase-9-p35/p12. This cleavage event inhibits homodimerization, reduces affinity for the apoptosome, and initiates the release of processed caspase-9 from the complex [23]. Feedback cleavage by caspase-3 at Asp-330 then generates caspase-9-p35/p10, which partially restores activity by removing the intersubunit linker that interferes with dimerization [68]. Throughout this process, the apoptosome also facilitates the formation of caspase-9/Apaf-1 heterodimers through interactions between the caspase-9 small subunit and the Apaf-1 NOD domain, creating an alternative activation mechanism that more efficiently activates procaspase-3 [23].
This sophisticated multivalent interaction network enables precise control over one of the most critical decisions in cellular fate, demonstrating how the apoptosome integrates multiple regulatory inputs to determine signaling outcomes.
The evolving understanding of apoptosome function from a simple dimerization platform to a sophisticated multivalent interaction hub has profound implications for both basic research and therapeutic development. The evidence from structural, biochemical, and cellular studies consistently demonstrates that caspase-9 activation involves a coordinated network of interactions that extend well beyond simple proximity-induced dimerization.
For researchers investigating apoptotic signaling, these findings underscore the importance of considering the full complexity of apoptosome architecture and the dynamic nature of its component interactions. The recognition that the apoptosome facilitates both homo- and heterodimeric interactions, employs stoichiometric mismatches to create specialized functional regions, and operates as a molecular timer through cleavage-regulated affinity changes provides a more comprehensive framework for designing experiments and interpreting results.
For drug development professionals, the multivalent nature of apoptosome interactions presents both challenges and opportunities. The identification of specific interaction interfaces, particularly the homo- and heterodimerization motifs, offers potential targets for therapeutic intervention in diseases involving dysregulated apoptosis. Additionally, the emerging role of caspase-9 in non-apoptotic processes, such as its recently identified function in promoting pulmonary fibrosis through β-catenin signaling [17], highlights the importance of understanding the full functional repertoire of this complex.
As structural biology techniques continue to advance, providing even higher resolution insights into these complexes, and chemical biology approaches develop more sophisticated tools for probing specific interactions, our understanding of multivalent interactions in the apoptosome will undoubtedly continue to evolve, potentially revealing new therapeutic avenues for manipulating this critical cellular decision-point.
The precise molecular mechanism governing caspase-9 activation represents a fundamental question in apoptosis research with significant implications for therapeutic development. For decades, two competing models have dominated the scientific discourse: the induced proximity model and the induced conformation model. The induced proximity hypothesis posits that the apoptosome primarily serves as a platform to concentrate caspase-9 monomers, driving their dimerization and subsequent autoactivation [1] [3]. In contrast, the induced conformation model argues that binding to the apoptosome induces specific structural changes in caspase-9 that directly enhance its catalytic activity [1] [4]. This guide objectively evaluates experimental evidence for both models, synthesizes key findings into a hybrid framework, and provides researchers with essential methodologies and reagents for further investigation.
Model Comparison: Theoretical Frameworks and Experimental Evidence
Core Principles of Competing Models
Induced Proximity Model: This model suggests that initiator caspases like caspase-9 autoactivate when brought into close proximity. The apoptosome, with its 7-fold symmetry, acts as a scaffold that increases the local concentration of caspase-9 zymogens, facilitating homodimerization through their intrinsic dimerization interfaces. The dimerization itself is considered the primary activation event [3] [4].
Induced Conformation Model: This model proposes that caspase-9 activation requires specific conformational changes induced by its binding to the apoptosome. The interaction between the caspase activation and recruitment domains (CARD) of Apaf-1 and caspase-9 creates a complementary interface that alters the enzyme's active site, enhancing its catalytic efficiency beyond what simple dimerization can achieve [1] [4].
Key Experimental Findings and Critical Evaluation
Table 1: Comparative Experimental Evidence for Caspase-9 Activation Models
| Experimental Approach | Prediction of Induced Proximity | Prediction of Induced Conformation | Key Findings | Interpretation & Limitations |
|---|---|---|---|---|
| Engineered Dimeric Caspase-9 [4] | Engineered dimer should exhibit activity comparable to apoptosome-activated caspase-9. | Engineered dimer would show only partial activity compared to apoptosome-activated caspase-9. | Dimeric caspase-9 showed increased activity over wild-type monomers, but only a fraction of Apaf-1-activated activity. Its activity was not enhanced by Apaf-1. | Dimerization is insufficient for full activation. Apaf-1 provides activation beyond mere proximity, supporting a conformational role. |
| CARD Domain Interactions [1] | Dimerization is the central event; CARD interaction is primarily for recruitment. | Specific CARD-CARD interactions directly induce activating conformational changes. | Structural studies revealed a multimeric CARD complex with three distinct interfaces essential for activation, beyond a simple 1:1 interaction. | The complexity of the CARD platform suggests it plays an active role in structuring caspase-9 for activation, beyond simple tethering. |
| Activity & Cleavage Status [1] | Cleavage of caspase-9 is essential for its activation. | Uncleaved caspase-9 can be fully active when bound to the apoptosome. | Uncleaved caspase-9 bound to the apoptosome exhibits significant catalytic activity, which is reduced upon cleavage and dissociation. | Activation is linked to apoptosome binding, not proteolytic cleavage, which may instead act as a "molecular timer" to limit activity [1]. |
| Nucleotide Binding Role [69] | Proximity is the sole requirement. | Binding events induce specific allosteric changes in the complex. | Cytochrome c promotes dATP/ATP binding to Apaf-1, which is crucial for apoptosome formation and caspase-9 activation. A non-hydrolyzable ATP analog could support activation. | Nucleotide binding is a key event induced by cytochrome c, leading to stable apoptosome formation, aligning with a conformational control mechanism. |
Deciphering the Activation Mechanism: Essential Experimental Methodologies
To investigate these models, researchers employ several key biochemical and cellular assays. Below are detailed protocols for critical experiments cited in the literature.
In Vitro Reconstitution of Apoptosome and Caspase-9 Activation Assay
This protocol is used to study the minimal components required for activation and to test the activity of engineered caspase-9 variants [4] [69].
- Objective: To reconstitute the apoptosome in vitro and quantitatively measure its ability to activate caspase-9.
- Materials:
- Purified recombinant Apaf-1, cytochrome c, procaspase-9, and effector caspases (e.g., caspase-3).
- dATP or ATP.
- Reaction buffer (e.g., 20 mM HEPES-KOH pH 7.5, 10 mM KCl, 1.5 mM MgCl2, 1 mM EDTA, 1 mM EGTA, 1 mM DTT).
- Caspase substrate (e.g., Ac-LEHD-AFC for caspase-9 or Ac-DEVD-AFC for effector caspases).
- Fluorometric plate reader.
- Procedure:
- Apoptosome Formation: Incubate Apaf-1 (e.g., 50-100 nM) with cytochrome c (10-50 μM) and dATP/ATP (1-5 mM) in reaction buffer for 30-60 minutes at 30°C.
- Caspase-9 Activation: Add purified, monomeric procaspase-9 (e.g., 20-50 nM) to the pre-formed apoptosome and incubate for an additional 30-60 minutes.
- Activity Measurement:
- Direct: Add the caspase-9-specific substrate Ac-LEHD-AFC directly to the activation mixture and monitor the release of the fluorescent AFC moiety over time.
- Downstream: To assess functional activation, add purified procaspase-3 and its substrate Ac-DEVD-AFC to the mixture. The cleavage of the DEVD substrate indicates successful caspase-9-mediated activation of the effector caspase.
- Key Application: This assay was pivotal in showing that an engineered, constitutively dimeric caspase-9 has low basal activity that is not significantly enhanced by addition of Apaf-1, in contrast to wild-type monomeric caspase-9 [4].
Single-Cell Analysis of iCasp9-Mediated Cell Death
This approach quantifies cell-to-cell variability in caspase activation and death, revealing how protein expression levels influence the efficacy of each model [70].
- Objective: To monitor the dynamics of induced caspase-9 dimerization, downstream caspase activation, and cell fate in single live cells.
- Materials:
- Cell line (e.g., HeLa) stably expressing iCasp9 (caspase-9 fused to a dimerization domain, e.g., DmrB) tagged with a fluorescent protein (e.g., mCherry).
- FRET-based Caspase-3 reporter (e.g., CFP-DEVDR-Venus).
- Chemical Inducer of Dimerization (CID), e.g., AP20187.
- Live-cell fluorescence microscope with environmental control.
- Image analysis software for single-cell tracking.
- Procedure:
- Cell Preparation: Seed cells expressing iCasp9-mCherry and the Caspase-3 FRET reporter into a multi-well imaging plate.
- Baseline Imaging: Acquire baseline images for mCherry (iCasp9), CFP, and Venus channels.
- Induction & Time-Lapse Imaging: Add CID to the medium and immediately begin time-lapse imaging, collecting data points every 15-30 minutes for 24-48 hours.
- Data Extraction:
- iCasp9 Dimerization: Track the fluorescence intensity of iCasp9-mCherry. A decrease indicates dimerization and potential activation [70].
- Caspase-3 Activation: Calculate the FRET ratio (CFP/Venus). A sharp increase indicates Caspase-3 cleavage and activation.
- Cell Fate: Correlate the FRET ratio dynamics with morphological changes of apoptosis (e.g., membrane blebbing, shrinkage).
- Key Application: This method demonstrated that heterogenous cell fates are dictated by the initial levels of iCasp9 and the ratio of XIAP to Caspase-3, providing a quantitative framework for understanding resistance in therapeutic contexts [70].
Visualizing the Pathways and Experimental Logic
The following diagrams illustrate the core concepts and experimental workflows discussed in this guide.
Caspase-9 Activation Models and Regulatory Network
Engineered Dimer Experiment Workflow
The Scientist's Toolkit: Key Research Reagents and Models
Table 2: Essential Reagents for Caspase-9 Activation Research
| Reagent / Model | Function / Purpose | Key Characteristics & Applications |
|---|---|---|
| Reconstituted Apoptosome System [69] | To study minimal components required for caspase-9 activation in vitro. | Purified recombinant Apaf-1, cytochrome c, and caspase-9. Allows for controlled manipulation of components (e.g., nucleotides, inhibitors). |
| Inducible Caspase-9 (iCasp9) [1] [70] | A controlled model for studying caspase-9 activation in cells. | Caspase-9 fused to a drug-inducible dimerization domain (e.g., DmrB). Activation is triggered by CID (AP20187/AP1903). Widely used in cell therapy as a safety switch. |
| Caspase-9 Fluorescent Reporters [70] | To visualize caspase-9 dimerization and activity in live cells. | iCasp9 tagged with mCherry (dimerization quenches signal). FRET-based caspase-3/7 reporters (e.g., CFP-DEVDR-Venus) for downstream activity. |
| Caspase-9 Inhibitors [1] [8] | To probe the functional role of caspase-9 in apoptotic pathways. | Z-LEHD-FMK: Irreversible peptide inhibitor. XIAP Bir3 Domain: Endogenous, highly selective inhibitor of the D315 cleaved form of caspase-9. |
| Genetic Models [1] | To determine physiological roles and non-apoptotic functions. | Caspase-9 KO Mice: Perinatal lethality, brain malformations. Caspase-9 null cells: Resistant to intrinsic apoptotic stimuli (UV, γ-irradiation, chemotherapeutics). |
| Phospho-specific Antibodies [1] | To study post-translational regulation of caspase-9. | Antibodies targeting inhibitory phosphorylation sites (e.g., Thr125), which is phosphorylated by ERK, CDK1, among others. |
Synthesis: The Emerging Hybrid Model and Its Implications
The accumulated experimental evidence strongly argues against a mutually exclusive interpretation of caspase-9 activation. Instead, the data converge on a hybrid model where both proximity-induced dimerization and conformation-inducing mechanisms are integrated. In this framework, the apoptosome's primary function is to serve as an allosteric activation platform, not merely a passive tethering device [1] [4]. It facilitates the initial dimerization of caspase-9 (proximity), but also, through specific multi-point CARD interactions and nucleotide-dependent structural changes, locks the caspase-9 dimer into a fully active conformation that cannot be achieved by dimerization alone [1] [69].
This integrated model has profound implications for drug development. It suggests that therapeutic strategies aimed at modulating apoptosis must consider both the assembly of the apoptosome and the subsequent allosteric activation of caspase-9. For instance, in cancers with defective apoptosis, agents that stabilize the active conformation of caspase-9 on the apoptosome could overcome resistance [1]. Conversely, in degenerative diseases, inhibitors targeting the specific interaction interfaces between Apaf-1 and caspase-9 might offer more precise control than general caspase inhibitors. The heterogeneity in cellular responses, driven by variable expression of caspase-9, XIAP, and other regulators, further underscores the need for a nuanced understanding of this hybrid mechanism to develop effective, personalized therapeutic interventions [8] [70].
Caspase-9 is universally recognized as the pivotal initiator caspase of the intrinsic apoptotic pathway, activated through the formation of the apoptosome complex in response to cellular stress. However, a growing body of evidence reveals that caspase-9's biological functions extend far beyond its classical role in cell death. This guide examines the non-apoptotic roles of caspase-9, focusing on its cross-talk with vital cellular processes including autophagy, differentiation, and fibrotic signaling, while framing these findings within the ongoing scientific debate regarding its activation mechanism: induced proximity versus induced conformation.
The Caspase-9 Activation Debate: A Theoretical Framework
The mechanism of caspase-9 activation is fundamental to understanding how it can participate in non-apoptotic processes without triggering full-blown apoptosis. Two primary models dominate this discourse, each supported by distinct experimental evidence.
- Table 1: Key Models of Caspase-9 Activation
| Model Name | Core Principle | Supporting Evidence | Implications for Non-Apoptotic Roles |
|---|---|---|---|
| Induced Proximity [1] [3] | The apoptosome serves as a platform to bring caspase-9 zymogens into close proximity, promoting homodimerization and auto-activation. | Engineered dimeric caspase-9 shows enhanced activity, though less than Apaf-1-activated wild-type [3]. | Suggests localized, transient dimerization could permit limited substrate cleavage without full apoptosis. |
| Induced Conformation [1] [3] | Binding to the apoptosome induces a specific conformational change in caspase-9 that drives its activation, beyond mere dimerization. | The crystal structure of an Apaf-1/caspase-9 CARD complex shows a specific complementary interface crucial for activation [1]. | implies that caspase-9's function could be modulated by protein interactions that alter its structure in different cellular contexts. |
The two models are not necessarily mutually exclusive, and the precise mechanism may involve elements of both [3]. This nuanced control of caspase-9 activity is a prerequisite for its involvement in the finely-tuned, non-lethal processes described below.
Non-Apoptotic Functions of Caspase-9: Mechanisms and Experimental Evidence
Caspase-9 engages in a diverse array of non-apoptotic processes, from cellular remodeling to signal transduction. The following table summarizes key functions, their mechanisms, and experimental approaches used to validate them.
- Table 2: Non-Apoptotic Roles of Caspase-9
| Functional Role | Mechanism / Pathway | Key Experimental Findings |
|---|---|---|
| Autophagy Regulation [71] | Forms a complex with Atg7, enhancing LC3-II formation and autophagosome biogenesis. Atg7 binding concurrently represses caspase-9's apoptotic activity. | siRNA knockdown of caspase-9 impairs autophagosome formation. The Atg7-caspase-9 interaction was confirmed via co-immunoprecipitation. |
| Cell Differentiation [1] [72] | Promotes skeletal myoblast differentiation; essential for spermatid individualization in Drosophila via localized, sub-lethal activation. | Caspase-9 knockdown inhibits bovine skeletal muscle differentiation [1]. Effector caspase activity is required for organelle removal in spermatids [72]. |
| Fibrosis Promotion [7] | Drives pulmonary fibrosis by inducing alveolar epithelial cell apoptosis and directly interacting with β-catenin to enhance its nuclear translocation and pro-fibrotic signaling. | Caspase-9 is upregulated in human PF lungs. Its inhibition reduces collagen in mouse models. β-catenin silencing reverses Caspase-9-induced fibrosis [7]. |
The diagram below illustrates the core signaling pathways through which caspase-9 exerts its non-apoptotic functions, highlighting its central role in cross-talk between different cellular processes.
Experimental Protocols for Key Findings
To facilitate replication and further research, here are the detailed methodologies for key experiments elucidating caspase-9's non-apoptotic roles.
Investigating the Caspase-9/Atg7 Complex in Autophagy
This protocol is adapted from research demonstrating the direct interaction between caspase-9 and the autophagy protein Atg7 [71].
- Objective: To confirm the formation of a protein complex between caspase-9 and Atg7 and assess its functional impact on autophagy and apoptosis.
- Key Reagents: Caspase-9 and Atg7 antibodies, Lipofectamine 2000, control and specific siRNAs against caspase-9, LC3-II antibody (autophagy marker), cell culture reagents.
- Procedure:
- Cell Culture and Transfection: Culture relevant cell lines (e.g., Hct116, MEFs). Transfect cells with caspase-9-specific siRNA or a non-targeting control siRNA using Lipofectamine 2000.
- Co-Immunoprecipitation (Co-IP): After 48 hours, lyse the cells. Incubate the cell lysates with an antibody against caspase-9 or a control IgG. Precipitate the immune complexes using Protein A/G beads.
- Immunoblotting: Resolve the precipitated proteins and total cell lysates by SDS-PAGE. Transfer to a membrane and perform Western blotting using antibodies against Atg7 and caspase-9 to detect interaction.
- Functional Autophagy Assay: In parallel, treat siRNA-transfected cells with autophagy inducers (e.g., rapamycin) or inhibitors. Analyze the conversion of LC3-I to the lipidated LC3-II form by Western blot as a measure of autophagic flux.
- Apoptosis Assay: Challenge the cells with an apoptotic stimulus (e.g., etoposide). Measure caspase-9 and caspase-3 activity using fluorogenic substrates or cleaved-form specific antibodies.
Delineating the Caspase-9/β-catenin Axis in Pulmonary Fibrosis
This method outlines the approach used to establish caspase-9's role in promoting fibrosis through β-catenin signaling [7].
- Objective: To determine if caspase-9 drives pulmonary fibrosis by activating β-catenin pro-fibrotic signaling.
- Key Reagents: Bleomycin, Caspase-9 inhibitor (Z-LEHD-FMK), β-catenin agonist (WAY-262,611), antibodies for Caspase-9, cleaved-Caspase-9, β-catenin, Collagen I, α-SMA, siRNA/plasmids for Caspase-9 and β-catenin.
- In Vivo Model (Mouse):
- Induce pulmonary fibrosis in C57BL/6 mice via intratracheal instillation of bleomycin (5 mg/kg).
- Treat experimental groups with either a caspase-9 inhibitor (Z-LEHD-FMK, 10 mg/kg, i.p.), a β-catenin agonist, or a negative control for 21 days.
- Harvest lung tissues for analysis. Assess fibrosis by Masson's trichrome (collagen deposition) and H&E staining (histology). Analyze protein expression of pro-fibrotic markers (Collagen I, α-SMA) and pathway components (caspase-9, β-catenin) by Western blot and immunohistochemistry.
- In Vitro Model (MLE-12 Cells):
- Culture mouse alveolar epithelial cells (MLE-12) and treat with TGF-β1 (10 ng/mL) to mimic fibrotic stimulation.
- Perform genetic manipulations: knockdown caspase-9 or β-catenin using siRNA; overexpress caspase-9 or a constitutively active β-catenin using plasmids.
- Measure outcomes: Apoptosis (TUNEL assay, cleaved caspase-3), epithelial-mesenchymal transition marker E-cadherin, and pro-fibrotic gene expression. Use co-immunoprecipitation and immunofluorescence to assess caspase-9/β-catenin interaction and β-catenin nuclear localization.
The Scientist's Toolkit: Essential Research Reagents
This table catalogs critical reagents for investigating the non-apoptotic roles of caspase-9, based on the experimental data cited.
- Table 3: Key Reagents for Caspase-9 Research
| Reagent Category | Specific Example | Function / Application |
|---|---|---|
| Pharmacological Inhibitors | Z-LEHD-FMK (Caspase-9 inhibitor) [7] | Selectively inhibits caspase-9 activity; used to probe its functional role in vivo and in vitro. |
| Genetic Tools | Caspase-9 specific siRNA/shRNA [71] [7] | Knocks down gene expression to study loss-of-function phenotypes. |
| Caspase-9 overexpression plasmids [7] | Enables study of gain-of-function and signaling activation. | |
| Caspase-9 knockout cells (e.g., MEFs) [71] | Provides a clean genetic background to validate specificity and for reconstitution experiments. | |
| Antibodies for Detection | Anti-Caspase-9 (full length) [71] [7] | Detects total protein levels and used for immunoprecipitation. |
| Anti-cleaved-Caspase-9 (Asp315) [7] | Specific marker for activated caspase-9. | |
| Anti-β-catenin [7] | Assesses interaction partner and nuclear translocation in fibrotic signaling. | |
| Anti-LC3-II [71] | Key marker for monitoring autophagosome formation and autophagic flux. | |
| Activity Assays | Fluorogenic caspase-9 substrates [73] | Directly measures enzymatic activity in cell lysates. |
| Animal Disease Models | Bleomycin-induced pulmonary fibrosis model [7] | Standard in vivo model for testing the role of caspase-9 in a fibrotic disease context. |
Caspase-9 is a multifunctional protease whose biological influence is not confined to apoptosis. It is a critical node of cross-talk, regulating autophagy, cellular differentiation, and disease processes like fibrosis. The precise, context-dependent regulation of its activity—whether explained by induced proximity, induced conformation, or a synthesis of both models—allows it to perform these non-apoptotic roles without triggering cell death. Understanding these pathways and the experimental tools available to probe them provides a solid foundation for developing novel therapeutic strategies that target the non-apoptotic functions of caspase-9 in cancer, fibrotic diseases, and beyond.
Caspases, a family of cysteine-aspartic proteases, are fundamental regulators of programmed cell death (PCD). Initiator caspases, which include caspase-2, -8, -9, and -10, stand at the apex of apoptotic signaling pathways and are characterized by their activation within large multi-protein complexes [2] [51]. For years, the induced proximity model has served as the dominant paradigm for explaining initiator caspase activation, proposing that dimerization driven by proximity is the central mechanism [12] [4]. However, emerging research reveals that caspase-9 activation is a notable exception, governed by a more complex mechanism that may be better explained by an induced conformation model [3] [4]. This guide provides a objective comparison of caspase-9 activation against other initiator caspases, synthesizing current structural, biochemical, and functional data to highlight these critical distinctions for researchers and drug development professionals.
Molecular Mechanisms of Initiator Caspase Activation
The Prevailing Model: Induced Proximity
The induced proximity model, initially proposed by Salvesen and colleagues, posits that initiator caspases are auto-activated when they are brought into close proximity with one another within activating protein complexes [12]. This clustering increases their local concentration, facilitating homodimerization—the formation of a complex containing two caspase molecules—which in turn triggers their catalytic activity [3]. This model is largely accepted for several initiator caspases, including caspase-8.
- Caspase-8:
- Activating Platform: The Death-Inducing Signaling Complex (DISC), formed by membrane-bound death receptors [62].
- Activation Mechanism: Within the DISC, the adapter protein FADD (Fas-associated death domain) recruits procaspase-8, leading to its dimerization and subsequent auto-activation through intermolecular processing [2] [62]. This process is a clear example of proximity-induced dimerization.
The Caspase-9 Exception: Induced Conformation
In contrast to caspase-8, activation of caspase-9, the initiator caspase of the intrinsic (mitochondrial) apoptotic pathway, cannot be fully explained by dimerization alone [3] [4].
- Activating Platform: The apoptosome, a heptameric complex composed of Apaf-1 (apoptotic protease activating factor-1) and cytochrome c [3] [62].
- Activation Mechanism: While the apoptosome does bring caspase-9 molecules into proximity, engineered dimeric forms of caspase-9 exhibit only a fraction of the catalytic activity of the Apaf-1-activated wild-type enzyme [4]. Furthermore, the activity of these engineered dimers cannot be significantly enhanced by Apaf-1 [4]. This evidence suggests that the apoptosome does not merely serve as a dimerization platform but induces a specific conformational change in caspase-9 that is essential for its full activation [3] [4].
The diagram below illustrates and contrasts the fundamental activation mechanisms for caspase-8 and caspase-9.
Comparative Analysis of Initiator Caspases
The table below provides a detailed, side-by-side comparison of the key regulatory and functional characteristics of major initiator caspases.
Table 1: Comparative Profile of Initiator Caspases
| Feature | Caspase-9 | Caspase-8 | Caspase-2 | Caspase-10 |
|---|---|---|---|---|
| Primary Pathway | Intrinsic Apoptosis [2] | Extrinsic Apoptosis [2] | Intrinsic Apoptosis / DNA Damage [2] | Extrinsic Apoptosis [2] |
| Activating Platform | Apoptosome (Apaf-1/cytochrome c) [3] [62] | DISC (Death Receptors, FADD) [2] [62] | PIDDosome (PIDD, RAIDD) [2] | DISC (Death Receptors, FADD) [2] |
| Pro-Domain | CARD [2] [51] | DED [2] [51] | CARD [2] | DED [2] |
| Activation Model | Induced Conformation [3] [4] | Induced Proximity (Dimerization) [62] [12] | Induced Proximity (presumed) | Induced Proximity (presumed) |
| Key Regulatory Role | Initiates mitochondrial pathway; cleaves caspases-3, -7 [2] [8] | Molecular switch between apoptosis, necroptosis, & pyroptosis [2] | DNA damage response; cell cycle control [2] | Involved in extrinsic apoptosis; may regulate caspase-8 [2] |
| Non-Apoptotic Functions | Mitochondrial homeostasis, corticospinal circuit remodeling, endothelial dysfunction [8] | Regulation of necroptosis and pyroptosis [2] | Inhibition of ferroptosis [2] | Implicated in pyroptosis and necroptosis [2] |
Key Experimental Evidence and Data
The distinct activation mechanism of caspase-9 is supported by robust biochemical and structural evidence that directly challenges the simple induced proximity model.
The Engineered Dimer Experiment
A seminal study by Shi and colleagues provided critical evidence for the induced conformation model. Researchers engineered a constitutively dimeric form of caspase-9 by mutating residues at its dimer interface to relieve steric hindrance [4].
Table 2: Activity Comparison of Wild-Type vs. Engineered Caspase-9
| Caspase-9 Form | Catalytic Activity | Response to Apaf-1 | Cell Death Induction |
|---|---|---|---|
| Wild-Type Monomer | Low basal activity [4] | Significantly enhanced [4] | Baseline level [4] |
| Engineered Dimer | Higher than monomer, but only a small fraction of Apaf-1-activated WT [3] [4] | Not significantly enhanced [4] | More efficient than WT monomer [4] |
| Apaf-1-Activated WT | High catalytic activity [4] | N/A | High efficiency [3] |
Experimental Protocol [4]:
- Protein Engineering: Site-directed mutagenesis was used to create a caspase-9 variant (F404E) designed to form a stable dimer by replacing a phenylalanine residue that causes steric clash at the dimer interface.
- Structural Validation: The crystal structure of the engineered dimer was solved and shown to closely resemble the wild-type caspase-9 dimer, confirming that the mutations did not cause major structural perturbations.
- Biochemical Assays: The enzymatic activity of the engineered dimer, wild-type monomer, and Apaf-1-activated wild-type caspase-9 was measured using a fluorogenic substrate (LEHD-AFC).
- Functional Cell-based Assay: The ability of each caspase-9 form to induce apoptosis was quantified upon expression in cells.
Interpretation: The fact that a stable, structurally sound dimer cannot replicate the full activity achieved on the apoptosome strongly indicates that Apaf-1 binding induces a specific, activating conformation in caspase-9 that goes beyond mere dimerization [3] [4].
DNA Origami Platform for Proximity Analysis
A modern, synthetic biology approach using DNA origami has further refined our understanding of caspase-9 activation. This method allows for the precise nanoscale organization of proteins to systematically study proximity effects [28].
Experimental Protocol [28]:
- Conjugate Synthesis: Caspase-9 catalytic domains were site-specifically conjugated to oligonucleotides via a non-canonical amino acid.
- Platform Assembly: DNA origami tiles were designed with protruding single-stranded DNA "handles" to hybridize and position caspase-9 conjugates at defined distances and geometries.
- Activity Measurement: The proteolytic activity of the scaffolded caspase-9 assemblies (dimers, trimers, tetramers) was measured and compared to free conjugates.
Key Findings: This study confirmed that proximity-induced dimerization is sufficient to trigger caspase-9 activity, supporting the proximity aspect of the model. However, it also revealed that oligomers of three and four enzymes exhibited enhanced activity, suggesting a multivalent effect that may be related to the heptameric nature of the native apoptosome [28]. This provides a potential bridge between the proximity and conformation models.
The Scientist's Toolkit: Key Research Reagents
The following table lists essential reagents and tools used in the featured experiments for studying caspase activation.
Table 3: Essential Research Reagents for Caspase Activation Studies
| Reagent / Tool | Function & Application | Key Feature |
|---|---|---|
| Fluorogenic Substrates (e.g., LEHD-AFC) | Quantitative measurement of caspase enzymatic activity. The substrate emits fluorescence upon cleavage [28]. | Allows real-time kinetic analysis of caspase activation in vitro. |
| Engineered Dimeric Caspase-9 (F404E) | A tool to directly test the role of dimerization independent of the apoptosome [4]. | Constitutively dimeric but lacks full activity of apoptosome-bound caspase-9. |
| DNA Origami Scaffold | A synthetic platform for precise spatial organization of caspase-9 monomers to study proximity effects [28]. | Programmable control over the number, position, and orientation of enzymes. |
| Caspase-9 Inhibitor Z-LEHD-FMK | A cell-permeable pharmacological inhibitor that selectively targets caspase-9 [17]. | Used in functional studies to probe the role of caspase-9 in cellular models of disease. |
| b-VAD-fmk (Biotinylated) | An irreversible, pan-caspase inhibitor that binds active caspases; allows affinity labeling and pull-down of active initiator caspases from cell lysates [73]. | Useful for identifying which initiator caspases are activated in response to specific apoptotic stimuli. |
The activation of initiator caspases is a critical control point in programmed cell death. While caspase-8 and -10 conform to the induced proximity model through dimerization in the DISC, caspase-9 operates under a distinct, more complex paradigm. Compelling evidence from engineered proteins and synthetic biology platforms demonstrates that its activation within the apoptosome requires an induced conformational change that is qualitatively different from and functionally superior to simple dimerization [3] [4] [28]. This fundamental mechanistic difference has profound implications for drug discovery, suggesting that therapeutic strategies targeting caspase-9 must aim to modulate its conformation within the apoptosome, rather than merely disrupting dimerization. A deep understanding of these comparative regulatory mechanisms is essential for developing precise interventions in cancer, neurodegenerative disorders, and other diseases characterized by dysregulated cell death.
Caspase-9, a pivotal initiator caspase in the intrinsic apoptotic pathway, functions as a critical signaling hub whose dysregulation contributes to numerous human diseases. Its activation occurs through a sophisticated molecular mechanism centered on the apoptosome, a heptameric complex composed of Apaf-1 and cytochrome c that forms in response to cellular stress [74]. For decades, the induced proximity model served as the prevailing explanation for caspase-9 activation, proposing that apoptosome assembly simply brings caspase-9 molecules into close proximity, facilitating homodimerization and subsequent autoactivation [3]. However, emerging research led by Yigong Shi and colleagues has challenged this paradigm, demonstrating through engineered dimeric caspase-9 that dimerization alone yields only fractional activity compared to Apaf-1-activated wild-type enzyme [3]. This evidence supports an alternative induced conformation model, wherein caspase-9 undergoes precise structural changes upon binding to the apoptosome that drive its activation [3]. These competing models—induced proximity versus induced conformation—provide essential frameworks for understanding how caspase-9 mutations disrupt normal function and contribute to pathological states including cancer, neurodegeneration, and fibrotic disease, while also illuminating potential mechanisms of therapeutic resistance.
Molecular Mechanisms: From Biochemical Models to Pathological Consequences
The Apoptosome Complex: Structural Foundations for Disease
The apoptosome serves as the central activation platform for caspase-9, forming only after mitochondrial outer membrane permeabilization releases cytochrome c into the cytosol [74]. This critical complex assembly represents the point where numerous cellular stress signals converge to initiate the intrinsic apoptosis pathway. The precise mechanism by which the apoptosome activates caspase-9 has profound implications for understanding disease pathogenesis. The induced proximity model emphasizes concentration-driven dimerization as the primary activation mechanism [12], while the induced conformation model highlights essential structural rearrangements in caspase-9 upon Apaf-1 binding [3]. Current evidence suggests these models are not mutually exclusive, with both proximity and conformational changes likely contributing to full caspase-9 activation [3]. This molecular understanding provides critical insights into how specific mutations in either caspase-9 or Apaf-1 might disrupt these precise activation mechanisms, potentially leading to pathological outcomes.
Caspase-9 in Cellular Homeostasis and Disease Pathogenesis
Beyond its established role in apoptosis, caspase-9 participates in diverse physiological processes through both catalytic and non-catalytic mechanisms, with dysregulation contributing to various disease states:
- Non-apoptotic Functions: Caspase-9 regulates cellular differentiation, innate immunity, mitochondrial homeostasis, and autophagy [74]. It mediates corticospinal circuit reorganization essential for skilled movement and cleaves semaphorin7a for proper axonal development [74] [8].
- Disease Associations: CASP9 polymorphisms link to cancers, neurological disorders, autoimmune pathologies, and lumbar disc disease [74]. Altered expression or activity occurs in neurodegeneration, retinal neuropathy, cardiomyopathies, and atherosclerosis [74] [8].
- Cell-Type Specific Effects: In ischemic injury, caspase-9 promotes neuronal apoptosis while driving non-apoptotic endothelial dysfunction contributing to capillary nonperfusion [74]. This functional diversity highlights the complex therapeutic landscape for caspase-9 modulation.
Table 1: Caspase-9 Activation Models and Their Pathological Implications
| Activation Model | Core Principle | Key Experimental Evidence | Disease Implications |
|---|---|---|---|
| Induced Proximity | Apoptosome increases local caspase-9 concentration, promoting homodimerization [3] | Engineered dimeric caspase-9 shows limited activity compared to Apaf-1-activated enzyme [3] | Mutations disrupting dimerization interfaces may cause immunodeficiencies |
| Induced Conformation | Apaf-1 binding induces structural changes that drive caspase-9 activation [3] | Dimeric caspase-9 shows no stimulation by Apaf-1, suggesting qualitative differences [3] | Conformational mutations may yield dysfunctional enzyme in neurodegeneration |
| Hybrid Model | Both proximity and conformational changes contribute to activation [3] | Crystal structure shows engineered caspase-9 resembles wild-type despite functional differences [3] | Compound mutations may cause more severe therapeutic resistance |
Disease Correlates: Activation Mechanisms in Human Pathology
Cancer and Therapy Resistance
In oncology, caspase-9 dysfunction represents a critical mechanism of tumor progression and therapeutic resistance. Alternative splicing generates caspase-9b, a dominant-negative isoform that competes with full-length caspase-9 for apoptosome binding, effectively inhibiting apoptosis [74]. This isoform is frequently upregulated in cancers, providing a mechanism for apoptosis evasion [74]. Additionally, caspase-9 mutations that favor the induced proximity model without proper conformational activation may permit sufficient survival signaling to resist chemotherapeutic agents. The development of therapeutic resistance often involves selection for caspase-9 variants with altered activation thresholds, particularly mutations at the Apaf-1 interaction interface that disrupt induced conformation mechanisms while preserving limited dimerization capability.
Neurodegenerative Disorders
In neurological contexts, caspase-9 activation features prominently in both acute and chronic neurodegeneration. In ischemic stroke, caspase-9 directly cleaves and activates caspase-6, driving axonal degeneration [74]. Beyond apoptotic roles, non-apoptotic caspase-9 activity regulates corticospinal circuit refinement, with deficient activation causing skilled movement deficits [74]. In retinal vein occlusion, caspase-9 inhibition confers stronger neuroprotection than VEGF neutralization, preserving electrophysiological function and reducing atrophy [75]. Neurodegenerative conditions like Alzheimer's and Parkinson's diseases may involve caspase-9 variants with altered activation kinetics, where mutations favoring premature proximity-induced activation without proper regulatory constraints could drive inappropriate neuronal loss.
Pulmonary and Fibrotic Diseases
Recent research has identified caspase-9 as a driver of pulmonary fibrosis through both apoptotic and non-apoptotic mechanisms. In fibrotic lung tissues, caspase-9 and cleaved-caspase-9 are significantly upregulated [17]. Mechanistically, caspase-9 interacts with β-catenin, enhancing its nuclear accumulation and promoting pro-fibrotic signaling [17]. Inhibition of caspase-9 reduces collagen deposition and improves lung architecture in bleomycin-induced pulmonary fibrosis models [17]. This caspase-9/β-catenin axis represents a novel signaling pathway in fibrotic pathogenesis, suggesting that conformational activation of caspase-9 may initiate non-apoptotic transcriptional programs contributing to tissue remodeling.
Table 2: Caspase-9 in Human Diseases: Genetic and Therapeutic Correlates
| Disease Category | Specific Conditions | Caspase-9 Alterations | Therapeutic Implications |
|---|---|---|---|
| Neurological Disorders | Ischemic stroke, Alzheimer's disease, Retinal vein occlusion [74] [75] | Increased activation, Altered cleavage patterns, Non-apoptotic signaling [74] | Caspase-9 inhibition shows superior neuroprotection to VEGF neutralization in RVO [75] |
| Fibrotic Diseases | Pulmonary fibrosis, Renal fibrosis, Hepatic fibrosis [17] | Upregulation in fibrotic tissues, Interaction with β-catenin [17] | Caspase-9 inhibition reduces collagen deposition; β-catenin activation reverses protection [17] |
| Cancer | Various solid and hematologic malignancies [74] | Caspase-9b isoform overexpression, Inactivating mutations [74] | Selective caspase-9 activation could bypass resistance mechanisms |
| Infectious Disease | Neurocysticercosis [76] | Parasite-induced caspase-9-mediated apoptosis in immune cells [76] | Caspase-9 inhibition may reduce inflammation-mediated pathology |
Experimental Analysis: Methodologies for Activation Mechanism Studies
Key Experimental Approaches
Understanding caspase-9 activation mechanisms requires sophisticated methodological approaches that can distinguish between proximity and conformational effects:
- Engineered Dimerization Studies: Researchers have created constitutively dimeric caspase-9 mutants to test induced proximity independently of apoptosome assembly. These studies revealed that dimerization produces only partial activity compared to Apaf-1-mediated activation [3], suggesting additional conformational requirements.
- Structural Biology Techniques: X-ray crystallography and cryo-electron microscopy provide high-resolution structural data comparing Apaf-1-bound and unbound caspase-9. These approaches reveal precise conformational changes during activation [3].
- Biochemical Reconstitution: Combining purified caspase-9 with assembled apoptosomes in cell-free systems allows precise dissection of activation kinetics and cleavage preferences, including analysis of neoepitope formation at D315 (autocleavage) versus D330 (caspase-3 cleavage) sites [74].
- Cellular Models of Disease: TGF-β1-treated alveolar epithelial cells (MLE-12) model pulmonary fibrosis, enabling study of caspase-9/β-catenin interactions [17], while retinal vein occlusion models reveal vascular and neuronal protection with caspase-9 inhibition [75].
Signaling Pathway Integration
The following diagram illustrates caspase-9 activation pathways and their connections to disease processes:
Research Reagent Solutions for Caspase-9 Investigation
Table 3: Essential Research Reagents for Caspase-9 Activation Studies
| Reagent / Tool | Type | Primary Research Application | Key Features & Limitations |
|---|---|---|---|
| Pen1-XBir3 | Cell-penetrating peptide inhibitor [75] | Selective caspase-9 inhibition in cellular and animal models | Comprises XIAP Bir3 domain linked to Penetratin-1; topical application effective in retinal models [75] |
| Z-LEHD-FMK | Pharmacologic caspase-9 inhibitor [17] | In vitro and in vivo inhibition studies | Irreversible inhibitor used in pulmonary fibrosis models (10 mg/kg in mice) [17] |
| Caspase-9b | Endogenous dominant-negative isoform [74] | Apoptosis resistance mechanisms | Naturally occurring inhibitor; competes for apoptosome binding; upregulated in cancers [74] |
| Engineered dimeric caspase-9 | Genetically modified caspase-9 [3] | Mechanism dissection | Tests induced proximity independently; shows partial activation without Apaf-1 [3] |
| Neoepitope-specific antibodies (D315/D330) | Cleavage-site specific antibodies [74] | Pathway activation analysis | Distinguish autocleavage (D315) from caspase-3 cleavage (D330); differential XIAP sensitivity [74] |
| Q-VD-OPh | Broad-spectrum caspase inhibitor [6] | Pan-caspase control studies | Improved permeability and reduced toxicity compared to Z-VAD-FMK [6] |
Therapeutic Implications: Targeting Activation Mechanisms
Caspase-9 Inhibition Strategies
Therapeutic targeting of caspase-9 has evolved significantly with improved understanding of its activation mechanisms:
- Selective Inhibition Approaches: The Bir3 domain of XIAP represents a natural highly selective caspase-9 inhibitor [74], which has been engineered into cell-penetrating formats like Pen1-XBir3 for topical application in retinal diseases [75]. In retinal vein occlusion models, this approach demonstrated superior protection against neurodegeneration and vascular dysfunction compared to VEGF neutralization [75].
- Pharmacological Challenges: Development of clinical caspase inhibitors has faced significant hurdles, with molecules like emricasan (IDN-6556), pralnacasan (VX-740), and belnacasan (VX-765) progressing to clinical trials but ultimately failing due to inadequate efficacy, toxicity concerns, or poor target specificity [6]. These challenges highlight the complexity of therapeutic caspase modulation in human disease.
- Alternative Therapeutic Strategies: Dominant-negative caspase-9 mutants represent a genetic inhibition approach that competes with endogenous caspase-9 for apoptosome binding [74]. Additionally, targeting regulatory phosphorylation sites or upstream pathway components may provide indirect methods for modulating caspase-9 activity with potentially improved therapeutic windows.
Activation Mechanism-Specific Therapeutics
The distinction between induced proximity and induced conformation models informs therapeutic design:
- Conformation-Targeting Approaches: Small molecules that stabilize caspase-9 in inactive conformations could selectively inhibit pathological activation while preserving physiological function. Conversely, compounds promoting activation conformations might overcome resistance in cancer contexts.
- Dimerization Interface Modulators: Molecules targeting the caspase-9 dimerization interface could specifically disrupt proximity-mediated activation, potentially addressing diseases where this mechanism predominates.
- Apoptosome-Directed Therapeutics: Interventions targeting Apaf-1 or apoptosome assembly could provide upstream control over caspase-9 activation, potentially with greater specificity than direct caspase inhibition.
The clinical correlates of caspase-9 activation mechanisms reveal a complex landscape where fundamental biochemical principles directly impact disease pathogenesis and therapeutic development. The ongoing evolution from simple induced proximity models toward more nuanced understanding incorporating induced conformational changes provides essential framework for connecting caspase-9 genetics and mutations to diverse pathological states. Future research directions should prioritize high-resolution structural studies of full apoptosome complexes, development of activation mechanism-specific chemical probes, and clinical validation of caspase-9 biomarkers for patient stratification. The integration of caspase-9's apoptotic functions with its emerging roles in cellular differentiation, mitochondrial homeostasis, and non-apoptotic signaling presents both challenges and opportunities for therapeutic innovation. As our understanding of caspase-9 activation mechanisms continues to mature, so too will our ability to target this critical protease with precision and efficacy across the spectrum of human disease.
Conclusion
The long-standing debate between the induced proximity and induced conformation models for caspase-9 activation has evolved from a dichotomy into a more integrated understanding. Current evidence strongly suggests that while dimerization is a necessary component, it is not sufficient for full catalytic activity. The apoptosome does not function as a passive platform but as an active allosteric regulator that induces specific conformational changes essential for robust caspase-9 activation. This refined hybrid model, supported by engineered dimer experiments and structural studies, has profound implications. For therapeutic development, it shifts the focus from merely promoting dimerization to targeting specific protein-protein interactions within the apoptosome. Future research, particularly high-resolution structural analysis of the full apoptosome complex, is critical to fully visualize the allosteric network. Understanding the precise molecular switch of caspase-9 activation opens new avenues for designing targeted therapies in cancer, neurodegenerative disorders, and other diseases characterized by dysregulated apoptosis.
References
- 1. Caspase-9: structure, mechanisms and clinical application [https://pmc.ncbi.nlm.nih.gov/articles/PMC5410359/]
- 2. Caspases as master regulators of programmed cell death [https://www.nature.com/articles/s12276-025-01470-9]
- 3. Proximity- or Conformation-Induced Caspase Activation? [https://pmc.ncbi.nlm.nih.gov/articles/PMC1088975/]
- 4. Engineering a Dimeric Caspase-9: A Re-evaluation of the ... [https://journals.plos.org/plosbiology/article?id=10.1371/journal.pbio.0030183]
- 5. Activation of caspase-9 on the apoptosome as studied by ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10743466/]
- 6. A long way to go: caspase inhibitors in clinical use [https://www.nature.com/articles/s41419-021-04240-3]
- 7. Caspase-9 activates β-catenin signaling to promote ... [https://translational-medicine.biomedcentral.com/articles/10.1186/s12967-025-07020-1]
- 8. Caspase-9: A Multimodal Therapeutic Target With Diverse ... [https://www.frontiersin.org/journals/pharmacology/articles/10.3389/fphar.2021.701301/full]
- 9. Caspase-9 inhibition triggers Hsp90-based chemotherapy ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10711858/]
- 10. Atomic structure of the apoptosome: mechanism of cytochrome c [https://pmc.ncbi.nlm.nih.gov/articles/PMC4691890/]
- 11. A near atomic structure of the active human apoptosome - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC5050015/]
- 12. Caspase Activation: Revisiting the Induced Proximity Model [https://www.sciencedirect.com/science/article/pii/S0092867404005756]
- 13. Apoptosome: a platform for the activation of initiator caspases [https://www.nature.com/articles/4402028]
- 14. Caspase-9 and APAF-1 form an active holoenzyme [https://genesdev.cshlp.org/content/13/24/3179.short]
- 15. CASPASE-9 CARD:CORE DOMAIN INTERACTIONS ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC5878146/]
- 16. Structure of an apoptosome-procaspase-9 CARD complex [https://pmc.ncbi.nlm.nih.gov/articles/PMC2874686/]
- 17. Caspase-9 activates β-catenin signaling to promote ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12406446/]
- 18. Implications for Assembly, Procaspase-9 Binding, and ... [https://www.sciencedirect.com/science/article/pii/S1097276502004422]
- 19. Caspase activation: The induced-proximity model [https://pmc.ncbi.nlm.nih.gov/articles/PMC34227/]
- 20. Engineering a dimeric caspase-9: a re-evaluation of ... - PubMed [https://pubmed.ncbi.nlm.nih.gov/15941357/]
- 21. Proximity-Based Modalities for Biology and Medicine - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC10375889/]
- 22. Induced proximity pushes beyond protein degraders, as ... [https://www.nature.com/articles/d41573-025-00037-7]
- 23. The Apaf-1 apoptosome induces formation of caspase-9 ... [https://www.nature.com/articles/ncomms13565]
- 24. Caspase-9 and APAF-1 form an active holoenzyme - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC317200/]
- 25. The Apoptosome Activates Caspase-9 by Dimerization [https://www.sciencedirect.com/science/article/pii/S1097276506001730]
- 26. Molecular determinants of caspase-9 activation by the Apaf-1 ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC4246342/]
- 27. Caspase activation: revisiting the induced proximity model [https://pubmed.ncbi.nlm.nih.gov/15210107/]
- 28. Proximity-induced caspase-9 activation on a DNA origami- ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC7080557/]
- 29. How Caspase 9 Antibody Works — In One Simple Flow ... [https://www.linkedin.com/pulse/how-caspase-9-antibody-works-one-simple-flow-2025-z32ef/]
- 30. Engineering a Dimeric Caspase-9: A Re-evaluation of the ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC1088972/]
- 31. Caspase-9: structure, mechanisms and clinical application [https://www.oncotarget.com/article/15098/text/]
- 32. Dimer formation drives the activation of the cell death protease ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC64668/]
- 33. Investigation of metallodrug/protein interaction by X-ray ... [https://pubs.rsc.org/en/content/articlehtml/2025/qi/d4qi03277b]
- 34. An NMR sample preparation case study: Considerations for the self-destructive protease caspase-6 | PLOS One [https://journals.plos.org/plosone/article?id=10.1371/journal.pone.0337291]
- 35. Crystal Structure of the Nod1 Caspase Activation and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC1821002/]
- 36. Proximity-induced caspase-9 activation on a DNA origami- ... [https://pubmed.ncbi.nlm.nih.gov/32190819/]
- 37. A modular DNA origami nanocompartment for engineering ... [https://www.nature.com/articles/s41565-024-01738-7]
- 38. DNA Nanodevices as Mechanical Probes of Protein ... [https://www.mdpi.com/2076-3417/11/6/2802]
- 39. New insights into apoptosome structure and function - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC6030056/]
- 40. Apoptosome Structure, Assembly, and Procaspase Activation [https://www.sciencedirect.com/science/article/pii/S096921261300083X]
- 41. A Systems Biology Analysis of Apoptosome Formation and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC4176236/]
- 42. A Systems Biology Analysis of Apoptosome Formation and ... [https://www.sciencedirect.com/science/article/pii/S0021925820484583]
- 43. Analysis of the composition, assembly kinetics and activity ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC424369/]
- 44. Learning from Deficiency: Gene Targeting of Caspases - Madame Curie Bioscience Database - NCBI Bookshelf [https://www.ncbi.nlm.nih.gov/books/NBK6153/]
- 45. A Whole Cell Assay to Measure Caspase-6 Activity by Detecting... [https://pmc.ncbi.nlm.nih.gov/articles/PMC3257251/]
- 46. Mapping specificity , cleavage entropy... | Nature Communications [https://www.nature.com/articles/s41467-021-21754-8?error=cookies_not_supported&code=4637d924-6495-4800-ab5a-92486aa0fa3e]
- 47. Dimer formation drives the activation of the cell death protease ... [https://pubmed.ncbi.nlm.nih.gov/11734640/]
- 48. Engineering a Dimeric Caspase-9: A Re-evaluation of the ... [https://journals.plos.org/plosbiology/article/figures?id=10.1371/journal.pbio.0030183]
- 49. - or Proximity -Induced Conformation ? Caspase Activation [https://journals.plos.org/plosbiology/article?id=10.1371/journal.pbio.0030198]
- 50. Caspase-9 [https://en.wikipedia.org/wiki/Caspase-9]
- 51. structural and molecular mechanisms and functions in cell ... [https://www.nature.com/articles/s41421-025-00791-3]
- 52. Protein posttranslational modifications in health and diseases [https://pmc.ncbi.nlm.nih.gov/articles/PMC10152985/]
- 53. The Chemical Biology of Protein Phosphorylation - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC3074175/]
- 54. Monitoring phosphorylation and acetylation of CRISPR ... [https://www.nature.com/articles/s41598-024-51887-x]
- 55. PTMNavigator: interactive visualization of differentially ... [https://www.nature.com/articles/s41467-024-55533-y]
- 56. Caspase-9 holoenzyme is a specific and optimal pro ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC2904439/]
- 57. Caspase-9 Activation of Procaspase-3 but not ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC8489496/]
- 58. Exploring caspase functions in mouse models | Apoptosis [https://link.springer.com/article/10.1007/s10495-024-01976-z]
- 59. Proteolytic Processing of the Caspase-9 Zymogen Is Required ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC3663534/]
- 60. Live imaging of apoptotic signaling flow using tunable ... [https://www.nature.com/articles/s41598-022-25286-z]
- 61. Live Cell Imaging of Caspase Activation for High Content ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC3613133/]
- 62. Regulation of the Apaf-1–caspase-9 apoptosome - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC2939798/]
- 63. Caspase-9 swings both ways in the apoptosome - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC5383358/]
- 64. The apoptosome activates caspase-9 by dimerization - PubMed [https://pubmed.ncbi.nlm.nih.gov/16630894/]
- 65. A near atomic structure of the active human apoptosome [https://elifesciences.org/articles/17755]
- 66. A docking model of key components of the DISC complex [https://www.sciencedirect.com/science/article/pii/S0014579301021627]
- 67. F-actin–rich territories coordinate apoptosome assembly and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10162420/]
- 68. Regulation of the Apaf-1/Caspase-9 Apoptosome by ... [https://www.sciencedirect.com/science/article/pii/S0021925820864242]
- 69. Cytochrome c Promotes Caspase-9 Activation by Inducing ... [https://www.sciencedirect.com/science/article/pii/S0021925820893958]
- 70. Cell-to-cell variability in inducible Caspase9-mediated cell ... [https://www.nature.com/articles/s41419-021-04468-z]
- 71. A Complex between Atg7 and Caspase-9 - PubMed Central [https://pmc.ncbi.nlm.nih.gov/articles/PMC3945314/]
- 72. Caspase-dependent non-apoptotic processes in ... [https://www.nature.com/articles/cdd201736]
- 73. Cell Biology Activation of Caspase-9, but Not ... [https://www.sciencedirect.com/science/article/pii/S002192581976219X]
- 74. Caspase-9: A Multimodal Therapeutic Target With Diverse ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC8299054/]
- 75. Caspase-9 inhibition confers stronger neuronal and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10336837/]
- 76. Helminthic larval stage induces cellular apoptosis via caspase ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12507911/]