Mitochondrial Membrane Potential: A Master Regulator of Metabolic Specialization and Partitioning in Health and Disease
This article explores the paradigm-shifting role of the mitochondrial membrane potential (MMP) beyond its canonical function in ATP production.
Mitochondrial Membrane Potential: A Master Regulator of Metabolic Specialization and Partitioning in Health and Disease
Abstract
This article explores the paradigm-shifting role of the mitochondrial membrane potential (MMP) beyond its canonical function in ATP production. We examine how MMP acts as a dynamic signaling hub that directs metabolic specialization by partitioning mitochondria into distinct subpopulations dedicated to either oxidative energy production or reductive biosynthesis. Targeting researchers and drug development professionals, this review synthesizes foundational concepts, current methodological approaches for assessing MMP, common challenges in its measurement, and validation strategies linking MMP dynamics to disease pathologies. We highlight how understanding MMP-mediated metabolic partitioning opens novel therapeutic avenues for cancer, neurodegenerative disorders, and metabolic diseases.
The Bioenergetic Signal: How MMP Governs Metabolic Fate and Compartmentalization
Mitochondrial membrane potential (MMP) is a central intermediate in oxidative energy metabolism, traditionally viewed as a simple driver of ATP synthesis. However, emerging research reveals that MMP serves as a dynamic signaling hub that integrates cellular status, regulates metabolic specialization, and directs mitochondrial quality control. This technical review synthesizes current understanding of MMP generation and dissipation mechanisms, highlighting their roles beyond maintaining protonmotive force. We provide quantitative analysis of MMP components, detailed experimental protocols for MMP assessment, and visualization of key signaling pathways. The findings underscore MMP's critical function in neuronal plasticity, metabolic partitioning, and cellular adaptation, offering new perspectives for therapeutic targeting in metabolic and neurodegenerative diseases.
The mitochondrial membrane potential (ΔΨm) represents a fundamental parameter of cellular energetic status, generated by charge separation across the inner mitochondrial membrane (IMM). According to the chemiosmotic theory established by Peter Mitchell, MMP constitutes the primary component of the protonmotive force (pmf), which couples electron transport chain (ETC) activity to ATP production [1]. The pmf consists of both an electrical gradient (ΔΨm) and a chemical proton gradient (ΔpH), with MMP typically contributing approximately 80% of the total pmf under physiological conditions [2] [3] [4].
While MMP's canonical role in driving ATP synthesis via ATP synthase is well-established, recent research has revealed that MMP dynamics extend far beyond this fundamental function. MMP undergoes rapid adjustments in response to acute changes in cellular energy demand and sustains modifications during developmental processes, positioning it as a key regulator of cellular signaling [2]. These dynamic potential changes influence reactive oxygen species (ROS) production, calcium handling, and mitochondrial quality control, enabling localized and time-sensitive regulation of cellular function [2] [1]. In specialized cells such as neurons, MMP changes coordinate synaptic plasticity by linking metabolic state to structural changes at synapses [2].
This review examines the mechanisms of MMP generation and dissipation, with particular emphasis on their implications for metabolic specialization and partitioning research. We provide quantitative frameworks for measuring MMP, detailed experimental approaches, and visualization of the complex signaling networks regulated by this fundamental bioenergetic parameter.
Theoretical Framework: Composition and Regulation of the Protonmotive Force
Components of the Protonmotive Force
The protonmotive force (pmf) represents the electrochemical potential gradient of protons across the inner mitochondrial membrane. Mathematically, the pmf is described by the following equation:
pmf = ΔΨ - ZΔpH
Where ΔΨ represents the electrical potential component (MMP), ΔpH represents the chemical proton gradient, and Z is a constant equal to approximately 59 mV at 25°C [3]. Under physiological conditions, the total pmf measures approximately 170-200 mV, with MMP typically ranging between -139 mV and -180 mV in various cell types, and ΔpH contributing approximately 30 mV (equivalent to 0.5 pH units) [2] [3] [5].
Table 1: Quantitative Distribution of Protonmotive Force Components Across Biological Systems
| System | Total pmf (mV) | ΔΨ Component (mV) | ΔpH Component (mV) | ΔΨ Contribution (%) | Reference |
|---|---|---|---|---|---|
| Isolated Mitochondria (Classical) | 190-200 | -160 to -180 | ~30 | 80-85% | [3] [5] |
| Cultured Rat Cortical Neurons | - | -139 ± 5 (Resting) | - | - | [5] |
| HeLa Cells | - | -108 to -158 (Regulated) | - | - | [6] |
| Theoretical Maximum (No ion transport) | - | ~99% of pmf | ~1% of pmf | ~99% | [3] |
The relative contribution of ΔΨ and ΔpH to the total pmf is not fixed but varies according to cellular conditions. Secondary transport of ions, particularly potassium, plays a crucial role in maintaining the physiological balance between these components [3]. Without such secondary ion transport, the pmf would exist almost exclusively as ΔΨ due to the low electrical capacitance of the IMM and the considerable pH buffering capacity of the mitochondrial matrix [3].
Spatial Heterogeneity of MMP
Recent super-resolution microscopy studies have revealed that MMP is not uniform across mitochondrial subcompartments. The inner mitochondrial membrane is divided into two structurally and functionally distinct domains: the cristae membrane (CM) and the inner boundary membrane (IBM), separated by the crista junction (CJ) [6].
The CM, which harbors the ETC complexes I, III, and IV, demonstrates a higher (more negative) membrane potential (ΔΨC) compared to the IBM (ΔΨIBM) [6]. This compartmentalization creates ultra-structures with different phospholipid and protein compositions, shapes, characteristics, and functions. The CJ serves as a critical barrier that regulates ion movement and ensures distinct electrical potentials, with evidence suggesting that it can seal and isolate the CM in terms of membrane potential [6].
Table 2: Experimental Measurements of Mitochondrial Membrane Potential Across Cell Types and Conditions
| Cell Type/Condition | MMP Value (mV) | Measurement Technique | Biological Significance | Reference |
|---|---|---|---|---|
| Cortical Neurons (Resting) | -139 ± 5 | TMRM fluorescence calibration | Baseline for neuronal metabolism | [5] |
| Cortical Neurons (Stimulated) | -126 to -154 | TMRM fluorescence calibration | Ca2+-dependent regulation | [5] |
| Cortical Neurons (High K+ Depolarization) | -108 ± 4 | TMRM fluorescence calibration | Response to increased ATP demand | [5] |
| Beta-cells (High Glucose) | Hyperpolarization | Fluorescent dyes | Stimulates insulin release | [4] |
| Cristae Membrane (CM) | Higher (more negative) than IBM | STED/SIM super-resolution microscopy | Site of proton pumping | [6] |
| Inner Boundary Membrane (IBM) | Lower (less negative) than CM | STED/SIM super-resolution microscopy | Interface with outer membrane | [6] |
This spatial heterogeneity of MMP has profound implications for mitochondrial function. The potential gradient between CM and IBM influences ATP production, ROS generation, and calcium handling, creating specialized microdomains within individual mitochondria [6].
Methodologies for Quantitative MMP Assessment
Fluorescence-Based Measurement Techniques
The quantitative assessment of MMP in living cells presents significant technical challenges due to the complex behavior of potentiometric probes and the need to account for multiple confounding factors. The following protocol outlines a rigorous approach for absolute quantification of MMP using tetramethylrhodamine methyl ester (TMRM), based on established methodologies [5] [4].
Protocol 1: Absolute Quantification of MMP in Cultured Cells Using TMRM
Principle This method utilizes the Nernstian distribution of lipophilic cations across energized membranes. TMRM accumulates in mitochondria according to the Nernst equation, with fluorescence intensity reflecting the equilibrium distribution between extracellular space, cytoplasm, and mitochondrial matrix [5].
Reagents and Equipment
- Tetramethylrhodamine methyl ester (TMRM)
- MitoTracker Green FM (MTG) for morphological reference
- Plasma membrane potential indicator (PMPI), e.g., bis-oxonol dye
- Extracellular buffer: 125 mM NaCl, 5 mM KCl, 1 mM MgCl2, 1 mM CaCl2, 20 mM HEPES, 5 mM glucose, pH 7.4
- Calibration solutions: High-K+ buffers with varying TMRM concentrations
- Imaging system: Epifluorescence or confocal microscope with temperature control
- Image analysis software capable of quantifying fluorescence intensities
Procedure
- Cell Preparation and Dye Loading
- Culture cells on appropriate substrates (e.g., poly-ornithine-coated coverslips for neurons)
- Load cells with 500 nM MTG for 30 minutes at 37°C to label mitochondrial morphology
- Incubate with 13.5 nM TMRM for 45-60 minutes at 37°C to achieve equilibrium distribution
- Include PMPI staining if simultaneous plasma membrane potential measurement is required
Image Acquisition
- Acquire simultaneous dual-channel images (MTG and TMRM) using appropriate filter sets
- Maintain constant imaging parameters (exposure time, laser power, gain) throughout experiment
- For time-course experiments, acquire images at regular intervals (e.g., every 30-60 seconds)
- Include calibration samples with known TMRM concentrations
Image Analysis and Data Processing
- Use MTG channel to define mitochondrial regions of interest (ROIs)
- Apply background subtraction to both MTG and TMRM channels
- Calculate TMRM fluorescence intensity within mitochondrial ROIs
- Account for matrix:cell volume ratio, dye binding coefficients, and activity coefficients
- Convert fluorescence intensities to absolute MMP values using calibration curve
Validation and Controls
- Confirm mitochondrial specificity with positive controls (e.g., oligomycin-induced hyperpolarization)
- Validate depolarization with FCCP (1-2 μM) as negative control
- Assess plasma membrane potential contribution using PMPI
- Determine signal linearity with TMRM concentration series
Calculation Absolute MMP values are calculated using a biophysical model of probe compartmentation and dynamics based on Eyring rate theory. The model accounts for ΔΨP-dependent redistribution, Nernstian behavior, matrix:cell volume ratio, high- and low-affinity binding, activity coefficients, background fluorescence, and optical dilution [5].
The standard error of the mean for absolute calibrated values of resting MMP, including all biological and systematic measurement errors introduced by calibration parameters, is typically less than 11 mV. Between samples treated differently, the equivalent error is approximately 5 mV [5].
Super-Resolution Analysis of Spatial MMP Gradients
Advanced imaging techniques now enable resolution of MMP gradients between mitochondrial subcompartments. The following protocol describes the assessment of spatial membrane potential gradients (SMPG) using structured illumination microscopy (SIM) [6].
Protocol 2: Analysis of Spatial Membrane Potential Gradients Using SIM
Principle This method exploits the differential distribution of TMRM between cristae membranes (CM) and inner boundary membranes (IBM) at varying dye concentrations. At low concentrations, TMRM preferentially accumulates in CM with higher membrane potential, while saturation at high concentrations enables distribution to IBM [6].
Reagents and Equipment
- TMRM and MitoTracker Green FM (MTG)
- Live cell imaging buffer
- Super-resolution microscope (e.g., N-SIM system)
- Image analysis software with custom algorithms
Procedure
- Sample Preparation and Staining
- Culture cells on high-precision coverslips suitable for super-resolution microscopy
- Stain cells with 500 nM MTG (constant) and varying TMRM concentrations (1.35-81 nM)
- Allow 45-60 minutes for dye equilibration at 37°C
Dual-Channel SIM Imaging
- Perform simultaneous dual-channel N-SIM imaging of MTG and TMRM
- Acquire z-stacks to capture full mitochondrial architecture
- Maintain physiological conditions (37°C, 5% CO2) throughout imaging
SMPG Analysis
Method I: IBM Association Index
- Use MTG channel to define mitochondrial boundaries via automated Otsu thresholding
- Generate IBM and CM regions by sequential shrinking and widening of boundaries
- Calculate fluorescence intensity ratio: IBM Association Index = TMRMIBM/TMRMCM
Method II: ΔFWHM Analysis
- Extract cross-section intensity profiles of MTG and TMRM signals
- Calculate full width at half maximum (FWHM) for both channels
- Determine ΔFWHM = FWHMMTG - FWHMTMRM
Dynamic SMPG Monitoring
- Stimulate cells with appropriate agonists (e.g., histamine for Ca2+ release)
- Acquire time-lapse SIM images pre- and post-stimulation
- Calculate changes in IBM Association Index or ΔFWHM over time
Interpretation A decrease in IBM Association Index or ΔFWHM following stimulation indicates relative hyperpolarization of CM compared to IBM, typically resulting from increased TCA cycle activity and enhanced proton pumping at cristae membranes [6].
MMP Generation: Mechanisms and Regulation
Electron Transport Chain and Proton Pumping
The primary mechanism for MMP generation involves vectorial proton translocation by the electron transport chain (ETC) complexes. Complexes I, III, and IV function as proton pumps, transferring electrons from reducing equivalents (NADH, FADH2) to molecular oxygen while moving protons from the mitochondrial matrix to the intermembrane space [2] [1].
Complex I (NADH:ubiquinone oxidoreductase) transfers four protons per two electrons, complex III (ubiquinol:cytochrome c oxidoreductase) transfers four protons but only two positive charges, and complex IV (cytochrome c oxidase) transfers two protons and four positive charges [3] [1]. This differential transfer of protons versus charges explains the varying sensitivity of ETC complexes to ΔΨ and ΔpH components of the pmf.
Multiple entry points exist for electrons into the ETC, including complexes I and II, as well as dihydroorotate dehydrogenase (DHODH), mitochondrial glycerol 3-phosphate dehydrogenase (mGPDH), and electron transfer flavoprotein (ETF) during fatty acid oxidation [1]. This diversity of electron inputs allows integration of various metabolic pathways into MMP regulation.
Regulation of MMP Generation
MMP generation is precisely regulated through multiple mechanisms:
Substrate Availability: The provision of reducing equivalents from carbohydrate, lipid, and amino acid metabolism directly influences ETC activity and proton pumping [1].
Calcium Signaling: Mitochondrial calcium uptake activates dehydrogenases in the tricarboxylic acid (TCA) cycle, increasing electron flow to the ETC and enhancing MMP generation [6] [7].
Cristae Junction Permeability: The dynamic regulation of CJ opening by proteins such as MICU1 and OPA1 controls ion access to cristae membranes, thereby influencing local MMP generation [6].
Electron Leak and ROS Production: Under conditions of high MMP, electron leak from the ETC increases, resulting in superoxide formation and potentially creating a feedback loop that modulates MMP generation [1].
MMP Dissipation: Pathways and Physiological Significance
Canonical Dissipation Pathways
MMP dissipation occurs through several regulated mechanisms that convert electrochemical energy into useful work or heat:
ATP Synthase: The primary energy-conserving pathway for MMP dissipation, wherein proton flux through the F0 subunit drives ATP synthesis in the F1 subunit [2] [4].
Proton Leak: Basal proton conductance across the IMM accounts for 20-50% of standard metabolic rate, representing a significant futile cycle that dissipates energy as heat [1].
Calcium Cycling: Electroneutral Ca2+ uptake via the mitochondrial calcium uniporter coupled with Ca2+ efflux mechanisms represents an energy-dissipating cycle [2].
Substrate Cycling: Various anion transport systems, including the ATP/ADP translocase and phosphate carrier, contribute to controlled MMP dissipation [3].
Regulated Uncoupling
Mitochondria possess specialized mechanisms for regulated uncoupling that fine-tune MMP under physiological conditions:
Uncoupling Proteins (UCPs): UCPs dissipate MMP as heat through controlled proton leak [2]. Genetic variants in UCPs (UCP2-UCP4) have been linked to metabolic and neurological disorders, highlighting their physiological importance [2].
Adenine Nucleotide Translocase (ANT): In addition to its primary role in ADP/ATP exchange, ANT can facilitate proton leak under specific conditions [1].
Lipid-Mediated Uncoupling: Fatty acids act as natural uncouplers by cycling in their protonated and anionic forms across the IMM, with this process potentially serving as a mitochondrial mediator of apoptotic signaling [7].
The balance between MMP generation and dissipation determines the steady-state MMP, which operates within a relatively narrow dynamic range in coupled mitochondria. This limited dynamic range reflects the thermodynamic constraints of the OXPHOS system, which becomes unstable at both very high and very low MMP values [4].
MMP in Metabolic Specialization and Signaling
Metabolic Compartmentalization
MMP plays a crucial role in establishing metabolically distinct mitochondrial subpopulations within individual cells. Classic work in cardiac muscle identified subsarcolemmal and interfibrillar mitochondria with differing respiratory capacity, protein composition, and stress sensitivity [2]. Recent research indicates that MMP dynamics contribute to this metabolic specialization through several mechanisms:
Metabolic Enzyme Partitioning: Elevated MMP enhances the activity and filamentation of enzymes such as pyrroline-5-carboxylate synthase (P5CS), promoting reductive biosynthesis and the formation of substrate-producing mitochondria [2].
Oxidative-Reductive Switching: MMP coordinates the switching between oxidative (ATP-producing) and reductive (biosynthetic) metabolic programs, allowing mitochondria to specialize according to cellular demands [2].
Nutrient Allocation: Regional variations in MMP may influence how mitochondrial fragments are sorted after fission, directing them toward either network reintegration or degradation based on functional capacity [2].
Quality Control and Fate Determination
MMP serves as a key determinant of mitochondrial fate through quality control mechanisms:
Mitophagy Regulation: Reduced MMP triggers PINK1 accumulation on the outer mitochondrial membrane, recruiting Parkin and initiating mitophagy to eliminate damaged organelles [2].
Fission-Fusion Dynamics: Following mitochondrial fission, fragments with higher MMP relative to baseline typically re-fuse with the network, while those with lower MMP are targeted for degradation [2].
Protein Import Control: MMP drives the import of nuclear-encoded mitochondrial proteins through the TIM23 complex, potentially allowing functional assessment during biogenesis [2].
These MMP-dependent quality control mechanisms ensure the maintenance of a healthy mitochondrial network while allowing metabolic specialization according to cellular requirements.
Research Reagent Solutions
Table 3: Essential Research Reagents for MMP Studies
| Reagent/Category | Specific Examples | Function/Application | Key Considerations |
|---|---|---|---|
| Potentiometric Dyes | TMRM, TMRE | MMP-dependent accumulation | Nernstian distribution; concentration-dependent saturation [5] [6] |
| JC-1 | J-aggregate formation at high MMP | Non-equilibrium accumulation; ratio metric but problematic for quantification [5] [4] | |
| Morphological Reference Dyes | MitoTracker Green FM | IMM reference marker | Potential-dependent accumulation but fixed after binding; used for morphology [6] |
| Plasma Membrane Potential Indicators | bis-oxonol dyes (PMPI) | ΔΨP measurement | Critical for accounting for plasma membrane potential effects [5] |
| MMP Modulators | Oligomycin | ATP synthase inhibitor | Induces hyperpolarization by reducing ΔΨm consumption [8] [4] |
| FCCP/CCCP | Protonophores | Complete depolarization; validate MMP dependence [8] [1] | |
| Rotenone, Antimycin A | ETC inhibitors (Complex I, III) | Inhibit MMP generation; test proton pump dependence [6] | |
| Ion Modulators | Histamine | IP3-generating agonist | Induces mitochondrial Ca2+ uptake and CM hyperpolarization [6] |
| High K+ medium | Plasma membrane depolarizer | Increases ATP demand and depolarizes ΔΨm [5] | |
| Advanced Imaging Systems | STED/SIM microscopy | Super-resolution SMPG analysis | Resolves cristae vs. IBM potential differences [6] |
MMP represents far more than a simple intermediate in ATP production, functioning as a dynamic regulatory hub that integrates cellular status and directs metabolic specialization. The spatial and temporal heterogeneity of MMP, particularly the gradients between cristae and inner boundary membranes, creates specialized microdomains that enable multifaceted regulation of mitochondrial function.
Future research in metabolic specialization and partitioning would benefit from increased attention to:
Single-Organelle Analysis: Developing approaches to monitor MMP dynamics in individual mitochondria within living cells would illuminate heterogeneity in metabolic specialization.
Cristae-Specific Targeting: Creating tools to specifically manipulate MMP in cristae versus inner boundary membranes would help elucidate compartment-specific functions.
Metabolic Memory: Investigating how historical MMP patterns influence long-term mitochondrial fate and specialization represents a promising avenue.
Therapeutic Targeting: Leveraging MMP dynamics for selective manipulation of metabolic pathways offers exciting possibilities for treating metabolic diseases, neurodegenerative disorders, and cancer.
The emerging paradigm positions MMP as a central integrator of cellular metabolism, with its generation and dissipation mechanisms coordinating everything from energy production to cell fate decisions. Understanding these complex dynamics will be essential for advancing both basic mitochondrial biology and therapeutic applications targeting metabolic regulation.
Matrix metalloproteinases (MMPs) represent a family of approximately 24 zinc-dependent endopeptidases traditionally recognized for their ability to degrade extracellular matrix (ECM) components, thereby facilitating tissue remodeling and resolution of excess matrix during fibrosis [9]. However, emerging research has unveiled a complex landscape of non-canonical functions that extend far beyond ECM degradation. These multifunctional enzymes are now recognized as crucial signaling modulators that influence cellular processes through both proteolytic and non-proteolytic mechanisms, impacting inflammation, cell migration, proliferation, and intracellular signaling cascades [10].
The conventional classification of MMPs—including collagenases (MMP-1, -8, -13), gelatinases (MMP-2, -9), stromelysins (MMP-3, -10, -11), matrilysins (MMP-7, -26), and membrane-type MMPs (MMP-14, -15, -16, -24)—reflects their structural diversity but fails to capture their functional complexity in pathophysiological contexts [9]. This whitepaper explores the paradigm shift in understanding MMPs from mere matrix-degrading enzymes to sophisticated signaling integrators, with particular emphasis on their implications for mitochondrial function and metabolic specialization in disease progression.
Non-Canonical MMP Functions in Cellular Signaling
Proteolytic Processing of Signaling Molecules
MMPs demonstrate remarkable capacity to process diverse non-matrix substrates, thereby activating latent signaling pathways and modulating cellular responses:
- Cytokine and Chemokine Modulation: MMP-1, -3, and -7 release TNF-α from the cell surface, while MMP-2 and -9 activate latent TGF-β1 and TGF-β2, thereby influencing inflammatory processes and fibrotic responses [9].
- Receptor Shedding and Activation: MMP-mediated cleavage of cell surface receptors, including cadherins and integrins, disrupts cell-to-cell and cell-to-matrix adhesion, facilitating cellular migration and invasion [11]. MMP-3 directly cleaves E-cadherin, promoting epithelial-to-mesenchymal transition (EMT) in carcinogenesis [11].
- Non-Canonical Notch Processing: Specific MMPs, including MMP-7 and MT1-MMP (MMP-14), process Notch receptors independently of the canonical ADAM proteases, leading to Notch intracellular domain (NICD) release, nuclear translocation, and subsequent target gene expression [11]. This non-canonical activation drives aggressive cancer traits such as invasion, metastasis, angiogenesis, and EMT.
Table 1: Non-Canonical Substrates and Signaling Pathways Activated by MMPs
| MMP | Non-Matrix Substrate | Signaling Consequence | Pathological Context |
|---|---|---|---|
| MMP-1, -3, -7 | Membrane-bound TNF-α | Release of active TNF-α | Inflammation regulation |
| MMP-2, -9 | Latent TGF-β1, TGF-β2 | TGF-β pathway activation | Fibrosis, immunomodulation |
| MMP-3 | E-cadherin | Loss of cell adhesion, EMT | Cancer metastasis |
| MMP-7, MT1-MMP | Notch receptor | Non-canonical Notch signaling | Cancer progression |
| MMP-8 | Unknown HSC activator | Increased COL1A1 expression, migration | Liver fibrosis progression |
Context-Dependent Pro-Fibrotic Functions
Contrary to their traditional anti-fibrotic role in ECM resolution, certain MMPs demonstrate pro-fibrotic activities in specific contexts. In hepatic stellate cells (HSCs), macrophage-derived MMP-8 (collagenase-2) promotes HSC activation, migration, and collagen type I, alpha 1 (COL1A1) expression, thereby contributing to liver fibrosis progression rather than resolution [12] [13]. Similarly, MMP-19 deficiency reduces liver fibrosis in murine models, suggesting a pro-fibrotic role for this enzyme despite its ECM-degrading capability [13].
Methodological Framework for Investigating Non-Canonical MMP Functions
Experimental Models and Analytical Techniques
Research into non-canonical MMP roles employs sophisticated methodological approaches to elucidate complex signaling networks:
- Cellular Migration Assays: Boyden chamber assays quantitatively measure MMP-mediated cell migration and invasion, as demonstrated in HSC migration studies [12] [13].
- Gene Expression Analysis: Real-time PCR with SYBR green enables quantification of MMP-responsive genes (e.g., COL1A1, α-SMA) following experimental manipulation [12] [13].
- Protein Profiling Arrays: Human MMP antibody arrays facilitate comprehensive profiling of MMP expression patterns in response to stimuli such as lipopolysaccharide (LPS) in macrophage models [12] [13].
- Zymography: Gelatin or casein zymography using SDS-PAGE gels containing substrate (1mg/ml) assesses MMP enzymatic activity in conditioned media or cell lysates [12] [13].
- Genetic Manipulation: Knockdown approaches (siRNA/shRNA) and genetic knockout models determine causal relationships between specific MMPs and functional outcomes in disease pathogenesis [14].
The Scientist's Toolkit: Essential Research Reagents
Table 2: Key Research Reagents for Investigating Non-Canonical MMP Functions
| Reagent/Tool | Application | Experimental Function |
|---|---|---|
| Human MMP Antibody Array (Abcam) | MMP expression profiling | Simultaneous detection of multiple MMPs in cell culture supernatants and lysates |
| Boyden Chamber Assay | Cell migration quantification | Measures MMP-mediated cellular migration and invasion capabilities |
| Gelatin/Casein Zymography | MMP activity assessment | Evaluates functional MMP activity through substrate degradation patterns |
| SYBR Green Real-Time PCR | Gene expression analysis | Quantifies expression of MMPs and MMP-regulated genes (COL1A1, α-SMA) |
| LPS (Lipopolysaccharide) | Macrophage stimulation | Induces MMP expression in macrophage models to study inflammatory responses |
| Recombinant Active MMPs | Functional studies | Direct application to cells to assess MMP-specific signaling effects |
MMP-Mitochondria Crosstalk: An Emerging Signaling Axis
Mitochondrial Membrane Potential as a Therapeutic Vulnerability
While direct mechanistic links between MMP activity and mitochondrial membrane potential (ΔΨm) regulation represent an emerging frontier, compelling parallel evidence suggests potential intersections. Studies on clonal hematopoiesis demonstrate that elevated ΔΨm constitutes a therapeutic vulnerability in mutant hematopoietic stem and progenitor cells (HSPCs) [15]. Dnmt3a-mutant HSPCs sustain elevated mitochondrial respiration associated with increased oxidative phosphorylation gene expression, high ΔΨm, and greater dependence on mitochondrial respiration compared to wild-type counterparts [15].
This bioenergetic vulnerability can be therapeutically exploited using long-chain alkyl-TPP molecules (MitoQ, d-TPP) that selectively accumulate in mitochondria with elevated ΔΨm, causing reduced mitochondrial respiration, mitochondrial-driven apoptosis, and ablation of competitive advantage in mutant HSPCs [15]. This paradigm highlights the potential for targeting metabolic specialization in disease contexts.
Methodological Considerations for ΔΨm Assessment
Accurate determination of mitochondrial membrane potential requires careful methodological implementation:
- Fluorescent Probe Selection: Tetramethylrhodamine methyl ester (TMRM), MitoSOX, and Rhod-2AM serve as standard fluorescent probes for assessing ΔΨm, reactive oxygen species, and calcium levels, respectively [16].
- Interpretation Caveats: ΔΨm has limited sensitivity and specificity for reporting changes in OXPHOS activity in coupled mitochondria, as fluorescent signals from commonly used dyes do not unequivocally indicate increased mitochondrial function [4].
- Complementary Approaches: Oxygen consumption measurements provide greater sensitivity for detecting OXPHOS changes compared to ΔΨm assessment alone, particularly when ΔΨm shifts are minimal despite functional alterations [4].
Therapeutic Targeting of Non-Canonical MMP Functions
MMP-Based Diagnostic and Therapeutic Strategies
The expanding understanding of non-canonical MMP functions opens innovative avenues for therapeutic intervention:
- MMP-Targeted Drug Delivery: MMP overexpression in pathological contexts can be exploited for targeted drug delivery systems utilizing MMP-cleavable substrates [9].
- Selective MMP Inhibition: Challenges in MMP inhibitor development include achieving subtype selectivity and minimizing metabolic side effects, necessitating sophisticated structural approaches [11].
- Biomarker Development: MMP-3 demonstrates particular promise as both a biomarker for early diagnosis and a therapeutic target for selective inhibition and modulation across inflammatory diseases, cardiovascular diseases, neurodegenerative disorders, and cancer [10].
- Druggable Genome Identification: Mendelian randomization approaches identify promising drug targets within the MMP family, such as MMP-25 for chronic periodontitis, highlighting the therapeutic potential of targeting specific MMPs [17].
Clinical Translation Challenges
Despite promising preclinical findings, therapeutic targeting of non-canonical MMP functions faces substantial translational challenges:
- Context-Dependent Actions: The dual pro- and anti-fibrotic functions of specific MMPs (e.g., MMP-8, -19) necessitate precise contextual understanding before therapeutic intervention [12] [13].
- Spatiotemporal Control: Successful therapeutic modulation requires exquisite spatiotemporal control to avoid disrupting physiological MMP functions in tissue homeostasis [10].
- Disease Stratification: Patient stratification based on MMP expression patterns and mitochondrial metabolic profiles may enhance therapeutic efficacy while minimizing adverse effects [15].
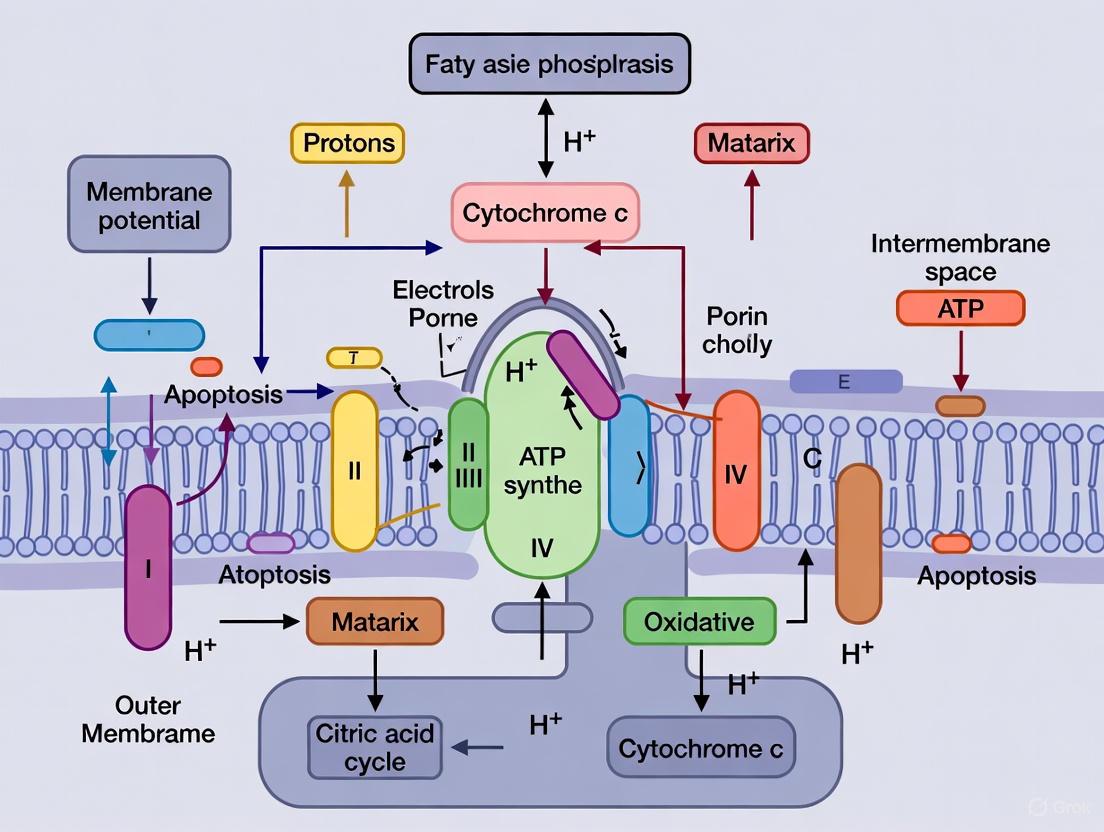
Visualizing Non-Canonical MMP Signaling Networks
Non-Canonical MMP Signaling to Mitochondria - This diagram illustrates how extracellular MMP activity activates intracellular signaling cascades that ultimately influence mitochondrial function and cellular fate decisions.
The evolving paradigm of MMPs as signaling hubs rather than merely matrix-degrading enzymes reveals complex regulatory networks with profound implications for cellular metabolism and specialization. The non-canonical functions of specific MMPs—including cytokine activation, receptor processing, and non-canonical protease activities—position these enzymes as critical integrators of extracellular cues and intracellular responses. Emerging evidence suggests potential connections between MMP-mediated signaling and mitochondrial metabolic reprogramming, particularly through the modulation of mitochondrial membrane potential and respiratory function.
Future research directions should prioritize elucidating the precise molecular mechanisms linking specific MMP activities to mitochondrial bioenergetics, developing sophisticated targeting strategies that account for context-dependent MMP functions, and exploring combinatorial approaches that simultaneously modulate MMP signaling and mitochondrial metabolism. Such integrated investigations promise to unlock novel therapeutic opportunities for diverse pathological conditions characterized by dysregulated extracellular proteolysis and metabolic specialization, from fibrotic disorders to cancer progression.
Mitochondrial membrane potential (MMP) transcends its canonical role as a mere driver of ATP synthesis, emerging as a pivotal regulator of mitochondrial metabolic specialization. Recent advancements elucidate how spatial and temporal dynamics of MMP facilitate the partitioning of mitochondria into distinct subpopulations dedicated to either oxidative phosphorylation (OXPHOS) or reductive biosynthesis. This whitepaper synthesizes current research demonstrating that gradients in MMP act as a fundamental bioelectric signal coordinating metabolic compartmentalization, influencing processes ranging from cellular reprogramming in cancer to synaptic plasticity in neurons. We provide a comprehensive technical overview of the mechanisms underlying MMP-driven metabolic partitioning, detailed experimental methodologies for its investigation, and quantitative frameworks for data interpretation, offering researchers a foundational guide for exploring this emerging paradigm in mitochondrial biology.
The classical view of mitochondria as homogeneous cellular powerhouses has been fundamentally challenged by evidence revealing significant functional heterogeneity among these organelles. Mitochondria can specialize metabolically, forming distinct subpopulations dedicated to specific biochemical outputs. This metabolic specialization is now recognized as a critical adaptive mechanism that allows cells to meet diverse energetic and biosynthetic demands. Central to this partitioning is the mitochondrial membrane potential (MMP), an electrochemical gradient across the inner mitochondrial membrane generated by proton pumping through electron transport chain (ETC) complexes I, III, and IV [2] [18].
The protonmotive force (PMF) driving ATP synthesis consists of both an electrical gradient (MMP, approximately -180 mV) and a chemical gradient (ΔpH, approximately 0.4 pH units), with MMP constituting the dominant component under physiological conditions [2]. Beyond its fundamental role in energy transduction, MMP serves as a dynamic signaling hub that integrates cellular status and directs mitochondrial functional specialization. Spatial and temporal variations in MMP influence reactive oxygen species production, calcium handling, and protein import machinery, ultimately enabling the emergence of metabolically distinct mitochondrial pools [2] [6].
This technical review examines the molecular mechanisms through which MMP gradients orchestrate metabolic partitioning, details advanced methodologies for quantifying and manipulating these processes, and discusses implications for therapeutic targeting in disease contexts characterized by metabolic dysregulation, including cancer and neurodegenerative disorders.
Molecular Mechanisms of MMP-Driven Metabolic Partitioning
Bioelectric Control of Enzyme Compartmentalization
The establishment of metabolically specialized mitochondrial subpopulations is fundamentally regulated by MMP-dependent protein import and enzyme activity. Recent research has identified specific metabolic enzymes whose localization and function are directly modulated by MMP dynamics:
Pyrroline-5-Carboxylate Synthase (P5CS): This rate-limiting enzyme in proline biosynthesis serves as a central regulatory node for mitochondrial metabolic switching. Under conditions of elevated MMP relative to baseline, P5CS activity is enhanced, promoting its assembly into filamentous structures that drive reductive biosynthesis pathways. Conversely, reduced MMP inhibits filamentation and limits substrate production for biosynthetic processes [2].
Import Machinery Sensitivity: The translocation of nuclear-encoded mitochondrial proteins depends on the presence of positively charged targeting signals that are electrophoretically pulled across the inner membrane by the MMP. Regional variations in MMP can therefore create heterogeneous protein composition across the mitochondrial network, establishing distinct functional identities [2].
Table 1: MMP-Sensitive Metabolic Enzymes in Mitochondrial Specialization
| Enzyme/Complex | Metabolic Pathway | MMP Response | Functional Outcome |
|---|---|---|---|
| P5CS | Proline Biosynthesis | Enhanced activity with elevated MMP | Drives reductive metabolism |
| ETC Complexes I, III, IV | OXPHOS | Generates MMP through proton pumping | Establishes bioelectric gradient |
| Protein Import Machinery | Protein Translocation | Dependent on MMP for electrophoretic pull | Determines mitochondrial proteome |
| Uncoupling Proteins (UCPs) | MMP Dissipation | Regulated dissipation of MMP | Prevents excessive hyperpolarization |
Spatial Organization of Mitochondrial Membranes
The inner mitochondrial membrane exhibits complex ultrastructural organization that creates microdomains with distinct MMP characteristics. Super-resolution microscopy techniques have revealed that the cristae membrane (CM) and inner boundary membrane (IBM) maintain different electrical potentials, with the CM typically exhibiting a higher (more negative) membrane potential (ΔΨC) compared to the IBM (ΔΨIBM) [6]. This potential difference is maintained by the crista junction (CJ), which acts as a diffusion barrier separating these compartments.
The regulation of CJ permeability is controlled by proteins including MICU1 and OPA1, which respond to cellular cues such as calcium concentrations. During calcium elevation, MICU1 oligomers disassemble, increasing CJ permeability and allowing enhanced communication between compartments [6]. This architectural specialization enables simultaneous maintenance of distinct bioenergetic environments within individual mitochondria, supporting concurrent operation of oxidative and reductive metabolic programs.
Figure 1: Spatial Organization of Mitochondrial Membrane Potential. The inner mitochondrial membrane is compartmentalized into cristae membranes (CM, high MMP) and inner boundary membranes (IBM, lower MMP), separated by regulated crista junctions (CJ). Calcium influx promotes CJ opening through MICU1 reorganization, while enhanced TCA cycle activity increases cristae hyperpolarization.
Regulatory Signaling Pathways
Multiple signaling cascades converge on mitochondria to modulate MMP and direct metabolic partitioning:
Calcium Signaling: Mitochondrial calcium elevation stimulates dehydrogenases of the tricarboxylic acid (TCA) cycle, enhancing electron flow through the ETC and consequently increasing MMP, particularly in the cristae membranes. This hyperpolarization creates a favorable environment for oxidative phosphorylation [6].
Metabolic Checkpoints: Reduced MMP acts as a definitive signal for mitochondrial quality control, triggering PINK1 accumulation and Parkin recruitment, which marks depolarized mitochondria for mitophagy. This quality control mechanism ensures removal of dysfunctional units while preserving specialized populations [2].
Cell Cycle Integration: MMP serves as a retrograde signal to regulate cell cycle progression. Decreased MMP delays G1-to-S phase transition in both mtDNA-deficient (ρ0) and control cells, while experimental restoration of MMP rescues normal cell cycle timing [19].
Quantitative Assessment of MMP Gradients and Metabolic Output
Measurement Techniques and Parameters
Advanced microscopy approaches have enabled quantitative assessment of spatial MMP gradients and their correlation with metabolic function:
Table 2: Quantitative Parameters of MMP Gradients and Metabolic Output
| Parameter | Measurement Technique | Typical Values | Biological Significance |
|---|---|---|---|
| ΔΨC - ΔΨIBM Gradient | SIM/STED microscopy with TMRM | 10-30 mV difference | Indicates cristae specialization capacity |
| IBM Association Index | SIM with MTG/TMRM ratio imaging | 0.3-0.7 (concentration-dependent) | Quantifies CJ permeability status |
| P5CS Filamentation Threshold | Fluorescence recovery after photobleaching | ~-160 to -180 mV MMP | Trigger for reductive metabolism activation |
| Oxygen Consumption Rate (OCR) | Seahorse XF Analyzer | Cell type-dependent | Oxidative phosphorylation capacity |
| TMRM Concentration Range | Fluorescence microscopy | 1.35-81 nM (saturation-dependent distribution) | Optimized for gradient detection |
Correlation Between MMP Gradients and ATP Production
Multi-parameter correlation measurements have established a direct relationship between spatial MMP gradients and mitochondrial energetic output:
Cristae Hyperpolarization: Histamine-induced calcium elevation in HeLa and EA.hy926 cells produces a measurable decrease in IBM association index (from approximately 0.55 to 0.35 within 5 minutes), indicating cristae-specific hyperpolarization that correlates with enhanced ATP production [6].
Uncoupling Effects: Treatment with uncouplers such as BAM15 or CCCP dissipates MMP gradients and produces a transient accumulation of cells in G1 phase, demonstrating the cell cycle impact of lost MMP compartmentalization [19].
Metabolic State Transitions: Single-cell analysis reveals that MMP thresholds determine mitochondrial fate decisions, with fragments maintaining MMP > -150 mV typically re-fusing with the network, while those depolarized below -120 mV are targeted for mitophagy [2].
Experimental Approaches for Investigating MMP-Driven Specialization
Super-Resolution Imaging of Spatial MMP Gradients
Protocol: Structured Illumination Microscopy (SIM) for Spatial Membrane Potential Gradient Analysis
Principle: This method exploits the potential-dependent distribution of cationic fluorophores like TMRM between cristae membranes (higher MMP) and inner boundary membranes (lower MMP) at varying dye concentrations [6].
Procedure:
- Cell Preparation and Staining:
- Culture cells on high-precision glass-bottom dishes
- Load with 500 nM MitoTracker Green FM (MTG) for 30 minutes at 37°C
- Stain with 13.5 nM TMRM for additional 20 minutes
- Maintain TMRM throughout imaging for equilibrium distribution
Image Acquisition:
- Perform simultaneous dual-channel SIM imaging
- Use 488 nm excitation for MTG, 561 nm for TMRM
- Acquire z-stacks with 0.1 μm intervals covering mitochondrial volume
- Maintain constant temperature (37°C) and CO₂ (5%)
IBM Association Index Calculation:
- Apply automated Otsu threshold to MTG channel to define mitochondrial boundaries
- Generate eroded mask (CM region) and dilated mask (IBM region)
- Calculate ratio: Mean TMRM intensityIBM / Mean TMRM intensityCM
- Values <0.4 indicate cristae hyperpolarization; >0.6 reflect reduced gradient
ΔFWHM Analysis:
- Extract cross-sectional intensity profiles for both channels
- Calculate full width at half maximum for MTG and TMRM signals
- Determine difference (ΔFWHM = FWHMMTG - FWHMTMRM)
- Larger ΔFWHM indicates greater TMRM accumulation in cristae
Technical Considerations: This approach requires careful optimization of TMRM concentrations (1.35-81 nM range) to avoid saturation artifacts. Low concentrations (1.35-5.4 nM) preferentially accumulate in cristae, while high concentrations (40.5-81 nM) saturate cristae and highlight IBM localization [6].
Functional Respiration and MMP Correlation Protocol
Multi-Parameter Assessment of Metabolic Specialization
Principle: Simultaneous measurement of MMP and oxygen consumption rate (OCR) enables direct correlation of bioenergetic status with oxidative phosphorylation capacity [6].
Procedure:
- Experimental Setup:
- Seed cells in specialized multi-well plates for correlative microscopy
- Load with 20 nM TMRM and 200 nM MTG
- Acquire baseline SIM images for MMP gradient analysis
ATP Production Monitoring:
- Transfer plates to microscope with environmental control
- Inject FRET-based ATP indicator (ATeam) via transfection or microinjection
- Monitor mitochondrial ATP production via FRET efficiency
Metabolic Perturbation:
- Apply 100 μM histamine to stimulate mitochondrial calcium uptake
- Acquire time-lapse SIM images every 30 seconds for 10 minutes
- Correlate IBM association index changes with ATP production dynamics
Inhibition Controls:
- Pre-treat with 2 μM rotenone (Complex I inhibitor) or 1 μM antimycin A (Complex III inhibitor)
- Confirm specificity of MMP responses to ETC activity
Data Interpretation: Cristae hyperpolarization (decreased IBM association index) preceding increased ATP production indicates MMP-driven OXPHOS enhancement. Disassociation of these parameters suggests metabolic uncoupling or dysfunction.
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagents for Investigating MMP-Driven Metabolic Partitioning
| Reagent Category | Specific Examples | Concentration Range | Primary Function |
|---|---|---|---|
| MMP-Sensitive Dyes | TMRM, TMRE | 1-100 nM (gradient studies) | Quantitative MMP measurement |
| JC-1 | 1-5 μM (ratio metric imaging) | Discrimination of high/low MMP | |
| Rhodamine 123 | 10-100 nM (flow cytometry) | Population-level MMP assessment | |
| Mitochondrial Morphology Markers | MitoTracker Green FM | 100-500 nM | IMM reference independent of MMP |
| MitoTracker Red CMXRos | 20-100 nM | MMP-dependent accumulation | |
| Metabolic Modulators | BAM15 | 1-10 μM | Mitochondrial uncoupler |
| Oligomycin | 1-5 μM | ATP synthase inhibitor | |
| Rotenone | 0.5-2 μM | Complex I inhibitor | |
| Genetic Encoded Sensors | ATeam (FRET-based ATP sensor) | Plasmid transfection | Mitochondrial ATP production |
| mt-cpYFP | Lentiviral transduction | Matrix pH and ROS detection | |
| Specialized Equipment | Super-resolution microscope (SIM/STED) | N/A | Spatial MMP gradient analysis |
| Seahorse XF Analyzer | N/A | Correlative OCR and MMP |
Pathophysiological Implications and Therapeutic Targeting
Cancer Metabolism and Metabolic Plasticity
The concept of MMP-driven metabolic partitioning provides important insights into cancer cell adaptability and therapeutic resistance:
Acute Myeloid Leukemia (AML): Patients with high mitochondrial DNA content exhibit increased OXPHOS dependence and MMP, correlating with chemoresistance to cytarabine-based regimens. Targeting mitochondrial metabolism with metformin overcomes this resistance, though cells adapt by increasing glycolysis and NAD+ production, creating a therapeutic vulnerability to NAMPT inhibition [20].
Metabolic Switching in Solid Tumors: Cancer cells leverage MMP thresholds to allocate resources between ATP production and biomass generation. Elevated MMP promotes P5CS filamentation and reductive metabolism, supporting nucleotide and amino acid synthesis for rapid proliferation [2].
Neurodegenerative Disorders and Neuronal Plasticity
Neurons exhibit sophisticated spatial regulation of MMP to support specialized functional requirements:
Synaptic Plasticity: Changes in MMP coordinate structural remodeling at synapses by linking metabolic state to protein synthesis. Mitochondrial recruitment to dendrites enables localized energy production supporting synaptic function [2].
Metabolic Compartmentalization in Neurons: Distinct mitochondrial subpopulations with different MMP characteristics maintain ion gradients in axon terminals while supporting biosynthetic requirements in cell bodies, with disruption contributing to neurodegenerative pathophysiology [2].
Future Directions and Technical Innovations
Emerging methodologies promise to further elucidate the intricacies of MMP-driven metabolic specialization:
Nanoparticle-Based Sensors: Fluorescent carbon dots and other nanomaterials combined with MMP-sensitive probes enhance contrast and photostability for long-term mitochondrial tracking [18].
Two-Photon and NIR Probes: Advanced fluorophores like KMG-501 enable deeper tissue penetration for in vivo MMP gradient assessment [18].
Mitochondrial-Targeted Gene Editing: CRISPR-based approaches specifically targeted to mitochondria offer potential for mechanistic dissection of MMP regulation, though challenges remain in delivery and membrane penetration [21].
AI-Enhanced Image Analysis: Machine learning algorithms are increasingly employed to interpret complex fluorescence patterns and identify subtle MMP heterogeneity not detectable by conventional analysis [22].
The continuing refinement of these technical approaches will undoubtedly uncover additional dimensions of MMP-mediated metabolic partitioning, providing novel insights into cellular adaptation mechanisms and revealing new therapeutic opportunities for diseases characterized by metabolic dysregulation.
Delta-1-pyrroline-5-carboxylate synthase (P5CS) is a bifunctional enzyme that serves as the rate-limiting enzyme in the mitochondrial biosynthesis of proline and ornithine. Recent research has established that P5CS functions as a dynamic molecular switch that couples mitochondrial membrane potential (MMP) to cellular metabolic partitioning. Under conditions of elevated MMP, P5CS undergoes filamentation, driving a metabolic shift toward reductive biosynthesis while simultaneously promoting the physical segregation of mitochondrial subpopulations. This review comprehensively examines the structural mechanisms of P5CS filament formation, its regulation by bioenergetic cues, and the functional consequences for metabolic specialization. We present detailed experimental protocols for investigating P5CS filamentation and analyze emerging therapeutic implications for cancer and other diseases characterized by metabolic dysregulation.
The mitochondrial membrane potential (MMP), generated by the electron transport chain (ETC), represents a fundamental bioenergetic parameter classically known for driving ATP synthesis through protonmotive force [2] [23]. Beyond this canonical role, MMP is increasingly recognized as a dynamic signaling hub that integrates cellular energy status with broader physiological outputs, including calcium handling, reactive oxygen species production, and mitochondrial quality control [2]. A particularly significant advancement in understanding MMP function reveals its capacity to direct metabolic specialization through spatial and functional compartmentalization within mitochondrial networks.
Central to this metabolic partitioning is the enzyme P5CS, which occupies a critical nodal position at the intersection of oxidative and reductive metabolic pathways [24] [25]. P5CS catalyzes the initial, rate-limiting steps in the synthesis of proline and ornithine from glutamate, competing with the tricarboxylic acid (TCA) cycle for this common substrate [24]. Recent evidence demonstrates that P5CS acts as a molecular switch that senses elevated MMP and responds by forming filamentous structures, thereby triggering a metabolic shift toward reductive biosynthesis while physically segregating these functions into specialized mitochondrial subpopulations [2] [24] [26].
Structural Mechanisms of P5CS Filamentation
Molecular Architecture of P5CS Filaments
P5CS is a bifunctional enzyme containing two catalytic domains: an N-terminal glutamate kinase (GK) domain that catalyzes glutamate phosphorylation, and a C-terminal glutamyl phosphate reductase (GPR) domain that catalyzes the NADPH-dependent reduction of γ-glutamyl phosphate to glutamate-γ-semialdehyde [27]. Structural analyses using cryo-electron microscopy (cryo-EM) have revealed that P5CS filamentation occurs through a conserved mechanism involving tetrameric assembly as the fundamental structural unit [27] [28].
Table 1: Structural Characteristics of P5CS Filaments Across Species
| Species | Filament Architecture | Key Structural Interfaces | Ligand Regulation |
|---|---|---|---|
| Drosophila melanogaster | Double helix with GK tetramer core and GPR dimer periphery [27] | GK tetramerization interface; GPR dimerization interface; helical interfaces [27] | Glutamate strongly promotes filamentation; ATP and NADPH provide additional stabilization [27] |
| Arabidopsis thaliana (P5CS2) | Orthogonal helical assembly with hook-like interfaces [28] | GK tetramer core; perpendicular hook interactions between helical units [28] | Glutamate and ATP equally promote filament formation; maximal length with all substrates [28] |
| Arabidopsis thaliana (P5CS1) | Non-perpendicular helical twist (~80°) [28] | Similar GK tetramer; distinct GPR positioning compared to P5CS2 [28] | NADPH alone can induce filaments; differential regulation compared to P5CS2 [28] |
Structural studies of Drosophila P5CS have demonstrated that filaments assemble through a double-helical arrangement with a diameter of approximately 180Å, where GK domain tetramers form the filament core while GPR dimers arrange in left-handed helices around the central axis [27]. The P5CS tetramer serves as the fundamental building block, with filament stability maintained through multiple interfaces including specific interactions between GK domains and connecting helical elements [27].
Functional Consequences of Filamentation
P5CS filament formation directly enhances catalytic efficiency through substrate channeling and allosteric regulation. Structural analyses reveal that filamentation enables coordinated conformational changes between GK and GPR domains that facilitate the transfer of reaction intermediates [27]. Point mutations that disrupt key filament interfaces dramatically reduce enzymatic activity without affecting substrate binding, confirming that filamentation is essential for optimal catalytic function [27].
Table 2: Functional Impact of P5CS Filamentation on Enzyme Activity
| Experimental Manipulation | Effect on Filamentation | Impact on Enzymatic Activity | Cellular Consequences |
|---|---|---|---|
| Point mutations disrupting filament interfaces (Drosophila) [27] | Complete disruption of filament assembly | ~70-80% reduction in catalytic activity [27] | Impaired proline biosynthesis; reduced cell proliferation under nutrient stress [27] |
| Glutamate supplementation [27] [28] | Promotes filament formation and elongation | Significant increase in proline production [27] [28] | Enhanced reductive biosynthesis; metabolic shift toward anabolism [24] |
| ATP and NADPH availability [28] | Synergistic effect with glutamate on filament stability | Maximal catalytic efficiency under complete substrate conditions [28] | Coordination between energy status and biosynthetic output [24] [28] |
The structural data collectively support a model wherein P5CS filamentation creates a specialized metabolic compartment that enhances catalytic efficiency through substrate channeling and allosteric communication between enzyme domains [27] [28]. This structural reorganization provides a mechanistic basis for the observed metabolic switching behavior in response to bioenergetic cues.
MMP Regulation of P5CS and Metabolic Specialization
MMP as a Primary Regulator of P5CS Activity
The mitochondrial membrane potential serves as a key upstream regulator of P5CS filamentation and activity. Under conditions of elevated MMP relative to baseline, P5CS enzymatic activity is enhanced, promoting the formation of filamentous assemblies that drive reductive biosynthesis [2]. Conversely, reduced MMP inhibits P5CS filamentation and shifts mitochondrial function toward core energetic processes such as oxidative phosphorylation (OXPHOS) [2]. This MMP sensitivity enables mitochondria to dynamically adjust their metabolic output based on bioenergetic status.
The mechanistic relationship between MMP and P5CS filamentation can be visualized as a regulatory circuit that controls metabolic partitioning:
Diagram 1: MMP Regulation of P5CS and Metabolic Partitioning. Elevated MMP induces P5CS filamentation, enhancing reductive biosynthesis and driving the formation of specialized mitochondrial subpopulations.
Formation of Metabolically Specialized Mitochondrial Subpopulations
Under conditions of high cellular ATP demand or nutrient stress, P5CS filamentation drives the physical segregation of mitochondria into distinct subpopulations with specialized functions [24] [25]. This compartmentalization creates a "division of labor" where:
- Biosynthetic Mitochondria: Contain filamentous P5CS, exhibit elevated MMP, lack cristae, and show reduced or absent ATP synthase expression. These mitochondria specialize in reductive biosynthesis of proline and ornithine [24] [26].
- Oxidative Mitochondria: Lack P5CS filaments, contain well-developed cristae with abundant ATP synthase, and specialize in OXPHOS-driven ATP production [24] [25].
This metabolic specialization is reversible and dynamically adjusts to changing cellular conditions. When proline or ornithine is supplemented externally, P5CS filaments disassemble, demonstrating the responsive nature of this compartmentalization system [24] [29].
Experimental Approaches for Investigating P5CS Filamentation
Methodologies for Inducing and Monitoring P5CS Filamentation
Table 3: Experimental Protocols for P5CS Filamentation Studies
| Method | Protocol Details | Key Measurements | Applications |
|---|---|---|---|
| Metabolic Stress Induction [24] | Culture cells in galactose medium or glucose-deficient medium to force OXPHOS dependence; Treatment with D-lactate (10-20mM) to stimulate ETC activity | intracellular proline quantification; NADH/NAD+ ratio; oxygen consumption rate [24] | Investigating P5CS response to bioenergetic stress; Linking filamentation to metabolic output |
| Filament Visualization [24] [27] | Immunofluorescence staining of endogenous P5CS; Expression of P5CS-GFP fusions; Correlative light and electron microscopy (CLEM) | Percentage of mitochondrial area containing P5CS; Filament length and distribution; Cristae ultrastructure [24] | Structural characterization of filaments; Spatial relationship to mitochondrial architecture |
| Genetic Manipulation [24] [26] | CRISPR-Cas9 knockout of P5CS; Expression of filament-disrupting mutants; Modulation of mitochondrial dynamics (DRP1/MFN knockout) | Proline auxotrophy assessment; Rescue experiments with CRISPR-resistant constructs; Metabolic tracing with [U-13C] glutamine [24] | Establishing necessity of filamentation; Testing functional consequences |
| Structural Analysis [27] [28] | Cryo-EM of purified P5CS with varying substrate combinations (glutamate, ATP, NADPH); Negative staining EM for initial screening | 3D reconstruction of filament structures (3.1-4.3Å resolution); Ligand binding site identification; Conformational changes [27] | Molecular mechanism of filamentation; Substrate channeling mechanisms |
The Scientist's Toolkit: Essential Research Reagents
Table 4: Key Reagents for P5CS Filamentation Research
| Reagent/Cell Line | Specific Application | Function in Experimental Design |
|---|---|---|
| Galactose Medium [24] | Force OXPHOS dependence | Increases mitochondrial ATP demand, inducing P5CS filament formation |
| [U-13C] Glutamine [24] | Metabolic flux analysis | Traces glutamate utilization into TCA cycle (oxidative) vs. proline/ornithine (reductive) pathways |
| P5CS Knockout MEFs [24] | Establish P5CS necessity | Demonstrate proline auxotrophy and provide background for rescue experiments |
| CRISPR-resistant P5CS constructs [24] | Complementation tests | Validate phenotype specificity and test structure-function relationships |
| Anti-P5CS antibodies [24] | Endogenous protein localization | Visualize native P5CS distribution and filament formation under different conditions |
| D-lactate [24] | ETC stimulation | Directly enhances electron transport chain activity without uncoupling |
| FCCP (uncoupler) [24] | Dissociate ATP production from ETC activity | Increases ETC activity without producing ATP, tests MMP-independent effects |
The experimental workflow for a comprehensive investigation of P5CS filamentation typically integrates multiple methodological approaches:
Diagram 2: Integrated Experimental Workflow for P5CS Filamentation Research. Comprehensive investigation requires complementary approaches including metabolic manipulation, genetic tools, visualization techniques, functional assays, and structural analysis.
Functional Consequences and Pathophysiological Implications
Mitochondrial Dynamics in Metabolic Partitioning
The formation of metabolically specialized mitochondrial subpopulations depends critically on balanced mitochondrial fusion and fission processes [24] [25] [26]. Experimental evidence demonstrates that:
- Fission-deficient cells (DRP1−/−) exhibit elongated mitochondria, fail to separate P5CS from ATP synthase, and show significantly reduced proline synthesis despite maintaining efficient OXPHOS [24] [26].
- Fusion-deficient cells (MFN1/2−/−) maintain proline synthesis capability but display impaired respiratory activity and fail to properly segregate P5CS from ATP synthase complexes [24] [25].
These findings establish that mitochondrial dynamics are not merely quality control mechanisms but actively participate in organizing metabolic compartmentalization. The fusion-fission cycle enables the physical separation of P5CS-containing biosynthetic mitochondria from OXPHOS-specialized mitochondria, allowing simultaneous operation of competing metabolic pathways [24] [25].
Implications for Disease Pathogenesis and Therapeutics
The discovery of P5CS-mediated metabolic partitioning has significant implications for understanding disease mechanisms and developing targeted therapies:
Cancer Metabolism: Pancreatic ductal adenocarcinoma cells exhibit prominent P5CS clustering in distinct mitochondrial subpopulations, while adjacent normal tissue lacks this compartmentalization [24] [26]. This adaptation enables cancer cells to maintain both energy production and biosynthetic capacity despite nutrient scarcity in the tumor microenvironment [24] [26]. Targeting P5CS filamentation may represent a novel therapeutic strategy to disrupt cancer metabolic adaptability.
Neurological Disorders: Recent evidence suggests parallels between mitochondrial alterations in migraine models and P5CS-related mitochondrial subsets, including cristae disruption and metabolic reprogramming [30]. The involvement of UCP4 polymorphisms in Alzheimer's disease and frontotemporal dementia further underscores the importance of MMP regulation in neuronal health [2] [30].
Metabolic Diseases: UCP3 polymorphisms associated with obesity demonstrate the physiological relevance of regulated MMP dissipation, with potential connections to P5CS-mediated metabolic partitioning under conditions of nutrient excess [2].
P5CS filamentation represents a sophisticated molecular mechanism that directly couples mitochondrial membrane potential to cellular metabolic fate decisions. By sensing elevated MMP and responding with structural reorganization into functional filaments, P5CS acts as a molecular switch that promotes reductive biosynthesis while simultaneously driving the physical compartmentalization of mitochondrial populations. This "divide and conquer" strategy enables cells to simultaneously maintain oxidative phosphorylation and reductive biosynthesis—metabolic pathways that would otherwise compete for shared substrates.
The emerging understanding of P5CS filamentation opens new avenues for therapeutic intervention in diseases characterized by metabolic dysregulation, particularly cancers that exploit this adaptability to survive in nutrient-poor environments. Future research directions should focus on elucidating the precise molecular mechanisms of MMP sensing by P5CS, developing pharmacological modulators of filamentation, and exploring the tissue-specific manifestations of this metabolic partitioning in health and disease. As a central integrator of bioenergetic status and biosynthetic output, P5CS represents both a fundamental mechanism of metabolic control and a promising target for precision metabolic therapeutics.
In muscle cells, mitochondria are not randomly distributed but are organized into distinct subpopulations based on their subcellular location. The two primary populations are the subsarcolemmal mitochondria (SSM), situated directly beneath the plasma membrane (sarcolemma), and the interfibrillar mitochondria (IFM), located between the contractile myofibrils [31] [32] [33]. This spatial organization is not merely structural; it underpins significant functional heterogeneity. These mitochondrial subpopulations differ in their biochemical properties, respiratory capacities, and responsiveness to metabolic stress and pharmacological agents [31] [32]. Within the context of mitochondrial membrane potential (ΔΨm)—the key component of the protonmotive force that drives ATP synthesis—this compartmentalization facilitates metabolic specialization and partitioning, allowing muscle cells to meet localized energy demands and respond adaptively to physiological and pathological challenges [2] [6] [33].
Structural and Biochemical Distinctions
The spatial segregation of SSM and IFM is accompanied by clear morphological and molecular differences, which form the basis for their functional specialization.
Ultrastructural and Morphological Differences
Electron microscopy reveals that in their native cellular environment, IFM are typically elongated and rod-shaped and contain a matrix with greater electron density. In contrast, SSM have a more rounded shape and a less electron-dense, "lighter" matrix [31]. In skeletal muscle, IFM form complex, highly branched networks that wrap around the I-band of the sarcomere, a morphology that maximizes the surface area-to-volume ratio to facilitate rapid ATP diffusion to the myofibrillar ATPases [33]. SSM, often found clustered between the sarcolemma and the myofibrils, are generally larger and less branched [33].
Proteomic and Enzymatic Variations
Biochemically, IFM consistently demonstrate a higher specific activity for key metabolic enzymes. The table below summarizes established biochemical differences between SSM and IFM isolated from cardiac muscle.
Table 1: Biochemical Properties of Subsarcolemmal and Interfibrillar Mitochondria
| Parameter | Subsarcolemmal Mitochondria (SSM) | Interfibrillar Mitochondria (IFM) | References |
|---|---|---|---|
| Succinate Dehydrogenase Activity | Lower | Higher | [32] |
| Citrate Synthase Activity | Lower | Higher | [32] |
| Carnitine Palmitoyltransferase Activity | Similar | Similar | [32] |
| α-Glycerophosphate Dehydrogenase Activity | Similar | Similar | [32] |
| Oxidation Rate of Various Substrates | Lower (~1.5x slower) | Higher | [32] |
| Respiratory Capacity | Lower | Higher | [31] |
| Protein Composition | Distinct | Distinct | [31] |
These biochemical profiles indicate that IFM possess a greater intrinsic capacity for oxidative metabolism, which is consistent with their primary role in directly fueling muscle contraction.
Functional Heterogeneity and Metabolic Specialization
The structural and enzymatic differences between SSM and IFM translate into specialized physiological roles, particularly in energy metabolism, calcium handling, and response to stress.
Bioenergetic Capacity and Membrane Potential
The higher enzymatic activity and oxidation rates in IFM contribute to a greater capacity for ATP production [32]. Furthermore, the mitochondrial membrane potential (ΔΨm), a critical indicator of mitochondrial energetic status and health, is not uniform across the cell. Recent super-resolution microscopy studies reveal that the inner mitochondrial membrane itself is compartmentalized into the cristae membrane (CM) and inner boundary membrane (IBM), which can maintain distinct electrical potentials (ΔΨC and ΔΨIBM, respectively) [6]. The CM, housing the proton-pumping complexes of the electron transport chain, typically exhibits a higher (more negative) membrane potential than the IBM [6]. This gradient is dynamically regulated; for instance, an increase in mitochondrial calcium uptake stimulates the TCA cycle and enhances proton pump activity, leading to a relative hyperpolarization of the cristae [6]. This intricate regulation of ΔΨm gradients ensures efficient ATP synthesis and links cellular signaling to energy production.
Differential Roles in Fatty Acid Metabolism
The functional specialization of mitochondrial subpopulations is particularly evident in skeletal muscle fatty acid metabolism. Studies show that oxidation rates of fatty acids are substantially higher in mitochondria isolated from red oxidative muscle fibers compared to white glycolytic fibers [34]. Moreover, SSM and IFM exhibit differential sensitivity to malonyl-CoA, a potent inhibitor of the fatty acid transport enzyme CPT1β. In one study, malonyl-CoA almost completely abolished fatty acid oxidation in SSM and IFM from white gastrocnemius muscle, but only partially inhibited oxidation in mitochondria from red gastrocnemius muscle [34]. Endurance training further accentuates this functional divergence, increasing palmitate oxidation rates more significantly in SSM (100% increase) than in IFM (46% increase) [34].
Differential Responsiveness to Stress and Pharmacological Agents
A critical aspect of their functional heterogeneity is their differential vulnerability and responsiveness to stimuli. SSM are consistently reported to be more vulnerable to ischemic injury and calcium overload than IFM [31]. This susceptibility makes them a prime target for protective therapeutics.
Research on diazoxide, a cardioprotective agent that targets mitochondrial potassium channels, demonstrates this principle. Diazoxide was found to be significantly more effective at protecting SSM against calcium-induced mitochondrial permeability transition and at restoring calcium-inhibited oxidative phosphorylation [31]. This indicates that SSM are the preferred target for this protective drug, highlighting how understanding mitochondrial subpopulations can inform targeted therapeutic strategies [31].
Experimental Methodologies for Isolation and Analysis
The study of SSM and IFM relies on specialized protocols for their separation and subsequent functional assessment.
Differential Isolation of SSM and IFM
The standard method for isolating distinct mitochondrial populations from cardiac muscle involves sequential mechanical and enzymatic digestion steps [31] [32].
Diagram Title: Mitochondrial Isolation Workflow
Key Steps Explained:
- Tissue Homogenization: Ventricular tissue is homogenized using a mechanical tissue processor like a Polytron. This process primarily ruptures the outer sarcolemma, releasing the SSM population [31] [32].
- SSM Isolation: The homogenate is subjected to differential centrifugation to pellet the released SSM [31].
- IFM Release: The remaining tissue pellet, now depleted of SSM but still containing the IFM entrenched within the myofibrillar matrix, is treated with a protease such as Nagarse. This enzymatic digestion breaks down the myofibrillar structures, releasing the IFM [31] [32].
- IFM Isolation: A second round of differential centrifugation is performed to pellet the purified IFM [31].
The integrity of the isolated mitochondria is typically confirmed using electron microscopy [31].
Core Functional Assays
Once isolated, the functional capacity of SSM and IFM is characterized using a suite of bioenergetic assays.
Table 2: Key Functional Assays for Mitochondrial Subpopulations
| Assay Parameter | Methodology | Key Insight | Common Reagents/Tools |
|---|---|---|---|
| Respiration | Clark-type oxygen electrode measuring O₂ consumption in response to substrates (e.g., succinate) and ADP [31]. | Direct measure of oxidative phosphorylation capacity. | Succinate, Pyruvate, ADP, MOPS buffer, KCl. |
| Membrane Potential (ΔΨm) | Tetraphenylphosphonium (TPP⁺)-sensitive electrode or potentiometric dyes (e.g., TMRM) [31] [6]. | Indicator of mitochondrial energetic state and protonmotive force. | TPP⁺ electrode, TMRM, Tetramethylrhodamine Methyl Ester. |
| Calcium Retention Capacity | Ca²⁺-sensitive minielectrode measuring cumulative Ca²⁺ uptake via sequential pulses until permeability transition [31]. | Assesses susceptibility to Ca²⁺-induced dysfunction and cell death. | Ca²⁺ electrode, Calcium chloride. |
| ATP Synthesis | HPLC or enzymatic coupled assays (e.g., hexokinase/glucose-6-phosphate dehydrogenase) to quantify ATP production [31]. | Direct measurement of functional energy output. | HClO₄, K₂CO₃, Hexokinase, Glucose-6-Phosphate Dehydrogenase, NADP⁺. |
| Citrate Synthase Activity | Spectrophotometric assay measuring conversion of acetyl-CoA and oxaloacetate [31]. | Marker of mitochondrial content and integrity. | 5,5′-Dithiobis-(2-nitrobenzoic acid), Acetyl-Co-A, Oxaloacetate. |
Research Toolkit: Essential Reagents and Materials
Table 3: Research Reagent Solutions for Mitochondrial Studies
| Reagent/Material | Function in Research | Application Example |
|---|---|---|
| Polytron Homogenizer | Mechanical disruption of tissue for selective release of SSM [31] [32]. | Initial step in differential isolation protocol. |
| Nagarse (Protease) | Enzymatic digestion of the myofibrillar matrix to release IFM [31] [32]. | Second step for IFM isolation after SSM removal. |
| Tetramethylrhodamine Methyl Ester (TMRM) | Potentiometric fluorescent dye for measuring mitochondrial membrane potential (ΔΨm) [6]. | Live-cell imaging of ΔΨm gradients and cristae-specific hyperpolarization. |
| Tetraphenylphosphonium (TPP⁺) Electrode | Direct quantitative measurement of ΔΨm in isolated mitochondrial suspensions [31]. | Functional characterization of bioenergetics in SSM vs. IFM. |
| Ca²⁺-Selective Minielectrode | Real-time monitoring of mitochondrial calcium uptake and flux [31]. | Determining Calcium Retention Capacity (CRC). |
| Clark-type Oxygen Electrode | Measurement of oxygen consumption as a direct readout of mitochondrial respiratory function [31]. | Assessing oxidative phosphorylation capacity with different substrates. |
| Diazoxide | Pharmacological activator of mitochondrial potassium channels; a cardioprotective agent [31]. | Probing differential drug responsiveness between SSM and IFM. |
Implications for Drug Development and Therapeutic Targeting
The distinct properties of mitochondrial subpopulations present both challenges and opportunities for pharmacology. The observed differential drug responsiveness, as seen with diazoxide, underscores that mitochondria are not a homogeneous target [31]. Effective therapeutic strategies may need to account for subcellular localization and the unique vulnerability of specific mitochondrial pools.
The higher ΔΨm of the cristae membrane and the general negative potential of the mitochondrial matrix are exploited for drug delivery. Delocalized lipophilic cations (DLCs), such as triphenylphosphonium (TPP⁺), can be conjugated to drugs to drive their accumulation within mitochondria [35] [36]. This strategy has been successfully used to develop mitochondrial-targeted antioxidants like MitoQ, which accumulates several hundred-fold inside mitochondria and has shown efficacy in models of ischemia-reperfusion injury and is in clinical trials for Parkinson's disease [35] [36].
Diagram Title: Targeted Drug Delivery to Mitochondria
Given the heightened vulnerability of SSM to stress, such targeted delivery systems could be particularly valuable for delivering protective agents directly to this susceptible population, offering a refined approach to treating conditions like heart failure, metabolic myopathies, and neurodegenerative diseases [31] [35] [36].
Mitochondrial quality control is a critical process for cellular homeostasis, integrating mitochondrial dynamics—fission and fusion—with the selective autophagy of mitochondria, known as mitophagy. Central to this integration is the mitochondrial membrane potential (ΔΨm), which acts as a key signaling hub and fate determinant for mitochondria following a fission event. This whitepaper delineates the established and emerging principles of how ΔΨm thresholds govern the decision for a mitochondrion to undergo repair via fusion or elimination via mitophagy. We synthesize current data into actionable guidelines and protocols, providing researchers with a framework to investigate these processes in the context of metabolic specialization and partitioning, with direct implications for therapeutic development in neurodegenerative diseases, metabolic syndromes, and aging.
The mitochondrial network is not a static entity but a dynamic population undergoing constant remodeling. The life cycle of an individual mitochondrion is punctuated by fission and fusion events, and its fate is critically dependent on its bioenergetic status, principally reflected by its ΔΨm [37]. The ΔΨm, generated by the electron transport chain, is not merely a proxy for energy production capacity but a dynamic signaling entity that regulates mitochondrial function, communication, and quality control [2]. Following a fission event, the resulting daughter mitochondria often exhibit asymmetric ΔΨm, creating a "fate decision" point [37]. The daughter unit that retains a high ΔΨm is fusion-competent and can re-join the network, effectively diluting any minor damage. Conversely, the depolarized daughter mitochondrion becomes fusion-incompetent and is targeted for degradation via mitophagy [37]. This review explores the quantitative thresholds and molecular mechanisms underlying this binary fate decision, framing it within the broader physiological context of metabolic specialization.
Core Mechanisms: The Interplay of Dynamics and Mitophagy
Fission Generates the Substrate for Quality Control
Contrary to spontaneous depolarization, mitochondrial fission is a primary mechanism for generating depolarized mitochondrial units. Prolonged tracking of individual mitochondria reveals remarkable ΔΨm stability during their solitary phase; significant depolarization is most frequently associated with fission events [37]. Electron microscope tomography and direct ΔΨm measurements show that fission often yields asymmetric daughter mitochondria, with one unit experiencing a transient or sustained depolarization [37]. This fission-induced depolarization creates the pre-autophagic pool—a population of depolarized, fusion-incompetent mitochondria that are primed for elimination.
Table 1: The Effects of Manipulating Mitochondrial Dynamics on Mitophagy
| Manipulation | Cell Type | Effect on Mitophagy | Key Findings |
|---|---|---|---|
| Fis1 RNAi | INS1 β-cells | ~70% reduction [37] | Specific reduction in autophagosomes containing mitochondria; no change in ER-phagy. |
| Drp1K38A (DN) | INS1 β-cells | ~75% reduction [37] | Suppresses mitochondrial autophagy specifically. |
| Fis1 Overexpression | HeLa, INS1 cells | Reduces mitochondrial mass [37] | Promotes mitochondrial loss, consistent with increased mitophagy. |
| Opa1 Overexpression | INS1 β-cells | ~63% reduction [37] | Inhibits mitophagy, likely by promoting fusion and rescuing depolarized units. |
The PINK1-Parkin Pathway and the ΔΨm Threshold
A core mechanism linking ΔΨm to mitophagy is the PINK1-Parkin pathway. On healthy, polarized mitochondria, PTEN-induced putative kinase 1 (PINK1) is continuously imported and degraded. Upon ΔΨm loss, this import is halted, leading to PINK1 accumulation on the outer mitochondrial membrane [2]. PINK1 then recruits the E3 ubiquitin ligase Parkin, which ubiquitinates mitochondrial surface proteins, marking the organelle for autophagic degradation via LC3 binding [2]. This pathway establishes ΔΨm as the primary signal for mitophagy initiation.
Metabolic Specialization and Compartmentalization
Beyond quality control, ΔΨm gradients facilitate metabolic specialization within the mitochondrial network. Recent research indicates that the inner mitochondrial membrane is not uniform; the cristae membrane (CM) can maintain a higher ΔΨm (ΔΨC) than the inner boundary membrane (IBM) [6]. This compartmentalization is regulated by cristae junctions (CJ) and allows for specialized functions. Furthermore, distinct mitochondrial subpopulations can be biased toward oxidative (ATP-producing) or reductive (biosynthetic) metabolism [2]. For instance, elevated ΔΨm can enhance the filamentation and activity of pyrroline-5-carboxylate synthase (P5CS), a key enzyme in proline biosynthesis, thereby promoting reductive metabolism [2]. This implies that ΔΨm thresholds may not only decide organelle fate but also direct metabolic programming.
The following diagram illustrates the core signaling pathway that integrates these processes, from mitochondrial fission to fate determination.
Quantitative Data and Experimental Evidence
The molecular interplay described above is supported by quantitative data from genetic and observational studies.
Table 2: Key Experimental Findings on MMP Thresholds and Fate Decisions
| Experimental Model | Key Finding | Quantitative/Measured Outcome |
|---|---|---|
| INS1 & COS7 Cells [37] | Fission frequently generates daughters with asymmetric ΔΨm. | ~5% of fission events produce sustained depolarization; majority show transient ΔΨm difference >5mV. |
| Δdnm1Δmgm1 Yeast [38] | Simultaneous loss of fission and fusion reduces mitophagy and shortens lifespan. | Replicative lifespan shortened to 10.8 gens (vs. 18.0 in WT); mitophagy impaired despite filamentous morphology. |
| HeLa & EA.hy926 Cells [6] | Cristae membrane (CM) has a higher ΔΨm than inner boundary membrane (IBM). | IBM association index and ΔFWHM methods quantify compartment-specific ΔΨm; Ca²⁺ influx hyperpolarizes CM. |
| Theoretical/Mechanistic [2] | MMP dictates protein import and influences metabolic enzyme partitioning. | Elevated MMP promotes P5CS filamentation, shifting mitochondria toward reductive biosynthesis. |
Methodologies for Investigating MMP and Quality Control
Protocol: Measuring Spatial Mitochondrial Membrane Potential Gradients
This protocol, adapted from scientific research, details the use of potentiometric dyes and super-resolution microscopy to visualize ΔΨm gradients across mitochondrial sub-compartments [6].
- Objective: To quantify differences in ΔΨm between the cristae membrane (CM) and inner boundary membrane (IBM).
- Principle: The distribution of the ΔΨm-sensitive dye TMRM is concentration-dependent. At low concentrations, it accumulates preferentially in the higher-potential CM, while at high concentrations, it saturates the CM and stains the IBM more prominently.
- Key Reagents:
- Tetramethylrhodamine Methyl Ester (TMRM): A cell-permeant, cationic fluorescent dye that accumulates in the mitochondrial matrix in a ΔΨm-dependent manner. Use a range of low concentrations (1.35 - 5.4 nM).
- MitoTracker Green FM (MTG): A dye that covalently binds to mitochondrial proteins, largely independent of ΔΨm, serving as a morphological reference for the inner mitochondrial membrane.
- Procedure:
- Cell Staining: Co-stain cells with 500 nM MTG and a low concentration of TMRM (e.g., 2.7 nM) in serum-free medium for 15-30 minutes at 37°C.
- Image Acquisition: Perform simultaneous dual-channel imaging using Structured Illumination Microscopy (SIM) or STED to achieve super-resolution.
- Image Analysis:
- IBM Association Index: An automated method using the MTG channel to define mitochondrial boundaries. The fluorescence intensity of TMRM in the IBM region is divided by its intensity in the CM region.
- ΔFWHM Method: A semi-automated method comparing the Full Width at Half Maximum (FWHM) of cross-sectional intensity profiles for MTG and TMRM. A larger ΔFWHM indicates greater TMRM accumulation in the CM.
- Interpretation: A decrease in the IBM association index or ΔFWHM after a stimulus (e.g., Ca²⁺ elevation) indicates relative hyperpolarization of the CM, linking bioenergetics to ultrastructure.
Protocol: Assessing Mitophagy Flux via PINK1/Parkin Translocation
- Objective: To monitor the initiation of mitophagy in response to ΔΨm loss.
- Principle: Depolarization prevents PINK1 import, leading to its accumulation on the OMM and subsequent recruitment of Parkin.
- Key Reagents:
- Cells expressing GFP-Parkin: A fluorescently tagged Parkin construct allows visualization of its cytosolic-to-mitochondrial translocation.
- ΔΨm-dissipating agents: Carbonyl cyanide m-chlorophenyl hydrazone (CCCP) or Oligomycin/Antimycin A can be used as positive controls to induce depolarization.
- Mitochondrial marker: A dye like MitoTracker Deep Red to label the total mitochondrial population.
- Procedure:
- Transfert cells with GFP-Parkin.
- Treat cells with the experimental compound or vehicle control.
- Fix and stain with a mitochondrial marker, or perform live-cell imaging.
- Image using confocal or high-content microscopy.
- Quantify the degree of Parkin colocalization with mitochondria.
- Interpretation: Increased colocalization of Parkin with mitochondria indicates activation of the mitophagy pathway in response to reduced ΔΨm.
The experimental workflow for investigating these processes, from cell preparation to data analysis, is summarized below.
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagents for Investigating MMP, Dynamics, and Mitophagy
| Reagent / Tool | Category | Primary Function in Research |
|---|---|---|
| TMRM / TMRE | Potentiometric Dye | ΔΨm-sensitive probe for live-cell imaging; distribution indicates polarization state [37] [6]. |
| MitoTracker Green FM | Mitochondrial Stain | ΔΨm-independent stain for labeling mitochondrial morphology as a spatial reference [6]. |
| CCCP | Pharmacological Agent | Protonophore that uncouples OXPHOS, dissipates ΔΨm, and induces PINK1/Parkin-mediated mitophagy. |
| Drp1 Inhibitors (e.g., Mdivi-1) | Small Molecule Inhibitor | Inhibits mitochondrial fission, used to test the necessity of fission for mitophagy [37]. |
| Opa1 Expression Constructs | Genetic Tool | Overexpression promotes fusion, rescuing depolarized mitochondria and suppressing mitophagy [37]. |
| GFP-Parkin | Fluorescent Biosensor | Live-cell reporting of mitophagy initiation via translocation from cytosol to mitochondria. |
The integration of mitochondrial fission, fusion, and mitophagy through ΔΨm thresholds represents a sophisticated cellular quality control system. The binary fate decision post-fission ensures the removal of damaged units while allowing for content mixing and metabolic flexibility. The emerging concepts of intramitochondrial ΔΨm gradients and metabolic partitioning add layers of complexity, suggesting that ΔΨm not only determines life-or-death decisions but also fine-tunes functional output. Future research must focus on precisely defining the quantitative ΔΨm thresholds for these decisions in different cell types and disease states, and on elucidating the molecular sensors that interpret these bioenergetic signals. Understanding these mechanisms in greater depth will open new avenues for therapeutic intervention in the vast range of pathologies associated with mitochondrial dysfunction.
Measuring the Charge: Advanced Techniques for Assessing MMP in Research and Drug Discovery
The mitochondrial membrane potential (ΔΨm) is a fundamental parameter in cellular bioenergetics, serving as a primary indicator of mitochondrial health and functional status. Generated by the proton-pumping activity of the electron transport chain, this electrochemical gradient across the inner mitochondrial membrane is essential for ATP synthesis, calcium homeostasis, and reactive oxygen species regulation [39] [40]. In the context of metabolic specialization and partitioning research, ΔΨm provides critical insights into how cells allocate energy resources, adapt to metabolic demands, and maintain physiological function under varying conditions. The accurate measurement of ΔΨm is therefore crucial for understanding cellular metabolism in health and disease, particularly in neurodegenerative disorders, cancer, and metabolic syndromes where mitochondrial dysfunction is a hallmark feature [41] [42].
Potentiometric fluorescent dyes represent the cornerstone technology for assessing ΔΨm in live cells. Among these, tetramethylrhodamine methyl ester (TMRM) and 5,5',6,6'-tetrachloro-1,1',3,3'-tetraethylbenzimidazolocarbocyanine iodide (JC-1) have emerged as two of the most widely utilized and technically distinct probes. These dyes enable researchers to track dynamic changes in mitochondrial physiology, from spontaneous fluctuations in individual mitochondria to population-wide shifts during apoptosis or metabolic adaptation [39] [43]. This technical guide provides a comprehensive examination of TMRM and JC-1 assays, detailing their fundamental properties, methodological considerations, and implementation across scales from traditional bench microscopy to high-throughput screening (HTS) platforms.
Fundamental Properties of TMRM and JC-1
Chemical and Optical Characteristics
TMRM and JC-1, while both targeting ΔΨm, operate through distinct mechanisms and offer complementary information about mitochondrial status.
TMRM is a cationic, lipophilic dye that accumulates in the mitochondrial matrix in a potential-dependent manner according to the Nernst equation [42]. In its most reliable application, TMRM is used in non-quenching mode at low concentrations (typically 5-50 nM), where fluorescence intensity directly correlates with ΔΨm – depolarization leads to dye redistribution and decreased fluorescence [42] [44]. TMRM exhibits excitation/emission maxima at approximately 548/573 nm [45]. Its key advantage lies in minimal perturbation of mitochondrial function, as it demonstrates low binding to mitochondrial membranes and negligible inhibition of the electron transport chain [42] [45].
JC-1 employs a unique dual-emission mechanism that provides an intrinsic rationetric measurement. At low membrane potentials or low concentrations, JC-1 exists as monomers that fluoresce green (emission ~529 nm). As ΔΨm increases, the dye accumulates in mitochondria and forms J-aggregates that emit red fluorescence (~590 nm) [40] [43]. This potential-dependent spectral shift enables quantitative assessment through the red/green fluorescence ratio, which is largely independent of mitochondrial morphology, dye concentration, and cell size [40] [46]. JC-1 has excitation maxima at 514 nm (for both forms) with dual emission peaks at 529 nm (monomer) and 590 nm (J-aggregate) [40].
Table 1: Fundamental Properties of TMRM and JC-1
| Property | TMRM | JC-1 |
|---|---|---|
| Detection Mechanism | Single-emission, intensity-based | Dual-emission, rationetric |
| Excitation/Emission | 548/573 nm [45] | 514/529 nm (monomer), 514/590 nm (J-aggregate) [40] |
| Signal Change with Depolarization | Decreased fluorescence [42] [44] | Decreased red/green ratio [40] [43] |
| Advantages | Minimal mitochondrial perturbation; Suitable for kinetics [42] [45] | Internal ratio control; Insensitive to dye loading [40] [46] |
| Limitations | Sensitive to loading conditions; Requires careful controls [42] | More complex data analysis; Potential dye precipitation [46] |
Mechanism of Action and Cellular Distribution
The operational principles of these dyes can be visualized through their distinct mechanisms of mitochondrial localization and signal generation:
Diagram 1: Mechanisms of TMRM and JC-1 Mitochondrial Accumulation and Signal Generation
Both dyes are lipophilic cations that readily cross cellular membranes and accumulate in the negatively charged mitochondrial matrix [43] [45]. For TMRM, this distribution follows a simple equilibrium determined by the Nernst potential, resulting in higher concentrations and fluorescence intensity in polarized mitochondria [42]. JC-1 exhibits a concentration-dependent equilibrium where the monomeric form predominates at low intramitochondrial concentrations (low ΔΨm), while J-aggregates form at higher concentrations (high ΔΨm) achieved through potential-dependent accumulation [40] [43]. During apoptosis or metabolic stress, one of the earliest events is mitochondrial depolarization, leading to TMRM redistribution throughout the cytosol with decreased fluorescence, or for JC-1, a shift from red J-aggregates to green monomers evidenced by a decreased red/green fluorescence ratio [40] [44].
Research Reagent Solutions
Successful implementation of TMRM and JC-1 assays requires appropriate selection of reagents and understanding of available commercial resources.
Table 2: Key Research Reagents for TMRM and JC-1 Assays
| Reagent / Kit | Primary Function | Application Context | Key Features |
|---|---|---|---|
| JC-1 Bulk Chemical (Thermo Fisher, T3168) [40] | ΔΨm detection by imaging/flow cytometry | Bench-scale experiments; Apoptosis studies | Flexible format for protocol development |
| MitoProbe JC-1 Assay Kit (Thermo Fisher, M34152) [40] | Optimized JC-1 assay for flow cytometry | High-throughput screening; Drug discovery | Includes CCCP depolarization control; Standardized protocol |
| TMRM Assay Kit (Abcam, ab228569) [44] | Complete TMRM staining solution | Flow cytometry, plate readers, microscopy | Includes TMRM reagent, buffer, and FCCP control |
| JC-1 MitoMP Detection Kit (Dojindo, MT09) [46] | ΔΨm detection with enhanced solubility | Multiple platforms: microscopy, flow, plate readers | Includes optimized imaging buffer; Improved dye dissolution |
| FCCP/CCCP (Various suppliers) [40] [44] [46] | Mitochondrial depolarization control | Validation of ΔΨm-dependent staining | Protonophore uncoupler; Essential for assay controls |
| Oligomycin (Various suppliers) [39] [42] | ATP synthase inhibition | Metabolic perturbation studies | Induces mitochondrial hyperpolarization |
Experimental Protocols: From Bench to HTS
Standard Staining Protocol for Adherent Cells
The following core protocol applies to both dyes with specific considerations for each:
Cell Preparation:
- Culture cells on appropriate sterile-treated surfaces (glass coverslips, multi-well plates)
- Ensure cells are 60-80% confluent at time of assay
- Replace culture medium with fresh pre-warmed assay buffer (e.g., HBSS with 20 mM HEPES) [39] [46]
Dye Loading:
- JC-1: Prepare 2-5 μM working concentration in assay buffer. Incubate cells for 20-30 minutes at 37°C protected from light [40] [46].
- TMRM: Prepare 20-200 nM working concentration in assay buffer. Incubate cells for 30-60 minutes at 37°C protected from light [42] [44].
Washing and Imaging:
- Gently rinse cells with warm assay buffer to remove excess dye
- Maintain dye in bath solution for TMRM experiments to prevent signal loss due to dye redistribution [42]
- For JC-1, imaging can proceed in dye-free buffer after equilibration [39] [46]
Critical Control Experiments:
- Depolarization control: Treat cells with 10-50 μM FCCP/CCCP during dye loading or after baseline measurement to confirm ΔΨm-dependent staining [40] [44] [46].
- Viability control: Include cell impermeant dyes (e.g., propidium iodide) to exclude dead cells from analysis.
- Solvent controls: Account for solvent effects (e.g., DMSO) when testing compounds.
High-Throughput Screening Implementation
Adapting ΔΨm assays to HTS platforms requires optimization for reproducibility, minimal perturbation, and compatibility with automation.
Plate Reader Configuration (TMRM):
- Seed neural cells (or other relevant type) at 1.0×10⁶ cells/ml in 48-well plates [47]
- Load with 50 nM TMRM in HBSS or physiological buffer
- Measure fluorescence every 2 minutes (Ex~548 nm/Em~573 nm) with temperature control at 37°C [47]
- Establish baseline for 3 cycles before compound addition
- Validate with FCCP (10 μM) as positive control for depolarization [47]
Flow Cytometry Configuration (JC-1):
- Prepare single-cell suspensions in appropriate buffer
- Load with 2 μM JC-1 for 15-30 minutes at 37°C, 5% CO₂ [40]
- Wash cells with phosphate-buffered saline before analysis
- Analyze using 488 nm excitation with 530 nm (green) and 585 nm (red) bandpass emission filters [40]
- Calculate population statistics based on red/green fluorescence ratio [40] [43]
HTS Experimental Workflow: The transition from bench-scale experiments to high-throughput screening follows a structured pathway to ensure data quality and reproducibility:
Diagram 2: High-Throughput Screening Workflow for TMRM and JC-1 Assays
Data Analysis and Interpretation
TMRM Data Analysis:
- For time-lapse imaging, normalize fluorescence intensity to baseline (F/F₀)
- Region of interest (ROI) analysis for individual mitochondria or cellular areas
- Calculate rate and extent of depolarization/hyperpolarization in response to stimuli
- Account for photobleaching through control experiments
JC-1 Data Analysis:
- Calculate ratio of red (J-aggregate) to green (monomer) fluorescence intensity
- For microscopy, create ratio images to visualize spatial heterogeneity
- For flow cytometry, plot green vs. red fluorescence to distinguish populations
- Establish threshold values for healthy vs. depolarized mitochondria
Troubleshooting Common Issues:
- Excessive photobleaching: Reduce illumination intensity, use neutral density filters, or optimize exposure time [39]
- Non-specific staining: Optimize dye concentration and include dead cell exclusion markers
- Dye precipitation (JC-1): Filter dye solution before use and ensure proper solvent composition [46]
- Incomplete depolarization with FCCP: Verify compound activity, concentration, and incubation time
Applications in Metabolic Research and Drug Discovery
The applications of TMRM and JC-1 assays span fundamental mitochondrial biology to drug discovery, particularly in the context of metabolic partitioning research.
Mapping Metabolic Specialization
TMRM has been instrumental in detecting spontaneous, transient depolarizations in individual mitochondria within neuronal processes, reflecting inherent oscillations between active and inactive states of oxidative phosphorylation [39]. These ΔΨm fluctuations appear to represent a physiological mechanism for optimizing the balance between respiration, ATP synthesis, and free radical production [39]. Such dynamic measurements are essential for understanding how mitochondria within specialized cellular compartments (e.g., neuronal processes, muscle sarcomeres) tailor their metabolic output to local requirements.
JC-1's rationetric properties make it particularly valuable for detecting heterogeneous mitochondrial populations within cells, such as in cancer research where subpopulations with different metabolic profiles (glycolytic vs. oxidative) coexist [40] [43]. The ability to distinguish these subcompartments based on ΔΨm provides critical insights into metabolic plasticity and specialization.
High-Content Screening for Mitochondrial Modulators
Recent advances have combined these dyes with high-content imaging and machine learning to identify small molecules that modulate mitochondrial function. A high-throughput screen measuring ΔΨm and ATP production in neurons identified structural classes including local anesthetics, isoflavones, COX-II inhibitors, and neurotransmitter system effectors that enhance mitochondrial function and protect against toxic insults relevant to neurodegenerative diseases [41]. This approach has been successfully applied to complex models including spheroids, isolated muscle fibers, and co-culture systems where automated image analysis can discriminate different cell types based on mitochondrial parameters [42].
The integration of ΔΨm measurements with other parameters (morphology, ROS production, calcium flux) creates multiparametric metabolic signatures that can classify compounds by mechanism of action or identify pathological patterns in patient-derived cells [41] [42]. These approaches are particularly powerful for understanding how cells partition metabolic resources under different conditions and how disease states disrupt this balance.
TMRM and JC-1 assays provide complementary approaches for assessing mitochondrial membrane potential from single organelles to population-level analyses. TMRM excels in kinetic measurements and high-temporal resolution studies with minimal perturbation, while JC-1 offers built-in rationetric quantification that controls for technical variables. The successful implementation of these assays requires careful attention to dye concentrations, appropriate controls, and platform-specific optimization. As research increasingly focuses on metabolic specialization and partitioning in health and disease, these potentiometric dyes will continue to be essential tools for deciphering mitochondrial function across biological scales, from detailed mechanistic studies to high-throughput drug discovery campaigns aimed at modulating mitochondrial function in pathological conditions.
Mitochondrial membrane potential (MMP) is a critical parameter of cellular health and function, representing the electrical gradient across the inner mitochondrial membrane. Recent research has revealed that MMP is not merely a uniform "battery" for ATP production but rather a dynamic, compartmentalized signaling hub that regulates fundamental cellular processes. The MMP facilitates metabolic specialization within mitochondrial subpopulations, directing resources toward either oxidative ATP production or reductive biosynthesis based on cellular demands [2]. This partitioning is particularly crucial in complex cellular environments like neurons, where MMP changes support synaptic plasticity and dendritic spine remodeling by linking metabolic state to structural changes at synapses [2].
Advances in super-resolution microscopy have revealed that the inner mitochondrial membrane maintains distinct electrical potentials across its sub-compartments—the cristae membrane (ΔΨC) and inner boundary membrane (ΔΨIBM)—separated by the crista junction barrier [6]. This compartmentalization creates ultrastructures with different phospholipid and protein compositions, enabling specialized functional domains within individual mitochondria [6]. The ability to screen for compounds that modulate these specialized domains requires sophisticated high-throughput screening (HTS) approaches that can capture the nuanced dynamics of MMP distribution and function.
Adapting MMP assays to 384-well platforms represents a critical advancement for drug discovery and basic research, enabling rapid assessment of mitochondrial function across thousands of experimental conditions. This technical guide provides comprehensive methodologies for implementing robust, quantitative MMP screening in 384-well formats, with particular emphasis on applications in metabolic partitioning research.
MMP Fundamentals and Technical Considerations
Biochemical Principles of Membrane Potential
The mitochondrial membrane potential is generated through electron transport chain activity, where complexes I, III, and IV pump protons from the mitochondrial matrix to the intermembrane space. This creates an electrochemical potential gradient known as protonmotive force (PMF), consisting of both an electrical component (MMP, approximately -180 mV) and a chemical component (ΔpH, approximately 0.4 pH units) [2]. Under physiological conditions, MMP serves as the primary contributor to the PMF, generating a driving force equivalent to a 1000-fold difference in proton concentration across the membrane [2].
The MMP dynamically regulates mitochondrial quality control mechanisms. Reduced MMP triggers the accumulation of PINK1, which recruits Parkin and LC3, marking dysfunctional mitochondria for degradation via mitophagy [2]. Additionally, MMP dictates protein import into mitochondria, as proteins with positively charged targeting signals are pulled across the inner membrane by the electrical driving force provided by MMP [2].
Spatial Organization of MMP
Recent super-resolution microscopy studies have revealed that MMP is not uniform across mitochondrial compartments. The cristae membranes (CM), which harbor the proton pumps (complexes I, III, and IV), typically maintain a higher (more negative) membrane potential (ΔΨC) compared to the inner boundary membranes (ΔΨIBM) [6]. The crista junction acts as a barrier that separates these compartments and regulates ion movement [6].
This compartmentalization has significant functional implications. Mitochondrial calcium elevation primarily hyperpolarizes the CM, likely through Ca²⁺-sensitive enhancement of TCA cycle activity and subsequent oxidative phosphorylation in the cristae [6]. This spatial specialization enables compartment-specific signaling and regulation, with the CJ potentially acting as a "membrane potential overflow valve" to protect mitochondrial integrity during excessive cristae hyperpolarization [6].
Adapting MMP Assays to 384-Well Platforms
Fluorescent Probe Selection and Optimization
Successful implementation of MMP assays in 384-well formats requires careful selection and optimization of potentiometric fluorescent dyes. The choice of dye concentration is particularly critical, as it significantly affects the ability to resolve spatial membrane potential gradients.
Table 1: TMRM Concentration Optimization for Spatial MMP Detection
| TMRM Concentration (nM) | Spatial Resolution | Primary Localization | Application |
|---|---|---|---|
| 1.35 - 2.7 | High | Cristae membrane | Detecting ΔΨC gradients |
| 5.4 - 13.5 | Moderate | Both compartments | General MMP screening |
| 40.5 - 81 | Low | Inner boundary membrane | Bulk MMP measurements |
As shown in Table 1, lower TMRM concentrations (1.35-2.7 nM) enable superior resolution of cristae-specific membrane potential, while higher concentrations saturate the cristae and provide primarily IBM signals [6]. This concentration-dependent distribution enables researchers to tailor their approach based on screening objectives—whether targeting general MMP changes or compartment-specific potential gradients.
Assay Validation and Quality Control
Robust assay validation is essential for high-throughput screening. Quality control metrics should be established during assay development to ensure reproducible and reliable results.
Table 2: HTS Quality Control Metrics for MMP Assays
| Parameter | Target Value | Calculation | Interpretation | ||
|---|---|---|---|---|---|
| Z'-factor | >0.5 | Z' = 1 - (3×σₚ + 3×σₙ)/ | μₚ - μₙ | Excellent assay robustness | |
| Signal/Background | >10:1 | S/B = μₚ/μₙ | Sufficient dynamic range | ||
| Assay Variability Ratio | <0.6 | AVR = σₚ/μₚ | Acceptable well-to-well variation |
These statistical parameters ensure assay robustness in 384-well formats. The Z'-factor is particularly important, with values above 0.5 indicating excellent assay quality suitable for high-throughput screening [48] [49]. Previous MMP-related screening assays have demonstrated Z'-factors of 0.93-0.95 with signal-to-background ratios exceeding 25:1 for optimized assays [49].
Experimental Protocols for 384-Well MMP Assays
Basic MMP Protocol Using TMRM
Materials:
- Tetramethylrhodamine methyl ester (TMRM)
- MitoTracker Green FM (MTG)
- Black, opaque-walled 384-well plates
- Assay buffer (50 mM Tris pH 7.5, 150 mM NaCl, 10 mM CaCl₂, 10 μM ZnCl₂, 0.05% Brij-35)
- Fluorescence plate reader capable of dual excitation/emission detection
Procedure:
- Cell Preparation: Seed cells at optimized density (typically 5-10×10³ cells/well) in 384-well plates and culture for appropriate duration.
- Staining: Load cells with 500 nM MTG and optimized TMRM concentration (see Table 1) in assay buffer.
- Incubation: Incubate plates at 37°C for 30-45 minutes to allow complete dye loading and distribution.
- Washing: Gently replace dye-containing medium with fresh assay buffer to remove excess dye.
- Baseline Reading: Acquire baseline fluorescence measurements using appropriate filter sets:
- MTG: excitation/emission ~490/516 nm
- TMRM: excitation/emission ~548/573 nm
- Treatment: Add experimental compounds using precision liquid handling systems.
- Endpoint/Kinetic Measurement: Acquire fluorescence readings at experimental endpoint or in kinetic mode to track temporal dynamics.
Data Analysis: Calculate normalized MMP values as TMRM/MTG ratios to account for variations in mitochondrial mass and dye loading efficiency. For spatial distribution analysis, employ the IBM association index or ΔFWHM methods as described in Section 5.2.
Calcium Stimulation Protocol for Metabolic Partitioning Studies
Purpose: To assess MMP dynamics in response to physiological stimulation that promotes metabolic specialization.
Additional Reagents:
- Histamine (or other Ca²⁺-mobilizing agonist)
- Rotenone (Complex I inhibitor)
- Antimycin A (Complex III inhibitor)
Procedure:
- Cell Preparation: Follow steps 1-5 of the Basic MMP Protocol.
- Inhibitor Pre-treatment: For control wells, pre-treat with rotenone (100 nM) or antimycin A (10 nM) for 30 minutes to inhibit electron transport chain activity.
- Stimulation: Add histamine (100 μM final concentration) to appropriate wells using multichannel pipettors or automated dispensers.
- Kinetic Measurement: Monitor TMRM and MTG fluorescence every 30-60 seconds for 15-20 minutes post-stimulation.
- Validation: Include control wells with inhibitors to confirm that observed changes depend on proton pump activity.
Interpretation: Histamine-induced ER Ca²⁺ release promotes mitochondrial Ca²⁺ uptake, which enhances TCA cycle activity and electron transport chain function. This typically produces cristae-specific hyperpolarization detectable as a decreased IBM association index or ΔFWHM [6]. Inhibition of this response by rotenone or antimycin A confirms the dependence on proton pump activity.
Advanced Applications and Data Analysis
Multi-Parameter Correlation Measurements
Advanced screening applications can simultaneously monitor MMP, ATP production, and mitochondrial morphology using correlative multi-parameter approaches:
Diagram 1: MMP Signaling in Metabolic Specialization
This integrated approach reveals how MMP gradients directly correlate with ATP production and mitochondrial dynamics. For instance, histamine-induced cristae hyperpolarization precedes CJ opening and mitochondrial fission, linking metabolic changes to structural remodeling [6].
Spatial MMP Distribution Analysis
Two complementary methods enable quantitative assessment of spatial MMP distribution in high-content screening:
IBM Association Index Method:
- Use MTG channel to define mitochondrial boundaries via automated Otsu thresholding.
- Generate IBM and CM regions through sequential shrinking and widening of borders.
- Calculate IBM Association Index = TMRMIBM/TMRMCM.
- Decreased index values indicate relative cristae hyperpolarization [6].
ΔFWHM Method:
- Extract cross-sectional intensity profiles for both MTG and TMRM.
- Calculate full width at half maximum (FWHM) for each profile.
- Determine ΔFWHM = FWHMMTG - FWHMTMRM.
- Increased ΔFWHM indicates greater TMRM accumulation in cristae [6].
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for MMP Screening in 384-Well Formats
| Reagent/Category | Specific Examples | Function/Application | Optimization Tips |
|---|---|---|---|
| Potentiometric Dyes | TMRM, TMRE | MMP detection | Concentration critical for spatial resolution |
| Mitochondrial Mass Markers | MitoTracker Green FM | Normalization reference | Lacks MMP sensitivity post-accumulation |
| FRET-Based ATP Biosensors | ATeam, QUEEN | Correlative ATP measurement | Requires genetically encoded sensors |
| Complex Inhibitors | Rotenone (CI), Antimycin A (CIII) | Mechanism validation | Confirm ETC dependence of responses |
| Calcium Agonists | Histamine, ATP | Physiological stimulation | Model physiological metabolic switching |
Adapting MMP assays to 384-well platforms represents a significant advancement in mitochondrial research and drug discovery. The ability to screen compound libraries for effects on MMP—particularly spatial MMP gradients—enables identification of novel modulators of metabolic partitioning and specialization. These approaches are especially valuable for investigating pathological conditions where metabolic reprogramming occurs, such as cancer, neurodegenerative diseases, and metabolic disorders [2].
Future developments in this field will likely include increased integration of multi-parameter correlative microscopy with high-throughput screening, combining the spatial resolution of super-resolution techniques with the statistical power of HTS [6]. Additionally, advances in artificial intelligence-driven analysis of pharmacotranscriptomic data may provide new insights into how MMP-modulating compounds influence global gene expression patterns and signaling pathways [50].
The methods outlined in this technical guide provide a robust foundation for implementing MMP screening in 384-well formats, with particular relevance to research on metabolic specialization and partitioning. By capturing the dynamic, compartmentalized nature of MMP, these approaches offer unprecedented insight into mitochondrial function in health and disease.
Mitochondrial function serves as a central biomarker in physiology and disease pathogenesis, from diabetes and long COVID to neurodegenerative conditions and reproductive aging [51] [52]. Traditionally, mitochondrial assessment has relied on isolated measurements, but this singular approach fails to capture the complexity of mitochondrial biology. The integration of respirometry, which measures oxygen consumption rates as an indicator of electron transport chain activity, with mitochondrial membrane potential (MMP) assessment, which quantifies the electrochemical gradient essential for ATP production, provides a multidimensional analytical framework [51] [2]. This combined approach is particularly valuable for investigating metabolic specialization and partitioning—the phenomenon where mitochondria within the same cell adopt specialized roles through dynamic regulation of their functional properties [2] [53].
The protonmotive force (PMF), generated by electron transport chain complexes I, III, and IV, consists of both an electrical gradient (MMP) and a chemical gradient (ΔpH) [2]. Under physiological conditions, MMP contributes approximately 75% of the total PMF, serving as the primary driving force for ATP synthesis while also functioning as a dynamic signaling hub that influences reactive oxygen species production, calcium handling, and mitochondrial quality control [2]. Recent research reveals that MMP is not uniform across mitochondrial networks but exhibits regional variations that facilitate metabolic compartmentalization, enabling distinct mitochondrial subpopulations to specialize in either oxidative ATP production or reductive biosynthetic processes [2]. This nuanced understanding of mitochondrial heterogeneity underscores the necessity of combining respirometry with MMP assessment to fully characterize mitochondrial functional states in health and disease.
Technical Foundations: Principles and Methodologies
Respirometry Methodologies and Platforms
Respirometry measures oxygen consumption rates, providing an integrative readout of cellular metabolism and mitochondrial function. Because respiration is coupled to ATP synthesis, respirometry can study any process that makes or consumes ATP so long as experimental conditions allow that process to control the overall oxygen consumption rate [54]. Two primary platforms dominate current respirometry research:
Table 1: Comparison of Respirometry Measurement Platforms
| Parameter | Chamber-Based Platinum Electrode | Plate-Based Fluorescence/Phosphorescence |
|---|---|---|
| Common Vendors | Oroboros Instruments, Hansatech Instruments, Rank Brothers | Agilent Seahorse XF Analyzer, Cayman Oxygen Consumption Rate Assay |
| Throughput | Single or dual chambers measuring 1-2 technical replicates sequentially | 96-well microplate format allowing simultaneous assessment of multiple experimental groups |
| Sample Requirement | Larger chamber volumes require increased material | Dramatically reduced sample needs, suitable for clinical biopsies |
| Additional Measurements | Can be multiplexed with electrodes for ROS, pH, Ca2+, MMP | Simultaneously measures extracellular acidification rates (glycolysis) |
| Data Output | Direct access to raw data for manual calculation | Proprietary software automatically calculates rates |
Chamber-based systems offer superior sensitivity for very low respiratory rates and enable unlimited injections of effector compounds, while plate-based systems provide higher throughput and require minimal sample material [54]. The experimental model system—isolated mitochondria, permeabilized cells, or intact cells—should be selected based on the research question. Isolated mitochondria are ideal for identifying intrinsic mitochondrial defects, while permeabilized cells preserve intracellular interactions and require less starting material [54].
MMP Assessment Techniques
MMP can be quantified using potentiometric dyes with flow cytometry or imaging approaches. Flow cytometry enables high-throughput multiparametric assessment of mitochondrial function in live cells, typically using combinations of MitoTracker Green (mitochondrial mass), MitoTracker Red (MMP), and MitoSOX (mitochondrial superoxide) [55]. A standard protocol involves staining cells with 100 nM MitoTracker Green and 50 nM MitoTracker Red in PBS with 2% FCS for 15 minutes at 37°C, followed by immediate acquisition on a flow cytometer [55]. Recent advances in imaging flow cytometry combine the statistical power of flow cytometry with visual confirmation, allowing simultaneous assessment of mitochondrial fragmentation, swelling, membrane potential, ROS production, and mass [56].
MMP typically ranges around -180 mV, generating a driving force equivalent to a 1000-fold difference in proton concentration across the inner mitochondrial membrane [2]. This charge separation not only powers ATP synthesis but also regulates protein import into mitochondria, as proteins with positively charged targeting sequences are pulled across the inner membrane by the electrical driving force provided by MMP [2]. The dependence of protein import on MMP creates a potential mechanism for metabolic specialization, as regional variations in MMP could influence mitochondrial composition and function [2].
Integrated Experimental Design: Combining Respirometry and MMP
Protocol for Concurrent Respirometry and MMP Assessment in Cultured Cells
The following integrated protocol enables researchers to obtain complementary respirometry and MMP data from the same cell population, providing a comprehensive functional assessment:
Cell Culture and Preparation:
- Culture primary buccal cells or fibroblasts in T25 flasks with DMEM + 10% FBS at 37°C with 5% CO₂ until ~80% confluency [57].
- For respirometry measurements, harvest cells using trypsin and resuspend in appropriate respiration medium (MiR05 or Z). Cell count should be determined using an automated cell counter [57].
- Split the cell suspension for parallel respirometry and MMP assays to enable direct comparison.
Respirometry Measurements:
- Perform high-resolution respirometry using an Oxygraph-2K system or Seahorse XF Analyzer [54] [57].
- Calibrate the instrument according to manufacturer specifications using sodium dithionite for zero oxygen calibration [57].
- Utilize substrate-uncoupler-inhibitor titration (SUIT) protocols to assess multiple mitochondrial functions sequentially:
- LEAK State: Basal respiration with complex I substrates (pyruvate + malate)
- OXPHOS Capacity: State 3 respiration after ADP addition
- ET Capacity: Maximum electron transport capacity after FCCP uncoupling
- Complex II Activity: Succinate addition with rotenone
- Residual Oxygen Consumption: Non-mitochondrial respiration after antimycin A inhibition [54]
MMP Assessment:
- For simultaneous MMP measurement during respirometry, aliquot cells and stain with 50 nM MitoTracker Red CMXRos or TMRE during the respiratory measurements [55] [52].
- Alternatively, perform parallel MMP assessment using flow cytometry with the same cell batch under identical conditions.
- Include controls for compensation and autofluorescence using fluorescence-minus-one (FMO) controls [55].
Data Normalization and Analysis:
- Normalize oxygen consumption rates to cell count, protein content, or citrate synthase activity [54].
- Express MMP as fluorescence intensity ratios relative to control conditions.
- Calculate coupling efficiency (1-LEAK/OXPHOS) and ATP production efficiency from the combined dataset.
Research Reagent Solutions
Table 2: Essential Reagents for Combined Respirometry and MMP Assays
| Reagent Category | Specific Examples | Function in Assays |
|---|---|---|
| Respiration Media | MiR05, Medium Z | Provide ionic and osmotic stability during respirometry; composition affects measured respiratory capacities [58] |
| MMP Indicators | MitoTracker Red CMXRos, TMRE, JC-1 | Potentiometric dyes that accumulate in mitochondria in proportion to MMP; allow quantification via fluorescence [55] [52] |
| Metabolic Substrates | Pyruvate, Malate, Succinate, Glutamate | Provide specific entry points to mitochondrial metabolic pathways; test function of different dehydrogenase complexes [54] |
| Inhibitors/Uncouplers | Rotenone, Antimycin A, Oligomycin, FCCP | Inhibit specific ETC complexes or uncouple respiration from ATP synthesis; enable dissection of ETC function [54] [57] |
Data Interpretation and Analysis in Metabolic Specialization Research
Integrated Parameters for Functional Classification
The combination of respirometry and MMP measurements enables identification of distinct mitochondrial functional phenotypes. Research on mitochondrial disease patients reveals that fibroblasts cluster into two overarching groups: a hypometabolic state with low-to-normal MMP and reduced respiration, and a hypermetabolic state with elevated MMP and respiration [56]. These phenotypes correlate with different molecular mechanisms—complex I stability defects in hypometabolic states versus altered proton pumping activity in hypermetabolic states [56].
In metabolic specialization, MMP serves as a regulatory switch between oxidative and reductive metabolism. Elevated MMP relative to baseline enhances the activity of enzymes like pyrroline-5-carboxylate synthase (P5CS), promoting the formation of filamentous assemblies that drive reductive biosynthesis [2]. Reduced MMP inhibits this filamentation and limits substrate production, shifting mitochondrial function toward core energetic functions like OXPHOS [2]. The combined assessment of respiration and MMP can therefore identify whether mitochondria are specializing in energy production or biosynthetic precursor generation.
Conceptual Framework for Metabolic Specialization
This diagram illustrates how MMP changes direct metabolic specialization toward either oxidative phosphorylation or reductive biosynthesis, driving different functional outcomes in cells.
Experimental Workflow for Combined Assessment
This workflow diagram outlines the parallel assessment of respirometry and MMP parameters, followed by data integration for comprehensive mitochondrial phenotyping.
Applications in Disease Research and Therapeutic Development
The combined respirometry-MMP approach provides critical insights into disease mechanisms and therapeutic interventions. In mitochondrial diseases, this integrated assessment identified two distinct functional phenotypes—hypometabolic and hypermetabolic—with the hypermetabolic cluster showing significantly more neuropathy, suggesting clinical relevance of this classification [56]. In reproductive medicine, MMP assessment in gametes combined with respirometry of cumulus cells offers insights into declining fertility associated with aging, where reduced MMP in aged oocytes indicates higher apoptotic susceptibility and decreased energy production [52].
The pharmaceutical industry can leverage this approach for drug safety screening, as many drug candidates exhibit mitochondrial toxicity that manifests as specific patterns of respiratory dysfunction and MMP dissipation. The combination of respirometry and MMP assessment can distinguish between compounds that directly inhibit electron transport chain complexes versus those that uncouple phosphorylation or induce mitochondrial permeability transition [54]. This distinction is crucial for lead optimization, as different MMP-respirometry profiles suggest different structural modifications to mitigate toxicity.
For neurodegenerative diseases like Alzheimer's and Parkinson's disease, where mitochondrial dysfunction is an early event, the combined assessment can identify specific defects in metabolic specialization. Research indicates that variants in uncoupling proteins (UCPs), which regulate MMP dissipation, are associated with neurodegenerative conditions, suggesting that altered MMP dynamics contribute to disease pathogenesis [2]. The integrated respirometry-MMP approach provides a framework for evaluating how these genetic variants affect mitochondrial bioenergetics and metabolic partitioning in neuronal subtypes with distinct energetic demands.
The synergistic application of respirometry and MMP assessment creates a powerful methodological framework for investigating mitochondrial function in health and disease. This combined approach moves beyond singular parameter measurement to capture the dynamic interplay between energy production, membrane potential regulation, and metabolic specialization. As research continues to reveal the complexity of mitochondrial networks and their role in cellular homeostasis, the integration of multiple analytical techniques will be essential for advancing our understanding of mitochondrial biology and developing targeted therapeutic interventions. The standardized protocols and analytical frameworks presented here provide researchers with a roadmap for implementing this comprehensive assessment strategy across diverse experimental systems and disease models.
The mitochondrial membrane potential (MMP), a fundamental bioenergetic parameter maintained by the electron transport chain, represents a powerful physiological force for targeted drug delivery. This whitepaper examines the strategic exploitation of delocalized lipophilic cations (DLCs) to harness the MMP for mitochondrial-specific targeting of therapeutic agents. With a typical potential of -180 mV relative to the cytoplasm, the MMP generates an electrochemical gradient that drives the accumulation of DLCs within the mitochondrial matrix, achieving concentrations 100-500-fold higher than in the extracellular space. This review comprehensively analyzes the underlying principles, key molecular frameworks, quantitative accumulation metrics, and advanced experimental methodologies for evaluating DLC-based mitochondrial targeting. Furthermore, we situate these targeting strategies within the emerging paradigm of mitochondrial metabolic specialization and compartmentalization, highlighting how MMP-driven drug delivery can be leveraged to address subcellular metabolic partitioning in disease states, particularly in oncology and neurodegenerative disorders.
Fundamental Generation and Maintenance of MMP
The mitochondrial membrane potential (ΔΨm) is an essential electrochemical gradient across the inner mitochondrial membrane (IMM) generated through the proton-pumping activity of electron transport chain (ETC) complexes I, III, and IV [2] [59]. During oxidative phosphorylation, these complexes transfer electrons through sequential redox reactions while translocating protons from the mitochondrial matrix to the intermembrane space, establishing both an electrical potential (ΔΨm) and a chemical pH gradient (ΔpH) that collectively constitute the proton motive force (PMF) [2]. Under physiological conditions, the MMP typically ranges between -180 mV to -200 mV, contributing approximately 75-80% of the total PMF, while the ΔpH accounts for the remaining 20-25% [2] [59]. This electrical gradient serves as the primary driving force for ATP synthesis through F1F0-ATP synthase and powers critical mitochondrial processes including protein import, metabolite exchange, and calcium homeostasis [2] [59].
MMP in Metabolic Specialization and Cellular Signaling
Beyond its canonical role in energy transduction, the MMP functions as a dynamic signaling hub that integrates cellular status with mitochondrial output [2]. Recent research has revealed that the MMP is not uniform across the mitochondrial network but exhibits spatial heterogeneity that facilitates metabolic specialization and compartmentalization [2]. Mitochondria can segregate into subpopulations with distinct metabolic priorities—some dedicated to ATP production via oxidative phosphorylation while others prioritize biosynthetic precursor synthesis through reductive metabolism [2]. This metabolic partitioning is regulated in part by MMP gradients, which influence the activity of key metabolic enzymes such as pyrroline-5-carboxylate synthase (P5CS) [2]. Elevated MMP enhances P5CS filamentation, promoting proline biosynthesis and reductive metabolic pathways, while reduced MMP inhibits this activity, shifting mitochondrial function toward oxidative energy production [2]. This MMP-mediated metabolic specialization has profound implications for cellular physiology and represents a critical framework for understanding mitochondrial contributions to complex diseases, including cancer.
Delocalized Lipophilic Cations: Fundamental Principles and Mechanisms
Theoretical Foundation for Mitochondrial Accumulation
Delocalized lipophilic cations (DLCs) represent a class of mitochondrial-targeting molecules characterized by a positively charged center surrounded by hydrophobic molecular structures. The fundamental mechanism driving their mitochondrial accumulation relies on both the highly negative MMP and the chemical properties of the DLCs themselves. According to the Nernst equation, the relationship between membrane potential and cation concentration gradient can be quantified as:
Membrane Potential (mV) = 61.5 log([cation]in/[cation]out) at 37°C [60]
This equation predicts that for every -61.5 mV of membrane potential, a tenfold increase in cation accumulation occurs within the mitochondrial matrix. Given the typical MMP of -180 mV, DLCs can achieve 100-500-fold higher concentrations in the mitochondrial matrix compared to the cytoplasm or extracellular space [60] [61]. The "delocalized" nature of the positive charge refers to its distribution across an extended π-orbital system, which reduces the energy penalty for membrane penetration and facilitates translocation across both the plasma membrane and mitochondrial membranes [61] [62].
Key Structural Classes of Delocalized Lipophilic Cations
Table 1: Major Classes of Delocalized Lipophilic Cations for Mitochondrial Targeting
| Class | Representative Molecules | Key Structural Features | Accumulation Mechanism | Therapeutic Applications |
|---|---|---|---|---|
| Triphenylphosphonium (TPP) | TPP-conjugated compounds, MitoQ | Positively charged phosphorus atom surrounded by three hydrophobic phenyl groups | Electrophoretic accumulation driven by MMP; lipophilic groups enable membrane penetration | Antioxidant delivery (MitoQ), chemotherapeutic targeting, photodynamic therapy [61] [62] |
| Rhodamine Derivatives | Rhodamine 123, TMRM, TMRE | Fluorophore with delocalized positive charge, lipophilic structure | Potential-dependent accumulation; fluorescence properties enable MMP monitoring | MMP measurement, imaging, phototherapy [61] [63] |
| Cyanine Dyes | JC-1, DiOC6(3) | Carbocyanine-based fluorophores with delocalized charge | Concentration-dependent fluorescence shift (JC-1); potential-sensitive accumulation | MMP assessment, apoptosis detection [59] [63] |
| Guanidinium-Based Molecular Transporters | Sorbitol-based guanidinium transporters | Multiple guanidine groups attached to carbohydrate scaffolds | Electrostatic attraction to negative MMP; enhanced by lipophilic modifications | Drug conjugates for mitochondrial dysfunction [60] |
The triphenylphosphonium (TPP) cation represents the most extensively characterized and utilized DLC for mitochondrial targeting [61] [62]. Its exceptional properties include remarkable chemical stability in biological systems, straightforward synthesis and purification, minimal chemical reactivity with cellular components, and negligible light absorption or fluorescence in the near-infrared spectrum, making it an ideal carrier for therapeutic applications [61]. The targeting process occurs through a multi-step mechanism: TPP first binds to the outer mitochondrial membrane, then traverses the inner mitochondrial membrane, and finally dissociates within the mitochondrial matrix, where it accumulates due to the highly negative potential [61].
Quantitative Assessment of DLC Accumulation and MMP Dependence
Mathematical Modeling of Mitochondrial Uptake
The accumulation of DLCs within mitochondria follows predictable physicochemical principles that can be quantitatively modeled. The Nernst equation provides the fundamental relationship between membrane potential and distribution ratio for monovalent cations. For a typical MMP of -180 mV, the concentration gradient can be calculated as:
-180 mV = 61.5 log([DLC]in/[DLC]out)
log([DLC]in/[DLC]out) = -180/61.5 ≈ -2.93
[DLC]in/[DLC]out ≈ 10^(-2.93) ≈ 1/850
This calculation indicates an approximately 850-fold accumulation within the mitochondrial matrix relative to the external environment [60]. In practice, measured accumulation ratios typically range from 100-500-fold due to factors including plasma membrane potential, binding to mitochondrial membranes, and active export mechanisms [61] [60]. The addition of multiple cationic charges further enhances this accumulation, with each charge contributing multiplicatively to the total targeting efficiency [60].
Table 2: Quantitative Accumulation Parameters for Selected DLCs
| DLC Compound | Theoretical Accumulation Ratio | Experimentally Measured Ratio | Key Influencing Factors | Primary Applications |
|---|---|---|---|---|
| TPP Derivatives | 100-1000-fold | 100-500-fold | Lipophilicity, molecular weight, number of cationic charges | Drug conjugates, antioxidant delivery [61] [62] |
| Rhodamine 123 | 100-500-fold | 50-200-fold | MMP sensitivity, fluorescence self-quenching | MMP monitoring, imaging [61] [63] |
| JC-1 Monomer | 100-500-fold | N/A | Concentration-dependent aggregation, fluorescence shift | MMP assessment (red/green ratio) [59] [63] |
| Guanidinium-Based Transporters | 100-500-fold per guanidinium | Varies with lipophilicity | Number of guanidine groups, lipophilic modifications | Drug delivery, mitochondrial dysfunction [60] |
Impact of Molecular Properties on Targeting Efficiency
The efficiency of mitochondrial accumulation depends critically on specific molecular properties of the DLCs. Lipophilicity represents a crucial parameter, as it determines the ability to permeate biological membranes. An optimal partition coefficient (log P) typically falls in the range of 1-3, balancing membrane penetration with sufficient aqueous solubility for distribution [60]. Molecular size and shape also significantly influence targeting efficiency, with smaller molecules generally exhibiting faster accumulation kinetics. The number and distribution of positive charges directly affect the magnitude of MMP-driven accumulation, with multivalent cations typically showing enhanced mitochondrial targeting but potentially reduced cellular uptake due to increased hydrophilicity [60]. Recent advances in molecular design have focused on optimizing these parameters through strategic incorporation of lipophilic groups such as alanine-naphthalene, which enhances membrane penetration while maintaining MMP-driven accumulation [60].
Advanced Experimental Methodologies for DLC Evaluation
Protocol 1: Quantitative Assessment of Mitochondrial Accumulation
Objective: Determine the mitochondrial accumulation ratio of a novel DLC-conjugated therapeutic agent.
Materials and Reagents:
- Test compound (DLC-therapeutic conjugate)
- Reference compounds (Rhodamine 123, TMRM)
- Cell culture (appropriate cell line)
- Mitochondrial isolation kit
- HPLC system with UV/fluorescence detection
- LC-MS/MS system for quantification
- Buffer solutions (isolation buffer, assay buffer)
Procedure:
- Cell Culture and Treatment: Culture cells to 80% confluence in appropriate medium. Treat with test compound at therapeutically relevant concentrations (typically 1-10 μM) for predetermined time points (0.5, 1, 2, 4 hours).
- Mitochondrial Isolation: Following treatment, harvest cells by trypsinization and wash with PBS. Isolate mitochondria using a differential centrifugation protocol: homogenize cells in isolation buffer (250 mM sucrose, 10 mM HEPES, 1 mM EGTA, pH 7.4), centrifuge at 800 × g for 10 minutes to remove nuclei and debris, then centrifuge supernatant at 10,000 × g for 15 minutes to pellet mitochondria [59].
- Sample Preparation: Lysate mitochondrial fractions and whole cell samples in extraction solvent. Prepare calibration standards in matched matrix.
- Quantitative Analysis: Analyze samples using HPLC-UV/fluorescence or LC-MS/MS. Calculate mitochondrial concentration based on standard curves and normalize to mitochondrial protein content.
- Data Analysis: Determine accumulation ratio as [DLC]mitochondria/[DLC]whole cell. Compare with reference compounds and theoretical values from Nernst equation.
Validation: Confirm mitochondrial purity by assessing marker enzymes (cytochrome c oxidase for mitochondria, lactate dehydrogenase for cytosol).
Protocol 2: MMP Dependency Assessment Using Uncouplers
Objective: Establish the MMP-dependence of DLC accumulation through pharmacological dissipation of MMP.
Materials and Reagents:
- Test DLC compounds
- MMP-sensitive fluorescent probes (JC-1, TMRM)
- Uncoupling agents (FCCP, CCCP)
- Fluorescence plate reader or flow cytometer
- Cell culture system
Procedure:
- Experimental Groups: Divide cells into four treatment groups: (1) vehicle control, (2) test DLC alone, (3) uncoupler alone, (4) test DLC + uncoupler.
- Pretreatment: Pre-incubate groups 3 and 4 with uncoupler (typically 10 μM FCCP) for 15 minutes to dissipate MMP before DLC addition.
- Compound Exposure: Treat groups 2 and 4 with test DLC at working concentration for predetermined time.
- Parallel MMP Monitoring: For validation, include cells loaded with JC-1 (5 μg/mL) or TMRM (100 nM) to confirm MMP dissipation by uncouplers through fluorescence measurement.
- Quantification: Measure intracellular DLC accumulation via fluorescence, HPLC, or LC-MS/MS.
- Data Interpretation: Calculate percentage reduction in DLC accumulation with uncoupler cotreatment. MMP-dependent accumulation typically shows >70% reduction with uncoupler treatment.
Protocol 3: Functional Assessment of Mitochondrial Targeting in Therapeutic Applications
Objective: Evaluate the functional consequences of DLC-mediated mitochondrial targeting on therapeutic efficacy.
Materials and Reagents:
- DLC-therapeutic conjugates and non-targeted controls
- Apoptosis detection kit (Annexin V/PI)
- ROS detection probes (DCFDA, MitoSOX Red)
- ATP quantification kit
- JC-1 or TMRM for MMP assessment
- Western blot equipment for cytochrome c release
Procedure:
- Therapeutic Efficacy Assessment: Treat cells with escalating concentrations of DLC-therapeutic conjugate versus non-targeted control. Assess viability using MTT or resazurin assays at 24, 48, and 72 hours.
- Mechanistic Studies:
- Apoptosis Induction: Detect phosphatidylserine externalization using Annexin V/PI staining and flow cytometry.
- Cytochrome c Release: Fractionate cells into mitochondrial and cytosolic fractions, then detect cytochrome c redistribution via Western blot.
- ROS Production: Measure mitochondrial superoxide production using MitoSOX Red and general ROS with DCFDA.
- MMP Changes: Monitor MMP dynamics using JC-1 (aggregate/monomer ratio) or TMRM fluorescence.
- ATP Levels: Quantify cellular ATP content using luciferase-based assays.
- Data Analysis: Compare dose-response curves, EC50 values, and mechanistic endpoints between targeted and non-targeted compounds.
Diagram 1: Mechanism of DLC Mitochondrial Accumulation. The process illustrates how delocalized lipophilic cations exploit electrochemical gradients for mitochondrial targeting.
The Scientist's Toolkit: Essential Reagents and Methodologies
Table 3: Essential Research Reagents for DLC and MMP Studies
| Reagent/Category | Specific Examples | Function/Application | Key Considerations |
|---|---|---|---|
| MMP-Sensitive Fluorescent Probes | JC-1, TMRM, TMRE, Rhodamine 123 | Quantitative and qualitative MMP assessment | JC-1 exhibits concentration-dependent fluorescence shift; TMRM provides more quantitative measurement; potential dye toxicity must be controlled [59] [63] |
| Pharmacological MMP Modulators | FCCP, CCCP (uncouplers), Oligomycin (ATP synthase inhibitor) | Experimental manipulation of MMP | Uncouplers dissipate MMP; oligomycin can hyperpolarize mitochondria; concentration optimization required [59] |
| Mitochondrial Isolation Kits | Commercial kits based on differential centrifugation | Obtain purified mitochondrial fractions for biochemical studies | Assess purity with compartment-specific markers; maintain cold conditions during isolation [59] |
| DLC Reference Compounds | TPP-conjugated probes, MitoQ, Rhodamine derivatives | Positive controls for mitochondrial targeting | Establish baseline accumulation parameters; validate experimental systems [61] [62] |
| Oxygen Consumption Assays | Seahorse XF Analyzer, Clark electrode | Integrated assessment of mitochondrial function | Correlate MMP with respiratory parameters; requires careful experimental design [59] |
| Apoptosis Detection Reagents | Annexin V/propidium iodide, caspase activity assays | Evaluate functional consequences of mitochondrial targeting | Distinguish early vs. late apoptosis; confirm mitochondrial pathway involvement [59] |
Emerging Applications and Future Perspectives
Advanced Nanocarrier Systems for Enhanced Mitochondrial Targeting
Recent advancements in nanotechnology have significantly expanded the potential of DLC-based mitochondrial targeting. Conventional DLCs face limitations in delivering large therapeutic cargoes, prompting the development of sophisticated nanocarrier systems functionalized with mitochondrial-targeting ligands [61] [62] [64]. These include liposomes, polymeric nanoparticles, dendrimers, and inorganic nanoparticles conjugated with TPP or other DLCs to improve drug loading capacity, bioavailability, and targeting precision [62] [64]. For instance, TPP-conjugated nanoparticles have demonstrated enhanced antitumor efficacy through mitochondrial-mediated apoptosis induction, with several platforms advancing through preclinical development [62]. These systems leverage the enhanced permeability and retention (EPR) effect for tumor accumulation, followed by DLC-mediated mitochondrial targeting for subcellular localization, representing a dual-targeting approach that addresses both tissue and organelle specificity [62] [64].
Integration with Metabolic Specialization Research
The emerging understanding of mitochondrial metabolic specialization and compartmentalization provides a sophisticated framework for advancing DLC-based targeting strategies [2]. As research reveals that mitochondria exist in functionally distinct subpopulations dedicated to specific metabolic tasks, opportunities arise for developing precision targeting approaches that address specific mitochondrial subtypes [2]. For instance, cancer cells often exhibit metabolic reprogramming with distinct mitochondrial subpopulations supporting biosynthetic versus bioenergetic demands [2] [65]. Strategic design of DLC-conjugated therapeutics could leverage these metabolic differences to target specific mitochondrial subsets, potentially overcoming resistance mechanisms and enhancing therapeutic selectivity. Furthermore, the recognition that MMP varies within mitochondrial networks and influences metabolic enzyme activity through effects on protein import and complex assembly provides additional avenues for therapeutic intervention [2].
Diagram 2: DLC Therapeutic Development Workflow. Comprehensive pathway from initial design to therapeutic application of delocalized lipophilic cations.
Delocalized lipophilic cations represent a powerful and well-established strategy for exploiting the mitochondrial membrane potential to achieve subcellular drug targeting. The fundamental physicochemical principles governing DLC accumulation—primarily described by the Nernst equation—enable predictable and substantial concentration within mitochondria, offering significant advantages for therapeutic applications where mitochondrial dysfunction contributes to disease pathology. As research continues to illuminate the complex roles of MMP in metabolic specialization, spatial organization, and stress adaptation, opportunities emerge for increasingly sophisticated targeting approaches that address specific mitochondrial subpopulations and functional states. The integration of DLC targeting with advanced nanocarrier systems and a growing understanding of mitochondrial biology promises to advance therapeutic strategies for cancer, neurodegenerative diseases, and metabolic disorders. However, challenges remain in optimizing delivery efficiency, minimizing off-target effects, and addressing the potential toxicity of cationic compounds, underscoring the need for continued innovation in this dynamic field.
Mitochondriotropic antioxidants represent a advanced class of compounds specifically engineered to mitigate oxidative damage within mitochondria, the primary source of cellular reactive oxygen species (ROS). Among these, MitoQ (mitoquinone mesylate) has emerged as a preeminent therapeutic candidate, demonstrating significant potential across a spectrum of diseases linked to mitochondrial dysfunction. MitoQ is a synthetic molecule comprising a ubiquinone antioxidant moiety covalently attached to a lipophilic triphenylphosphonium (TPP+) cation via a 10-carbon aliphatic chain [66] [67]. This strategic design enables the molecule to cross phospholipid bilayers and accumulate several hundred-fold within the mitochondrial matrix, driven by the substantial mitochondrial membrane potential (ΔΨm, typically -150 to -180 mV) [68]. This accumulation far exceeds the capabilities of untargeted antioxidants, positioning MitoQ as a potent inhibitor of mitochondrial oxidative damage [69].
The therapeutic relevance of MitoQ is rooted in the critical role of mitochondrial ROS in pathophysiology. Under normal physiological conditions, cells maintain redox balance through enzymatic and non-enzymatic antioxidant systems. However, excessive mitochondrial ROS can attack lipids, proteins, and DNA, leading to severe and irreversible oxidative damage implicated in a wide array of disorders, including neurodegenerative diseases, cardiovascular conditions, cancer, and aging [70]. MitoQ's mechanism of action is multifaceted: its ubiquinol form directly scavenges ROS, preventing oxidative damage to mitochondrial components, including membrane lipids and mitochondrial DNA (mtDNA) [67] [69]. Furthermore, by modulating the mitochondrial redox environment, MitoQ influences crucial cellular processes such as apoptosis, autophagy, and inflammatory signaling, thereby exerting protective effects in various disease models [68] [71].
Molecular Mechanisms and Mitochondrial Membrane Potential
The Central Role of Mitochondrial Membrane Potential
The mitochondrial membrane potential (ΔΨm) is a critical component of the proton motive force used to generate ATP through oxidative phosphorylation. This electrochemical gradient, established by proton pumping across the inner mitochondrial membrane during electron transport through complexes I, III, and IV, creates a negative interior environment that exerts a strong driving force for the uptake of lipophilic cations [68]. MitoQ exploits this fundamental physiological property through its TPP+ moiety, which enables the molecule to permeate lipid bilayers and accumulate substantially within the mitochondrial matrix, where concentrations can reach 1000-fold higher than in the incubation medium [72]. This targeting mechanism ensures that MitoQ is positioned precisely where its antioxidant activity is most needed—at the primary site of cellular ROS generation.
Recent research has revealed that ΔΨm is not merely a passive targeting mechanism but also a critical factor in metabolic specialization and cellular partitioning. Studies in hematopoietic stem and progenitor cells (HSPCs) with DNMT3A mutations, which drive clonal hematopoiesis, have demonstrated that these mutant cells maintain an elevated ΔΨm compared to wild-type cells [73]. This elevated membrane potential is associated with enhanced mitochondrial respiration, increased spare respiratory capacity, and resistance to aging-related microenvironmental changes. The heightened ΔΨm in mutant HSPCs creates a self-reinforcing cycle: it drives greater accumulation of TPP+-conjugated compounds like MitoQ, which in turn can selectively modulate the function of these cells based on their metabolic state [73]. This phenomenon illustrates how mitochondrial membrane potential serves as both a therapeutic target and a means for achieving cell-type specific interventions.
Complex Bioenergetic Interactions and Signaling Pathways
Upon accumulation in mitochondria, MitoQ exhibits complex effects on mitochondrial bioenergetics that extend beyond its antioxidant properties. Research indicates that MitoQ can induce a "pseudo-mitochondrial membrane potential" (PMMP) by introducing exogenous positive charges into the intermembrane space [68]. This PMMP is maintained by the cationic TPP+ moieties of accumulated MitoQ molecules rather than by proton pumping, which leads to inhibition of respiratory chain complexes I, III, and IV, reduced proton production, and decreased oxygen consumption [68]. The resulting impairment of proton backflow through ATP synthase decreases ATP production while increasing AMP levels, subsequently activating AMPK and inhibiting the MTOR pathway to induce autophagy [68].
The following diagram illustrates MitoQ's structure, mitochondrial accumulation, and downstream effects on cellular signaling pathways:
Simultaneously, MitoQ activates the Nrf2/ARE signaling pathway, a critical regulator of cellular antioxidant responses [69]. Under oxidative stress conditions, MitoQ promotes Nrf2 translocation to the nucleus, where it binds to antioxidant response elements (ARE) and upregulates cytoprotective genes including heme oxygenase-1 (HO-1), NAD(P)H quinone dehydrogenase 1 (NQO-1), and the catalytic subunit of γ-glutamylcysteine ligase (γ-GCLC) [69]. This pathway activation enhances cellular resilience to oxidative stress and contributes to the stabilization of mitochondrial transcription factor A (TFAM), thereby protecting mitochondrial DNA from oxidative damage and maintaining mitochondrial function [69].
Therapeutic Applications and Experimental Evidence
MitoQ has demonstrated significant potential across diverse disease models, with compelling evidence supporting its therapeutic utility in conditions characterized by mitochondrial oxidative stress. The table below summarizes key quantitative findings from preclinical and clinical studies:
Table 1: Therapeutic Effects of MitoQ in Experimental Models
| Disease Area | Model System | Dosage/Concentration | Key Effects | Mechanistic Insights |
|---|---|---|---|---|
| Hearing Loss [70] | Idh2⁻/⁻ mice | ex vivo administration | Near-complete neutralization of H₂O₂-induced ototoxicity; restored hair cell survival | Increased NADPH and GSH levels; reduced ROS-mediated apoptosis |
| Clonal Hematopoiesis [73] | Dnmt3aR878H/+ mouse model | in vivo and ex vivo | Reduced mitochondrial respiration; ablated competitive advantage of mutant HSPCs | Selective accumulation in high-ΔΨm cells; induced mitochondrial-driven apoptosis |
| Radiosensitization [72] | Human breast cancer models (MDA-MB-231) | 250-500 nM; 18 mg/kg per os in mice | Inhibited mitochondrial OCR; delayed tumor growth with radiotherapy | Reduced oxygen consumption; increased tumor oxygenation |
| Intestinal Ischemia-Reperfusion [69] | Mouse I/R model; IEC-6 cells | 4 mg/kg IV in mice; 0.1-1.0 μM in vitro | Stabilized intestinal barrier; reduced hyperpermeability and apoptosis | Preserved mtDNA copy number via Nrf2/ARE pathway activation |
| Cardiomyocyte Protection [71] | Human iPSC-derived cardiomyocytes | 1 μM | Attenuated H₂O₂-induced ROS; prevented mitochondrial hyperpolarization and cell death | Preserved mitochondrial structure and regulation |
| Septic Shock [74] | Human clinical trial (n=42) | 20 mg twice daily for 5 days | Improved oxidative stress biomarkers (GPx, CAT, SOD, MDA) | Modulated oxidative stress; reduced vasopressor requirements |
Cardiovascular and Metabolic Applications
In cardiometabolic diseases, MitoQ has demonstrated significant protective effects against oxidative stress-induced damage in cardiomyocytes. Studies using human induced pluripotent stem cell-derived cardiomyocytes (hiPSC-CM) and H9C2 rat cardiomyoblasts have shown that MitoQ (1 μM) effectively blunts hydrogen peroxide-induced overproduction of ROS, mitochondrial hyperpolarization, and cell death [71]. Importantly, these cytoprotective effects are attributed to the ubiquinone moiety rather than the TPP+ targeting component, as dodecyl-TPP (dTPP) control failed to provide similar protection [71]. MitoQ also preserves mitochondrial network structure and reduces fragmentation under oxidative stress conditions, suggesting benefits for mitochondrial quality control mechanisms relevant to heart failure and other cardiac conditions [71].
A recent scoping review of MitoQ's effects on cardiometabolic diseases highlighted its potential to improve metabolic parameters, including enhanced insulin secretion and improved lipid profiles, though the authors noted that translation to clinical application requires further investigation to determine appropriate dosing and target populations [66] [75]. The review emphasized that while MitoQ has shown promise in modulating mitochondrial function and oxidative stress in preclinical models, most human evidence remains preliminary, necessitating larger, well-powered clinical trials to establish efficacy [75].
Oncological Applications
In cancer biology, MitoQ exhibits dual functionality—both protecting against oxidative damage in normal tissues and sensitizing malignant cells to conventional therapies. In breast cancer models, MitoQ at clinically achievable nanomolar concentrations (250-500 nM) completely abrogates the oxygen consumption rate (OCR) in multiple human cancer cell lines, including MDA-MB-231 and MCF7 cells [72]. This inhibition of mitochondrial respiration induces a glycolytic switch characterized by increased glucose uptake and lactate release, which in hypoxic tumors improves oxygenation and consequently enhances radiosensitivity [72]. Notably, this effect appears specific to MitoQ, as other mitochondria-targeted antioxidants like MitoTEMPO and SKQ1 do not produce comparable inhibition of cellular respiration [72].
The selective targeting of cancer cells with elevated mitochondrial membrane potential represents another promising application. In Dnmt3a-mutant clonal hematopoiesis, mutant HSPCs with elevated ΔΨm show preferential accumulation of TPP+-conjugated compounds like MitoQ, leading to selective reduction of their mitochondrial respiration and ablation of their competitive advantage [73]. This approach exploits intrinsic metabolic differences between cell populations, suggesting a strategy for targeting malignant or pre-malignant cells based on their bioenergetic profiles rather than specific molecular targets.
Hepatoprotective and Gastrointestinal Applications
MitoQ has demonstrated efficacy in protecting against liver dysfunction and intestinal injury in various models. In hepatic contexts, MitoQ improves high-fat-induced liver dysfunction through its antioxidant properties, reducing oxidative damage and preventing hepatic fat accumulation without necessarily altering liver fat content or mitochondrial bioenergetics [66]. In the gastrointestinal system, MitoQ (4 mg/kg IV) provides significant protection against intestinal ischemia-reperfusion injury by preserving mtDNA integrity, maintaining mitochondrial transcription factor A (TFAM) levels, and activating the Nrf2/ARE pathway [69]. These effects stabilize the intestinal barrier, reduce hyperpermeability, attenuate inflammatory responses, and decrease epithelial apoptosis, collectively preventing the translocation of bacteria and antigens from the intestinal lumen to the systemic circulation [69].
Experimental Protocols and Methodologies
Standard In Vitro Assessment of MitoQ Effects
To evaluate MitoQ's effects on cellular bioenergetics, researchers typically employ extracellular flux analysis to measure oxygen consumption rates (OCR) and extracellular acidification rates (ECAR). The standard protocol involves seeding cells in specialized microplates, allowing attachment, and then treating with MitoQ at concentrations ranging from 62.5 nM to 1 μM for 24 hours [72]. OCR measurements are taken under basal conditions and following sequential injection of mitochondrial modulators: oligomycin (ATP synthase inhibitor), FCCP (mitochondrial uncoupler), and rotenone/antimycin A (complex I and III inhibitors) [72]. This approach allows quantification of basal respiration, ATP-linked respiration, proton leak, maximal respiratory capacity, and spare respiratory capacity. Parallel measurement of ECAR provides insight into glycolytic flux, revealing MitoQ-induced metabolic shifts toward glycolysis when oxidative phosphorylation is inhibited [72].
For assessment of mitochondrial membrane potential, researchers commonly use fluorescent probes such as tetramethyl rhodamine ethyl ester (TMRE) or JC-1. Cells are treated with MitoQ for specified durations (typically 30 minutes to 1 hour), loaded with the potential-sensitive dye, and analyzed by flow cytometry or fluorescence microscopy [68] [73]. JC-1 exhibits potential-dependent accumulation in mitochondria, indicated by a fluorescence emission shift from green (~529 nm) to red (~590 nm) as the dye forms J-aggregates in energized mitochondria. The red/green fluorescence intensity ratio provides a quantitative measure of ΔΨm [68]. This methodology is particularly important for identifying cell populations with elevated mitochondrial membrane potential that may show enhanced MitoQ accumulation and response.
In Vivo Administration and Disease Models
In murine models, MitoQ is typically administered via intraperitoneal injection, intravenous injection, or oral gavage. For ischemia-reperfusion studies, a common protocol involves intravenous administration of 4 mg/kg MitoQ (adsorbed to β-cyclodextran in 100 μL saline) 15 minutes before the induction of ischemia [69]. In cancer therapy studies, oral administration of 18 mg/kg has been used in combination with radiotherapy [72]. For long-term studies investigating age-related conditions, MitoQ may be administered in drinking water (typically 100-500 μM) for extended periods ranging from weeks to months [73].
The following workflow illustrates a standard experimental approach for evaluating MitoQ effects in disease models:
Assessment of MitoQ's effects typically includes evaluation of oxidative stress biomarkers (SOD, catalase, GPx, MDA), mtDNA damage (8-hydroxyguanine content, mtDNA copy number, mitochondrial transcript levels), and mitochondrial function (respiratory complex activities, ATP production, mitochondrial membrane potential) [70] [74] [69]. Tissue-specific functional assessments are also critical, such as measurement of intestinal permeability using FITC-dextran in intestinal injury models, auditory brainstem response measurements in hearing loss studies, and tumor growth monitoring with caliper measurements or imaging in oncology models [70] [72] [69].
Research Reagent Solutions
Table 2: Essential Research Reagents for MitoQ Studies
| Reagent/Category | Specific Examples | Research Applications | Key Considerations |
|---|---|---|---|
| MitoQ Formulations [72] [69] | MitoQ (mitoquinone mesylate); Adsorbed to β-cyclodextran | In vivo administration; cellular studies | Water-soluble formulations improve bioavailability for in vivo use |
| Control Compounds [71] | dodecyl-TPP (dTPP); Untargeted ubiquinone | Distinguishing targeted vs. antioxidant effects | dTPP controls for TPP+ moiety effects without antioxidant capacity |
| Mitochondrial Dyes [68] [73] | JC-1; TMRE; MitoSOX Red | ΔΨm measurement; mitochondrial ROS detection | JC-1 provides ratio-metric measurement of membrane potential |
| Metabolic Assay Kits [72] | Seahorse XF ATP Rate Assay; SCENITH kit | Cellular bioenergetics; metabolic dependency | SCENITH allows single-cell analysis of metabolic dependencies |
| Oxidative Stress Biomarkers [74] [69] | SOD, CAT, GPx activity kits; MDA detection | Quantifying oxidative stress status | MDA measures lipid peroxidation as marker of oxidative damage |
| Pathway Inhibitors/Activators [68] [69] | Compound C (AMPK inhibitor); Nrf2 siRNA | Mechanistic pathway dissection | siRNA knockdown confirms pathway specificity |
| Cell Type-Specific Models [70] [73] [71] | Idh2⁻/⁻ mice; Dnmt3aR878H/+ HSPCs; hiPSC-CMs | Disease-specific mechanistic studies | Patient-derived cells enhance translational relevance |
MitoQ represents a paradigm shift in antioxidant therapy, demonstrating that subcellular targeting significantly enhances therapeutic efficacy while potentially reducing off-target effects. Its mechanism of action extends beyond simple antioxidant activity to include modulation of mitochondrial bioenergetics, induction of cytoprotective signaling pathways, and selective effects on cells with elevated mitochondrial membrane potential. The growing body of evidence supporting MitoQ's benefits across diverse disease models—including cardiovascular conditions, neurodegenerative disorders, cancer, hepatic diseases, and metabolic syndromes—highlights the fundamental role of mitochondrial oxidative stress in pathophysiology and the therapeutic potential of targeted interventions.
Future research directions should focus on optimizing dosing regimens for specific clinical applications, understanding the long-term effects of MitoQ administration, and identifying patient populations most likely to benefit from therapy. The selective accumulation of MitoQ in cells with elevated ΔΨm presents a particularly promising approach for targeting pathological cells without affecting normal tissues, as demonstrated in clonal hematopoiesis and cancer models [73] [72]. As research continues to elucidate the complex relationships between mitochondrial membrane potential, metabolic specialization, and disease progression, mitochondriotropic antioxidants like MitoQ are poised to play an increasingly important role in therapeutic development for conditions characterized by mitochondrial dysfunction.
Peripheral blood mononuclear cells (PBMCs) are increasingly recognized as valuable surrogate biomarkers in clinical and translational research. Comprising lymphocytes, monocytes, and dendritic cells, these accessible cells reflect systemic physiological and pathological processes, offering a "window" into the body's metabolic and immune status. This technical guide comprehensively examines the advantages and limitations of using PBMCs as surrogates, with particular emphasis on their application in studying mitochondrial membrane potential within metabolic specialization and partitioning research. We synthesize current methodologies, analytical frameworks, and validation approaches to provide researchers and drug development professionals with practical tools for implementing PBMC-based biomarker strategies in preclinical and clinical studies.
Peripheral blood mononuclear cells (PBMCs) represent a heterogeneous population of circulating immune cells, including lymphocytes (T cells, B cells, and NK cells), monocytes, and dendritic cells, that are isolated from peripheral blood via density gradient centrifugation [76] [77]. The use of PBMCs as surrogate biomarkers has gained substantial traction across diverse research domains, from oncology to metabolic disorders and neurodegenerative diseases. Their utility stems from their accessibility, dynamic responsiveness to physiological perturbations, and ability to mirror molecular alterations in less accessible tissues [76].
In the specific context of metabolic specialization and partitioning research, PBMCs offer a unique model system for investigating mitochondrial regulation and cellular bioenergetics. Mitochondrial membrane potential (ΔΨm), the electrochemical gradient across the inner mitochondrial membrane, serves as a crucial indicator of mitochondrial health and functional capacity. As key regulators of immunometabolic processes, PBMCs exhibit mitochondrial adaptations that reflect systemic metabolic states, making them valuable surrogates for investigating tissue-specific metabolic partitioning [78] [79]. The growing body of evidence supporting PBMCs as "surrogate biopsies" positions them as powerful tools for accelerating biomarker discovery and clinical translation in metabolic diseases [76].
Advantages of PBMC-Based Approaches
Accessibility and Practical Utility
The most significant advantage of PBMCs lies in their straightforward accessibility through minimally invasive venipuncture procedures, which can be performed by various healthcare professionals with minimal risk to patients [76]. This accessibility enables:
- Repeat sampling: Longitudinal monitoring of disease progression or treatment response through serial blood collections
- Immediate processing: Rapid isolation using standardized density gradient centrifugation protocols
- Cryopreservation feasibility: Successful maintenance of viability and functionality after long-term storage in biobanks
This practical utility makes PBMCs particularly suitable for large-scale clinical trials and longitudinal studies where repeated tissue sampling would be impractical or unethical [76].
Molecular Profiling Capabilities
PBMCs serve as a rich source for multi-omics investigations, providing comprehensive molecular profiling opportunities:
- Transcriptomic analysis: RNA sequencing and gene expression profiling reveal pathway alterations in response to disease states or interventions [80]
- Epigenetic characterization: Assessment of DNA methylation patterns and chromatin modifications that may reflect environmental exposures
- Proteomic profiling: Identification of protein expression and post-translational modifications relevant to disease mechanisms
- Metabolomic analysis: Investigation of metabolic fluxes and pathway activities
Single-cell RNA sequencing technologies have further enhanced the resolution of PBMC analyses, enabling the identification of distinct cellular subpopulations and their specific responses to pathological conditions [81].
Reflection of Systemic Processes
Perhaps the most compelling scientific advantage of PBMCs is their capacity to reflect systemic physiological and pathological processes. As circulating cells, PBMCs interact with various tissues and organs, acquiring molecular signatures that mirror broader biological states:
- Immune system monitoring: PBMCs naturally participate in immune surveillance and inflammatory responses, making them ideal for studying immunometabolic diseases [78]
- Metabolic status indication: Mitochondrial function in PBMCs correlates with systemic metabolic health and disease states [78] [82]
- Therapeutic response assessment: PBMCs can reveal drug target engagement and pharmacodynamic effects [77]
Relevance to Mitochondrial Research
In the specific context of mitochondrial membrane potential and metabolic partitioning research, PBMCs offer distinct advantages:
- Bioenergetic profiling: PBMCs exhibit measurable alterations in mitochondrial respiration parameters under different metabolic conditions [78]
- Dynamic responsiveness: PBMC mitochondrial function changes rapidly in response to nutritional interventions, exercise, and disease states
- Correlation with tissue metabolism: Studies demonstrate associations between PBMC mitochondrial parameters and metabolic phenotypes in other tissues [78]
Table 1: Key Advantages of PBMCs in Translational Research
| Advantage Category | Specific Benefits | Research Applications |
|---|---|---|
| Accessibility | Minimal invasive collection; Repeat sampling; Cost-effective processing | Large cohort studies; Longitudinal monitoring; Clinical trials |
| Molecular Profiling | Multi-omics compatibility; Single-cell resolution; High-content data generation | Biomarker discovery; Pathway analysis; Patient stratification |
| Systemic Reflection | Immune system representation; Metabolic status indication; Drug response assessment | Immunometabolic studies; Therapeutic monitoring; Pharmacodynamics |
| Mitochondrial Relevance | Bioenergetic profiling capability; Dynamic responsiveness; Tissue correlation | Metabolic specialization research; Mitochondrial dysfunction assessment |
Limitations and Technical Considerations
Biological Variability and Confounding Factors
The interpretation of PBMC-based data requires careful consideration of numerous biological and technical confounding factors:
- Cellular heterogeneity: The composition of PBMC populations varies significantly between individuals and under different physiological conditions, potentially obscuring specific signals [81]
- Diurnal rhythms: Gene expression and metabolic functions in PBMCs exhibit circadian oscillations that must be accounted for in study design
- Medication effects: Numerous drugs, particularly immunomodulators and metabolic agents, can alter PBMC biology [83]
- Comorbidities: Inflammatory conditions, infections, and metabolic disorders can independently influence PBMC parameters [83]
For core studies focusing on specific disease mechanisms, experts recommend strict exclusion criteria, including the absence of inflammatory or autoimmune diseases, active infections, recent surgeries or vaccinations, and use of immunomodulatory medications [83].
Technical Challenges in Processing and Analysis
Standardization of methodological approaches remains a significant challenge in PBMC research:
- Isolation variability: Density gradient centrifugation protocols can yield different cellular compositions and activation states based on technical execution
- Sample stability: Time between blood draw and processing significantly impacts RNA quality, protein phosphorylation, and mitochondrial function [77]
- Cryopreservation effects: Freezing and thawing procedures can alter cell viability, surface marker expression, and functional responses
Mitochondrial function assessment presents particular technical challenges, as respiration parameters are highly sensitive to processing conditions, substrate availability, and assay temperature [51].
Representation of Tissue-Specific Processes
A fundamental limitation of PBMC-based approaches concerns their capacity to accurately reflect tissue-specific processes:
- Metabolic partitioning: PBMCs may not fully recapitulate the specialized metabolic adaptations of non-immune tissues, such as skeletal muscle, liver, or adipose tissue
- Disease-specific alterations: While PBMCs show promise as surrogates in various conditions, the degree to which they mirror tissue-specific pathology varies by disease and individual
- Microenvironmental influences: PBMCs lack exposure to the unique tissue microenvironments that shape cellular metabolism and function in parenchymal organs
Table 2: Key Limitations of PBMC-Based Approaches and Mitigation Strategies
| Limitation Category | Specific Challenges | Potential Mitigation Strategies |
|---|---|---|
| Biological Variability | Cellular heterogeneity; Diurnal rhythms; Medication effects; Comorbidities | Strict inclusion/exclusion criteria; Uniform sampling times; Covariate adjustment in analysis |
| Technical Challenges | Isolation variability; Sample stability; Cryopreservation effects; Assay standardization | SOP development; Sample processing within 2-4 hours; Validation of freezing protocols |
| Tissue Representation | Metabolic partitioning differences; Disease-specific alterations; Microenvironmental influences | Correlation with tissue biopsies when feasible; Focus on systemic rather than tissue-specific processes |
Methodologies for PBMC Analysis in Metabolic Research
PBMC Isolation and Processing Protocols
Standardized protocols for PBMC isolation are critical for generating reproducible data:
Density Gradient Centrifugation Protocol
- Collect venous blood in EDTA or heparin Vacutainer tubes
- Dilute blood 1:1 with phosphate-buffered saline (PBS)
- Carefully layer diluted blood over Ficoll-Paque PLUS solution
- Centrifuge at 400-800 × g for 20-30 minutes at room temperature with brake disengaged
- Collect PBMC layer at the plasma-Ficoll interface
- Wash cells twice with PBS or culture medium
- Perform cell counting and viability assessment using trypan blue exclusion
For mitochondrial function assays, immediate processing is essential, as delays significantly impact respiratory parameters [82].
Assessment of Mitochondrial Membrane Potential
Mitochondrial membrane potential (ΔΨm) serves as a key metric of mitochondrial health in PBMCs:
Flow Cytometric Analysis of ΔΨm
- Resuspend fresh PBMCs at 1×10^6 cells/mL in culture medium
- Load cells with ΔΨm-sensitive fluorescent dyes (e.g., JC-1, TMRM, or TMRE)
- Incubate for 15-30 minutes at 37°C in the dark
- Wash cells to remove unincorporated dye
- Analyze fluorescence by flow cytometry using appropriate excitation/emission settings
- Include carbonyl cyanide m-chlorophenyl hydrazone (CCCP) as a depolarization control
JC-1 exhibits potential-dependent accumulation in mitochondria, indicated by a fluorescence emission shift from green (~529 nm) to red (~590 nm). The red/green fluorescence ratio provides a quantitative measure of ΔΨm that is largely independent of mitochondrial size, shape, and density [82].
Multiparameter Flow Cytometric Assays
Advanced flow cytometric approaches enable simultaneous assessment of multiple parameters in PBMC subpopulations:
DNA Damage Response Assessment [77]
- Fix PBMCs in 0.4% paraformaldehyde at 4°C overnight
- Permeabilize cells with ice-cold methanol or commercial permeabilization buffers
- Stain with primary antibodies against targets of interest (e.g., γH2AX for DNA damage)
- Incubate with fluorophore-conjugated secondary antibodies
- Analyze using a flow cytometer capable of detecting multiple fluorophores
- Use isotype controls and fluorescence-minus-one (FMO) controls for gating
This approach can be adapted to simultaneously assess mitochondrial membrane potential, mitochondrial mass, and cell surface markers to evaluate mitochondrial parameters in specific immune cell subsets.
Figure 1: Experimental Workflow for PBMC Isolation and Mitochondrial Analysis
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Research Reagents for PBMC Mitochondrial Studies
| Reagent Category | Specific Examples | Function & Application |
|---|---|---|
| Blood Collection Tubes | EDTA Vacutainers; Heparin Vacutainers; PAXgene Blood RNA Tubes | Anticoagulation and RNA stabilization for different downstream applications |
| Cell Isolation Media | Ficoll-Paque PLUS; Lymphoprep; Histopaque-1077 | Density gradient media for PBMC separation from whole blood |
| Viability Assessment | Trypan blue; Propidium iodide; LIVE/DEAD fixable dyes | Determination of cell viability before functional assays |
| Mitochondrial Dyes | JC-1; Tetramethylrhodamine (TMRM/ TMRE); MitoTracker dyes | Assessment of mitochondrial membrane potential and mass |
| Antibodies for Subsetting | Anti-CD3 (T cells); Anti-CD19 (B cells); Anti-CD14 (monocytes) | Identification of specific PBMC subpopulations by flow cytometry |
| DNA Damage Markers | Anti-γH2AX; Anti-MRE11; Anti-RAD51 | Assessment of DNA damage response pathways |
| Metabolic Probes | 2-NBDG (glucose uptake); MitoSOX (mitochondrial ROS) | Evaluation of metabolic activity and oxidative stress |
| Cell Preservation | DMSO; FBS; Cryostor solutions | Cryopreservation of PBMCs for long-term storage |
Applications in Disease Research and Clinical Translation
Oncology Applications
PBMCs have demonstrated significant utility in oncology research, particularly in assessing therapy response and resistance mechanisms:
- DNA damage response evaluation: PBMCs can reflect functional DNA repair capacity, which may predict response to DNA-damaging agents like platinum chemotherapy and PARP inhibitors [77]
- Immunomonitoring: Changes in PBMC subpopulations and their functional states provide insights into antitumor immune responses and immunotherapy efficacy
- Mitochondrial function assessment: PBMCs from lung cancer patients show significant mitochondrial dysfunction, suggesting their potential as biomarkers for cancer detection and monitoring [82]
In a study of high-grade serous ovarian cancer patients, PBMCs from non-responders to PARP inhibitor therapy showed significantly higher pre-treatment levels of γH2AX (DNA damage marker) and higher γH2AX/MRE11 ratios compared to responders, suggesting the potential for PBMC-based predictive biomarkers [77].
Metabolic Disease Research
PBMCs serve as valuable surrogates in metabolic disease research, reflecting systemic metabolic alterations:
- Obesity and weight loss interventions: Transcriptomic analysis of PBMCs reveals distinct gene expression patterns between hyper- and hypo-responders to caloric restriction [80]
- Mitochondrial functional assessment: PBMC mitochondrial respiration parameters correlate with metabolic health status and may reflect tissue-level mitochondrial adaptations [78]
- Inflammatory signaling: PBMCs mediate the interface between metabolism and inflammation, a key axis in metabolic disease pathogenesis
A recent study identified 1,581 differentially expressed genes in PBMCs from hyper- versus hypo-responders to weight loss interventions, with collagen biosynthesis pathways significantly enriched among downregulated genes in hypo-responders [80].
Neurodegenerative Disorders
Growing evidence supports the involvement of peripheral immunity in neurodegenerative diseases, positioning PBMCs as valuable peripheral windows into central nervous system pathology:
- Parkinson's disease: Studies demonstrate alterations in both innate and adaptive immune compartments in PBMCs from Parkinson's patients
- Immune profiling: Comprehensive immunophenotyping of PBMCs may enable immune-based stratification for clinical trials and treatment monitoring
Expert recommendations for Parkinson's disease research emphasize strict exclusion criteria for core studies, including absence of inflammatory/autoimmune diseases, active infections, and recent immunomodulatory medication use to minimize confounding factors [83].
PBMCs represent powerful surrogate biomarkers that balance accessibility with biological relevance, particularly in the context of mitochondrial research and metabolic specialization studies. Their capacity to reflect systemic processes while enabling detailed molecular and functional characterization positions them as invaluable tools for translational research and drug development.
Future advancements in PBMC-based approaches will likely focus on several key areas:
- Standardization of methodologies across laboratories to enhance reproducibility and comparability
- Single-cell multi-omics integration to resolve cellular heterogeneity and identify specific subpopulation responses
- Dynamic functional assessments that capture real-time metabolic adaptations in response to perturbations
- Correlation with tissue-specific analyses to validate PBMCs as faithful surrogates for particular disease contexts
As technologies advance and our understanding of PBMC biology deepens, these accessible cells will continue to provide critical insights into mitochondrial regulation, metabolic partitioning, and disease mechanisms, accelerating the development of targeted therapies and personalized medicine approaches.
Navigating Experimental Challenges: Pitfalls in MMP Measurement and Data Interpretation
Mitochondrial heterogeneity manifests at multiple interconnected levels, creating a complex landscape that researchers must navigate. Cellular heterogeneity refers to the variations in mitochondrial features between different cells, while mitochondrial heterogeneity encompasses variations within the mitochondrial population of a single cell [84] [85]. This heterogeneity extends to technical variability introduced by measurement tools and experimental conditions, which can significantly impact data interpretation. Within the specific context of mitochondrial membrane potential (ΔΨm) research, understanding these sources of variation is crucial for studying metabolic specialization and partitioning—the process by which mitochondria adopt specialized metabolic roles within cells [2].
The phenomenon of mitochondrial heterogeneity is driven by diverse factors including genetic variation, dynamic structural adaptations, and responses to localized cellular environments [84] [85]. Mitochondrial DNA (mtDNA) exhibits significant genetic variation both between cells and within individual cells, a state known as heteroplasmy [84]. Meanwhile, the mitochondrial proteome, lipid composition, and morphological characteristics demonstrate remarkable plasticity in response to physiological demands and pathological conditions [84]. This review systematically addresses the sources, measurement approaches, and functional implications of heterogeneity in mitochondrial research, with particular emphasis on its role in metabolic specialization.
Dimensions of Mitochondrial Heterogeneity
Genetic and Component Heterogeneity
Table 1: Sources and Characteristics of Mitochondrial Heterogeneity
| Dimension of Heterogeneity | Key Features | Functional Consequences | Experimental Assessment Methods |
|---|---|---|---|
| Genetic (mtDNA) | Heteroplasmy (mixed mutant/wild-type populations), microheteroplasmy (pervasive mutations), copy number variation [84] [85] | Bioenergetic output variability, disease susceptibility, threshold effects in phenotype expression [84] [85] | Single-cell sequencing, digital PCR, deep resequencing [84] |
| Proteomic & Lipidomic | Variable protein complexes stoichiometry, distinct lipid compositions (e.g., cardiolipin) [84] | Altered electron transport chain efficiency, membrane fluidity, supercomplex formation [84] | Mass spectrometry, blue native PAGE, protein import assays [84] |
| Morphological | Dynamic continuum between fragmented and fused states, varying cristae density [84] [2] | Quality control, apoptosis susceptibility, metabolic efficiency [84] [2] | Electron microscopy, live-cell imaging with targeted fluorophores [84] |
| Membrane Potential (ΔΨm) | Intramitochondrial gradients (cristae vs. IBM), inter-mitochondrial variation, transient depolarization [2] [6] | ATP production capacity, protein import efficiency, metabolic specialization [2] [6] | TMRM, JC-1, super-resolution potential mapping [2] [6] |
Spatial and Functional Heterogeneity in Membrane Potential
Mitochondrial membrane potential (ΔΨm) represents a key parameter exhibiting significant heterogeneity at multiple spatial scales. Across the mitochondrial population within single cells, individual organelles maintain different ΔΨm values depending on their functional state, with depolarized mitochondria targeted for degradation via mitophagy [2]. Remarkably, even within single mitochondria, electrical gradients exist between the cristae membranes (CM) and inner boundary membranes (IBM), with the CM typically maintaining a more hyperpolarized state (ΔΨC) compared to the IBM (ΔΨIBM) [6].
This intramitochondrial potential gradient is functionally significant, as it creates compartmentalized electrical domains regulated by cristae junctions (CJ) that act as barriers for ion movement [6]. The CJ serves as a potential overflow valve protecting mitochondrial integrity during excessive cristae hyperpolarization [6]. During calcium (Ca2+) uptake, the CM experiences greater hyperpolarization, likely due to Ca2+-sensitive enhancement of TCA cycle activity and subsequent oxidative phosphorylation in the cristae [6]. This spatial heterogeneity in ΔΨm enables functional specialization within individual mitochondria, with cristae functioning as the primary sites for ATP production while the IBM facilitates communication with the cytosol and other organelles.
Methodological Approaches for Resolving Heterogeneity
Advanced Imaging and Analytical Techniques
Super-resolution microscopy techniques have become indispensable for resolving mitochondrial heterogeneity, particularly for analyzing ΔΨm gradients within individual mitochondria. Structured Illumination Microscopy (SIM) and Stimulated Emission Depletion (STED) microscopy enable visualization of the spatial distribution of ΔΨm across mitochondrial subcompartments [6]. These approaches typically employ the potential-sensitive dye TMRM (tetramethylrhodamine methyl ester) in combination with IMM-specific reference markers such as MitoTracker Green FM (MTG) [6].
Table 2: Key Research Reagent Solutions for Assessing Mitochondrial Heterogeneity
| Research Tool | Specific Application | Key Utility in Heterogeneity Research | Technical Considerations |
|---|---|---|---|
| TMRM | Measurement of mitochondrial membrane potential [16] [6] | Quantifies ΔΨm heterogeneity; concentration-dependent distribution reveals intramitochondrial gradients [6] | Low concentrations (1.35-5.4 nM) preferentially accumulate in cristae; higher concentrations saturate both CM and IBM [6] |
| MitoTracker Green FM | Mitochondrial morphology reference [6] | Provides spatial reference for IMM structure; enables ratio imaging with potentiometric dyes [6] | Potential-independent IMM binding; enables normalization of potential-sensitive dye distribution [6] |
| MitoSOX | Detection of mitochondrial ROS [16] | Assesses heterogeneity in ROS production across mitochondrial populations [16] | Specific for superoxide; signal intensity correlates with oxidative stress heterogeneity [16] |
| Rhod-2 AM | Measurement of mitochondrial calcium [16] | Evaluates heterogeneity in Ca2+ handling capacity [16] | Accumulates in mitochondrial matrix; requires careful calibration for quantitative measurements [16] |
Two analytical methods have been developed to quantify these intramitochondrial potential gradients:
IBM Association Index: This fully automated method uses the MTG channel to define mitochondrial boundaries through Otsu thresholding. By systematically shrinking and widening these boundaries, it creates distinct regions of interest for the IBM and CM, then calculates their fluorescence intensity ratio [6].
ΔFWHM Method: This semi-automated approach analyzes the full width at half maximum (FWHM) of cross-sectional intensity profiles of both MTG and TMRM. The difference between their FWHMs indicates the extent of TMRM accumulation in cristae versus IBM regions [6].
The experimental workflow for assessing ΔΨm heterogeneity involves dual-channel super-resolution imaging of cells co-loaded with MTG (typically 500 nM) and concentration-optimized TMRM (1.35-81 nM, with 13.5 nM often used for dynamic studies) [6]. Following image acquisition, the distribution of TMRM fluorescence relative to the MTG reference is quantified using either the IBM association index or ΔFWHM method to determine relative ΔΨm differences between cristae and IBM compartments [6].
Correlative Multi-Parameter Microscopy
To address the complex interplay between different aspects of mitochondrial heterogeneity, correlative multi-parameter microscopy approaches have been developed that simultaneously monitor ΔΨm gradients, ATP production, and mitochondrial morphology [6]. This integrated methodology revealed that mitochondrial Ca2+ elevation preferentially hyperpolarizes the CM, likely through Ca2+-sensitive stimulation of TCA cycle activity and subsequent oxidative phosphorylation in cristae [6]. Furthermore, these studies demonstrated a direct correlation between CM hyperpolarization and enhanced ATP production, highlighting the functional significance of intramitochondrial ΔΨm gradients [6].
Diagram 1: Experimental workflow for resolving mitochondrial heterogeneity using super-resolution microscopy and multi-parameter analysis. CM: Cristae Membrane; IBM: Inner Boundary Membrane.
Functional Consequences: Metabolic Specialization and Partitioning
Membrane Potential in Metabolic Compartmentalization
Mitochondrial heterogeneity, particularly in ΔΨm, serves as a fundamental driver of metabolic specialization within cells. Research has revealed that mitochondria can be partitioned into distinct subpopulations dedicated to specific metabolic roles, with ΔΨm acting as a key regulator of this specialization [2]. Elevated ΔΨm promotes the filamentous assembly of enzymes such as pyrroline-5-carboxylate synthase (P5CS), driving reductive biosynthesis pathways including proline synthesis [2]. Conversely, reduced ΔΨm conditions inhibit this filamentation and shift mitochondrial function toward oxidative ATP production via OXPHOS [2].
This ΔΨm-dependent metabolic partitioning enables cells to maintain simultaneous oxidative and reductive programs in separate mitochondrial subpopulations [2]. The dynamic interplay between these specialized populations allows cells to rapidly adapt to changing metabolic demands. In neuronal systems, ΔΨm changes coordinate synaptic plasticity by linking metabolic state to structural changes at synapses, with mitochondrial recruitment to dendrites supporting localized protein synthesis for synaptic function [2]. This metabolic compartmentalization has particular significance in pathological contexts such as cancer, where enhanced reductive biosynthesis supports rapid cellular proliferation [2].
Quality Control and Population Dynamics
Mitochondrial heterogeneity plays a crucial role in quality control processes that maintain cellular health. The dynamic balance between mitochondrial fission and fusion creates a heterogeneous population that enables selective segregation of dysfunctional components [2]. Following fission events, the resulting daughter mitochondria exhibit different ΔΨm values, which determines their subsequent fate—those maintaining higher ΔΨm typically rejoin the network through fusion, while those with reduced ΔΨm are targeted for degradation via mitophagy [2].
This quality control mechanism relies on the heterogeneity of ΔΨm across the mitochondrial population to distinguish functional from dysfunctional organelles. The PINK1-Parkin pathway exemplifies this process, with reduced ΔΨm leading to PINK1 accumulation on the mitochondrial outer membrane, subsequent Parkin recruitment, and ultimately mitophagy [2]. This continuous monitoring and selective removal of depolarized mitochondria maintains the overall health of the mitochondrial network despite ongoing heterogeneity.
Diagram 2: Functional consequences of mitochondrial heterogeneity in metabolic specialization, quality control, and spatial organization.
Therapeutic Implications and Research Translation
Targeting Mitochondrial Heterogeneity in Drug Development
The growing understanding of mitochondrial heterogeneity has significant implications for therapeutic development, particularly for neurodegenerative diseases, metabolic disorders, and cancer. The neurodegenerative disorder therapeutics market is projected to reach USD 38.2 billion by 2029, with mitochondrial-targeted therapies representing a rapidly expanding segment [86]. Specifically, mitochondrial-based therapeutics for central nervous system indications are expected to grow at a remarkable CAGR of 97% from 2023 to 2029 [86].
Current therapeutic strategies targeting mitochondrial heterogeneity include:
Metabolic Optimization: Fine-tuning metabolic activities to stabilize energy production in neurons, including approaches to enhance electron transport chain efficiency [86].
Quality Control Enhancement: Promoting mitochondrial biogenesis and modulating mitophagy to maintain healthy mitochondrial populations [86].
Membrane Potential Stabilization: Developing compounds that modulate ΔΨm to prevent excessive heterogeneity that compromises cellular function [2] [87].
Antioxidant Approaches: Utilizing compounds like MitoQ and Coenzyme Q10 to mitigate heterogeneity driven by oxidative stress [87] [88].
The high failure rate of neurodegenerative drugs in clinical trials underscores the importance of considering mitochondrial heterogeneity in therapeutic development [86]. Precision medicine approaches that account for individual variations in mitochondrial profiles represent a promising direction for future therapies [86].
Technical Considerations for Therapeutic Screening
When developing screening platforms for mitochondrial-targeted therapies, researchers must account for technical variability in ΔΨm measurements. Key considerations include:
Dye Concentration Effects: TMRM distribution between CM and IBM compartments is concentration-dependent, with low concentrations (1.35-5.4 nM) preferentially accumulating in cristae, while higher concentrations (40.5-81 nM) saturate both compartments [6]. This necessitates careful optimization for each experimental system.
Cell Type Variations: Different cell lines exhibit distinct ΔΨm characteristics—highly glycolytic cells like HeLa show different response patterns compared to more oxidative cells like EA.hy926 [6].
Dynamic Responses: ΔΨm heterogeneity changes rapidly in response to stimuli such as Ca2+ elevation, requiring high-temporal resolution measurements to capture relevant dynamics [6].
Standardized assessment protocols that control for these technical variables are essential for reliable evaluation of therapeutic compounds targeting mitochondrial heterogeneity, particularly those aimed at modulating metabolic specialization in disease contexts.
Mitochondrial heterogeneity represents a fundamental biological phenomenon with profound implications for cellular function, metabolic specialization, and disease pathogenesis. The multidimensional nature of this heterogeneity—spanning genetic, structural, functional, and spatial domains—creates both challenges and opportunities for research and therapeutic development. Advanced methodological approaches, particularly super-resolution microscopy and multi-parameter correlation analysis, now enable researchers to resolve ΔΨm gradients at unprecedented spatial scales, revealing intricate relationships between membrane potential heterogeneity, metabolic partitioning, and quality control processes. As the field advances, integrating these sophisticated measurement techniques with therapeutic screening platforms will accelerate the development of targeted interventions that modulate mitochondrial heterogeneity to treat diverse human diseases.
The isolation of high-purity peripheral blood mononuclear cells (PBMCs) represents a critical preprocessing step for a wide array of downstream applications in immunology, drug development, and cellular research. Among various contamination challenges, platelet adherence remains a particularly pervasive issue that can profoundly compromise experimental outcomes. Platelets, or thrombocytes, are inherently sticky cell fragments that readily activate and adhere to mononuclear cells, especially monocytes, forming aggregates that impede accurate cell counting, reduce viability, and alter functional properties [89]. When PBMCs are utilized for mitochondrial function studies—a growing focus in metabolic research—platelet contamination introduces significant confounding variables by altering the metabolic readouts of the purified cell population.
The presence of platelets and their released factors can artificially influence cellular activation states, potentially masking true mitochondrial responses or inducing artifactual signatures [90]. For researchers investigating mitochondrial membrane potential in the context of metabolic specialization, ensuring a pristine PBMC population free from platelet contamination is therefore not merely a technical preference but a fundamental prerequisite for data integrity. This technical guide examines the sources of platelet contamination, provides optimized isolation protocols, details validation methodologies, and discusses the critical implications for mitochondrial function research.
Technical Challenges: How Platelet Contamination Compromises Research
Platelet contamination interferes with PBMC research through multiple mechanistic pathways, each presenting distinct challenges for downstream applications:
Physical Interference: Platelets form aggregates with PBMCs, particularly monocytes, creating cell clumps that obstruct accurate cell counting and flow cytometry analysis by altering light scatter properties [89]. These aggregates also reduce cell recovery during isolation procedures by entrapping viable cells in the pellet during washing steps.
Biological Activation: Activated platelets release a multitude of soluble factors including cytokines, chemokines, and growth factors that can pre-activate immune cells, potentially altering their transcriptional profile and responsiveness to experimental stimuli [90]. This is particularly problematic for assays measuring immune cell function such as proliferation, cytokine production, or metabolic adaptation.
Metabolic Interference: For research focused on mitochondrial membrane potential (ΔΨm) and metabolic partitioning, platelet contamination introduces significant confounding variables. Platelets themselves possess functional mitochondria and exhibit dynamic metabolic shifts upon activation, potentially skewing bulk measurements of PBMC mitochondrial function [91].
The sticky nature of platelets means they readily adhere to monocytes and other adhesive cell types in the PBMC fraction, creating a contamination problem that standard density gradient centrifugation alone cannot resolve [89]. Without specific interventions to address platelet removal, researchers risk conducting experiments on an impure cell population whose characteristics may not represent true biological responses.
Methodological Approaches: Optimized Protocols for Platelet Removal
Standardized Density Gradient Centrifugation with Modified Washing
The conventional Ficoll-Paque density gradient centrifugation effectively separates mononuclear cells from granulocytes and erythrocytes but does not adequately remove platelets, which co-localize with the PBMC layer due to their similar density [92]. The following optimized protocol incorporates specific modifications to minimize platelet retention:
Critical Steps for Platelet Reduction:
Blood Collection Considerations: Use acid citrate dextrose (ACD) as an anticoagulant rather than heparin, as it better preserves platelet integrity and reduces activation during collection. Employ a wide-bore needle (e.g., 19G) and avoid vigorous traction on syringe plungers to prevent mechanical platelet activation [89].
Temperature Optimization: Maintain reagents and samples at 18-20°C throughout the isolation procedure. Lower temperatures reduce platelet aggregation but can compromise PBMC separation efficiency, making 18-20°C the optimal range [92].
Low-Speed Centrifugation Washes: Following density gradient separation, perform washing steps at 120 × g for 10 minutes with the centrifuge brake disengaged [89]. These slow-spin conditions allow platelet suspension in the supernatant while pelleted PBMCs remain intact. Perfect balance of centrifuge buckets is essential at this low g-force to prevent pellet disruption.
Multiple Washes: Conduct at least three sequential wash steps with Ca++/Mg++-free phosphate-buffered saline (PBS) supplemented with 2% fetal bovine serum (FBS). The protein content helps prevent cell clumping while the absence of divalent cations minimizes platelet activation [89].
Advanced Techniques for High-Purity Applications
For research applications requiring exceptionally low platelet contamination, such as single-cell sequencing or precise mitochondrial function assays, additional strategies may be employed:
Immunomagnetic Depletion: Commercial systems like the EasySep Direct Human PBMC Isolation Kit incorporate antibodies targeting platelet-specific markers (e.g., CD41/CD61) for immunomagnetic depletion directly from whole blood, bypassing the need for multiple washing steps [89]. Alternatively, a platelet removal cocktail can be used following density gradient separation to further deplete residual platelets.
Sucrose Gradient Enhancement: For extreme platelet removal, a secondary centrifugation through a 4-20% sucrose gradient can be employed after initial PBMC isolation. Platelets remain at the sucrose interface while PBMCs pellet, effectively separating the populations [92].
Automated Isolation Systems: Emerging technologies like the FlowMagic platform utilize proprietary insert systems that demonstrate reduced contamination profiles, though independent validation specific to platelet removal remains limited [93].
Quality Assessment: Validating Platelet Removal and PBMC Function
Quantitative Contamination Assessment
Table 1: Methods for Assessing Platelet Contamination in PBMC Preparations
| Method | Target | Measurement Approach | Acceptance Threshold | Advantages |
|---|---|---|---|---|
| Flow Cytometry | CD41a/CD61+ events | Percentage of positive events in PBMC gate | <2% platelets in CD45+ population | High specificity, quantitative |
| Microscopy | Visual platelet rosettes | Counting cells with >5 adherent platelets | <5% of monocytes affected | Simple, no special equipment |
| Automated Cell Counting | Size-based discrimination | Platelet count in PBMC suspension | <10 platelets per 100 PBMCs | Rapid, high-throughput |
| ATP Assay | Metabolic activity | Luminescence-based ATP detection | Context-dependent baseline | Functional assessment |
Flow cytometry represents the gold standard for quantifying platelet contamination, utilizing antibodies against platelet-specific surface markers CD41a (GPIIb/IIIa) or CD61 (integrin beta-3) in conjunction with CD45 for leukocyte identification [89]. Samples should demonstrate less than 2% platelet positivity within the CD45+ population for most applications. For mitochondrial studies, even lower thresholds may be necessary.
Microscopic examination of Wright-Giemsa stained cytospin preparations allows visual identification of platelet rosettes on monocyte surfaces. While less quantitative, this method provides直观 evidence of contamination and can identify samples where platelets remain adherent despite low free platelet counts [89].
Functional Validation for Mitochondrial Research
For PBMCs destined for mitochondrial function assessment, additional validation is recommended:
Baseline Activation State: Measure expression of activation markers (e.g., CD69 on T cells, CD83 on monocytes) to ensure platelet contamination has not induced premature immune activation, which independently alters mitochondrial metabolism [90].
Mitochondrial Membrane Potential (ΔΨm) Consistency: Assess variability in ΔΨm using fluorescent probes such as TMRM (tetramethylrhodamine methyl ester) across multiple donors and isolations. High variability may indicate inconsistent platelet contamination or cellular activation [91].
Respiratory Capacity: Utilize extracellular flux analyzers to measure basal and maximal oxygen consumption rates, as platelet mitochondria can contribute to overall oxygen consumption measurements in contaminated samples [4].
Research Reagent Solutions: Essential Materials for Platelet-Free PBMC Isolation
Table 2: Key Reagents for Minimizing Platelet Contamination in PBMC Isolation
| Reagent/Category | Specific Examples | Function | Considerations for Mitochondrial Studies |
|---|---|---|---|
| Anticoagulants | Acid Citrate Dextrose (ACD) | Prevents coagulation while preserving platelet integrity | Superior to EDTA for cellular metabolism |
| Density Gradient Media | Ficoll-Paque PLUS, Lymphoprep | Separates PBMCs based on density | Use at 18-20°C for optimal separation |
| Wash Buffers | D-PBS without Ca++/Mg++ | Removes platelets without activation | Add 2% FBS to prevent cell clumping |
| Platelet Depletion Reagents | EasySep Platelet Removal Cocktail | Immunomagnetic depletion of platelets | Validated for compatibility with PBMCs |
| Mitochondrial Probes | TMRM, MitoSOX, Rhod-2AM | Assess mitochondrial function | Use validated concentrations to avoid artifacts [91] |
| Cell Viability Assays | Trypan blue, propidium iodide | Determine PBMC viability and counting accuracy | Platelet contamination falsely elevates counts |
Implications for Mitochondrial Membrane Potential Research
The investigation of mitochondrial membrane potential (ΔΨm) in the context of metabolic specialization and partitioning research demands exceptionally pure PBMC populations. Platelet contamination introduces several critical confounding factors that can compromise data interpretation:
Direct Metabolic Contribution: Platelets contain functional mitochondria capable of oxidative phosphorylation and maintaining ΔΨm [23]. While individual platelet mitochondrial content is low, significant contamination introduces a non-PBMC source of mitochondrial signals when using bulk measurement techniques.
Probe Uptake Interference: Fluorescent dyes used to measure ΔΨm (e.g., TMRM, JC-1) and other mitochondrial parameters (MitoSOX for ROS, Rhod-2AM for calcium) accumulate in all mitochondria present, not just those in PBMCs [91] [4]. The resulting composite signal thus reflects a weighted average of PBMC and platelet mitochondria, with potential discordance in their metabolic states.
Activation-Induced Metabolic Shifts: Platelet-derived microvesicles and soluble factors can activate PBMCs, particularly monocytes, inducing rapid metabolic reprogramming from oxidative phosphorylation to glycolysis—a shift accompanied by alterations in ΔΨm [94]. This pre-activation masks true baseline metabolic states and responsiveness to experimental stimuli.
Recent research on metabolic specialization in immune cells has revealed that distinct lymphocyte subsets maintain characteristic mitochondrial profiles that support their functional specialization [23] [4]. Platelet contamination obscures these subtle inter-population differences by introducing biological noise and potentially activating specific cell subsets, thereby reducing the resolution needed to detect authentic metabolic partitioning patterns.
Mitigating platelet contamination represents an essential component of rigorous PBMC isolation protocols, particularly for sophisticated applications like mitochondrial membrane potential assessment. The methodologies outlined herein—incorporating optimized density gradient centrifugation, strategic washing techniques, and appropriate quality assessment—provide researchers with a comprehensive framework for obtaining high-purity PBMC preparations. As research into metabolic specialization advances, with increasing focus on how mitochondrial membrane potential governs immune cell function and fate decisions, the elimination of technical artifacts stemming from platelet contamination becomes increasingly critical. By implementing these standardized approaches, researchers can enhance the reliability and interpretability of their mitochondrial function data, ensuring that observed phenomena reflect genuine biological processes rather than isolation artifacts.
Mitochondrial membrane potential (MMP) is a key parameter reflecting the health and functional state of mitochondria, serving as a central regulator of cellular bioenergetics. Beyond its canonical role in driving ATP synthesis, the MMP acts as a dynamic signaling hub that influences reactive oxygen species production, calcium handling, and mitochondrial quality control [2]. Recent research has revealed that the MMP facilitates metabolic specialization and compartmentalization within cells, where regional variations in membrane potential enable the emergence of functionally distinct mitochondrial subpopulations dedicated to specific metabolic tasks [2]. This partitioning is particularly important in pathological conditions such as cancer, where augmented substrate production supports rapid cellular proliferation.
Accurate measurement of MMP is therefore essential for understanding fundamental cellular processes, yet researchers face significant technical challenges related to the dynamic range limitations of potentiometric dyes. These fluorescent probes are prone to misinterpretation, as oversimplified interpretation of their signals can lead to important mistakes in both data acquisition and conclusions [4]. This technical guide addresses the critical optimization parameters for dye loading and concentration to ensure accurate assessment of mitochondrial function in the context of metabolic specialization research.
Quantitative Analysis of Mitochondrial Dyes
The selection of appropriate dyes and their optimal concentrations is fundamental to obtaining reliable MMP measurements. Different dyes offer varying advantages depending on the experimental requirements, including membrane potential sensitivity, photostability, and compatibility with specific analytical platforms.
Table 1: Comparative Analysis of Mitochondrial Dyes and Their Optimal Loading Conditions
| Dye Name | Primary Application | MMP Dependence | Optimal Concentration Range | Key Advantages | Technical Considerations |
|---|---|---|---|---|---|
| LDS 698 [95] | Tracking subtle MMP changes in live cells | High | Not specified | High sensitivity & specificity; Photostability; Non-toxicity | Suitable for microscopy, flow cytometry, and plate reader assays |
| MitoTracker Green FM [96] | Mitochondrial mass assessment | Independent | 50 nM (for T cells) | Insensitive to MMP changes; Bright signal; High photo-stability | Labels total mitochondrial pool (healthy and unhealthy) |
| TMRM [97] | High-throughput MMP screening | Dependent | Protocol-dependent washing | Suitable for 96/384-well plates; Quantitative MMP measurement | Requires washing steps to remove excess dye |
| TMRE [73] | MMP measurement in primary cells | Dependent | Not specified | Compatible with primary HSPCs; Low toxicity | Used for functional analysis in stem cell populations |
| MitoTracker Red (MTR) [98] | Mitochondrial localization studies | Dependent | 50 nM (for specific labeling) | Commercial availability; Reacts with thiols on mitochondrial proteins | Significant non-specific labeling concerns |
Table 2: Photobleaching and Phototoxicity Profiles of Common Mitochondrial Dyes [99]
| Dye Name | Phototoxicity Level | Key Morphological Impacts | Functional Impacts | Recommended Use Cases |
|---|---|---|---|---|
| NAO | High | Transformation from tubular to spherical shape; Reduced cristae density | Loss of MMP | Limited use for live-cell imaging |
| MTG (MitoTracker Green) | Moderate | Less morphological transformation | Better MMP preservation | General morphology studies |
| TMRE | Moderate | Less morphological transformation | Better MMP preservation | Prolonged live-cell imaging |
Experimental Protocols for Dye Optimization
Concentration Optimization for MitoTracker Green FM in T Cells
The following protocol establishes a standardized approach for determining optimal dye concentration using Imaging Flow Cytometry (IFC) [96]:
Cell Preparation: Isolate and stimulate T cells according to experimental requirements. For murine T cells, use wild-type OT-I TCR transgenic mice.
Initial Microscopy Validation:
- Label live T cells with 200 nM MitoTracker Green FM dye
- Confirm mitochondrial localization around the nucleus using widefield microscopy
- Establish expected staining pattern for subsequent IFC validation
Imaging Flow Cytometry Concentration Titration:
- Test a range of dye concentrations (e.g., 12.5 nM to 200 nM)
- For each concentration, standardize acquisition parameters including:
- Laser voltage (e.g., 200 mW, 440 mW, 560 mW)
- Fluorescence intensity values
- Processing parameters
- Include unstained controls for background subtraction
Optimization Criteria Assessment:
- Confirm mitochondrial localization matches microscopy patterns
- Eliminate false positive or negative staining
- Use Raw Max Pixel values to standardize dye concentration and voltage
- Establish 50 nM as optimal concentration for T cells [96]
Validation with Functional Assays:
- Combine with membrane potential-sensitive dyes (e.g., MitoTracker Red/Deep Red) to assess different physiological states
- Correlate staining patterns with functional readouts
TMRM-Based High-Throughput Screening Assay
This protocol enables MMP measurement in living cells in 96- or 384-well plates for drug screening applications [97]:
Cell Seeding and Treatment:
- Seed cells in appropriate multi-well plates
- Treat with test compounds according to experimental design
Dye Loading and Incubation:
- Incubate cells with TMRM fluorescent potentiometric probe
- Optimize incubation time based on cell type and dye penetration
Washing Procedure:
- Thoroughly wash cells to remove free compounds and probe
- Perform multiple washes to eliminate background fluorescence
Fluorescence Measurement:
- Measure retained TMRM fluorescence using an LJL Analyst fluorescence reader or similar instrument
- Ensure fluorescence intensity is proportional to MMP
Quality Control:
- Include control compounds (e.g., carbonylcyanide m-chlorophenyl hydrazone, dinitrophenol)
- Validate assay performance with Z' factor > 0.5 [97]
- Establish IC₅₀ values for reference compounds
LDS 698 Staining Protocol for Live-Cell Imaging
A novel application of LDS 698, a hemicyanine solid-state laser dye, provides high sensitivity for detecting subtle MMP changes [95]:
Dye Preparation: Prepare fresh staining solution according to manufacturer recommendations.
Cell Staining:
- Incubate live cells with LDS 698 dye
- Optimize incubation time and temperature for specific cell types
Multi-Modal Imaging Applications:
- Perform fluorescence microscopy with appropriate filter sets
- Conduct flow cytometry analysis for population-level assessment
- Utilize plate reader assays for high-throughput screening
Prolonged Live-Cell Imaging:
- Leverage dye robustness and non-toxicity for time-lapse experiments
- Monitor mitochondrial morphology and membrane potential dynamics
- Quantify changes in response to experimental manipulations
Critical Considerations for Dye Specificity and Limitations
Non-Specific Staining of Mitochondrial Dyes
A significant concern in MMP research is the non-specific accumulation of potentiometric dyes in cellular compartments other than mitochondria. Rosamine-based dyes like Mitotracker Red contain a mild thiol-reactive chloromethyl moiety that accumulates in mitochondria depending on MMP, but also reacts with thiols on proteins and peptides present in other organelles [98]. Thiol-containing peptides are present in numerous proteins located in mitochondria, endoplasmic reticulum, Golgi apparatus, and cell membrane [98]. While healthy mitochondria exhibit the highest membrane potential and thus the strongest signal, dyes can also be attracted and accumulated by other membrane structures, often resulting in weak signals that are frequently disregarded as background noise.
Horizontal Mitochondrial Transfer Artifacts
Mitotracker Red has commonly been employed to visualize mitochondrial localization and track mitochondrial movement between neighboring cells. However, comprehensive investigation reveals that mitochondrial dye transfer does not equal actual mitochondrial transfer [98]. When comparing the transfer efficiency of MTR with mito-targeted GFP, most recipient cells received the MTR signal rather than the genuine mitochondrial protein marker [98]. This discrepancy highlights that dye transfer can occur independently of actual organelle transfer, potentially leading to misinterpretation of intercellular mitochondrial trafficking studies.
Interpretation Challenges in OXPHOS Research
MMP measurements have low sensitivity and specificity for reporting changes in OXPHOS activity in coupled mitochondria [4]. The finite range of ΔΨm in coupled mitochondria creates inherent limitations for detection sensitivity. Furthermore, divergent changes in OXPHOS can be associated with the same MMP shift [4]. For example:
- Increased oxygen consumption with decreased MMP: Occurs when ATP synthesis increases and Δp consumption is not perfectly compensated by elevated electron transport
- Increased oxygen consumption with increased MMP: Occurs when the elevation in electron transfer rates exceeds ATP synthase capacity
These complexities necessitate complementary measurements of oxygen consumption rates alongside MMP assessment for accurate interpretation of mitochondrial function.
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Research Reagents for MMP Studies in Metabolic Specialization
| Reagent/Category | Specific Examples | Function/Application | Key Considerations |
|---|---|---|---|
| MMP-Sensitive Dyes | TMRM, TMRE, Mitotracker Red, Rhodamine-based probes | Quantitative MMP measurement | Membrane potential-dependent accumulation; Require careful concentration optimization |
| MMP-Independent Dyes | MitoTracker Green FM | Mitochondrial mass assessment | Insensitive to MMP; Labels total mitochondrial pool |
| Novel High-Sensitivity Probes | LDS 698 | Detecting subtle MMP changes | High photostability; Suitable for prolonged live-cell imaging |
| Inhibitors/Uncouplers | Oligomycin, FCCP, Rotenone, Antimycin A | Assay validation and controls | Essential for establishing assay dynamic range and specificity |
| Validation Tools | Mito-targeted GFP (COX8a, TOM20) | Specific mitochondrial labeling | Critical for verifying dye specificity and function |
| Advanced Imaging Platforms | Airyscan super-resolution, Imaging Flow Cytometry | High-resolution mitochondrial analysis | Enable quantification of morphology and function simultaneously |
Optimizing dye loading and concentration is not merely a technical exercise but a fundamental requirement for generating reliable data in mitochondrial membrane potential research. The dynamic range issues associated with potentiometric dyes demand rigorous validation and careful interpretation, particularly in the context of metabolic specialization where subtle changes in MMP regulate critical cellular decisions. By implementing the standardized protocols and considerations outlined in this technical guide, researchers can advance our understanding of how mitochondrial membrane potential serves as a central regulator of cellular function in health and disease. Future methodological developments should focus on enhancing dye specificity, improving quantitative interpretation, and enabling multi-parameter assessment of mitochondrial function in complex biological systems.
The study of mitochondrial dynamics, particularly the expression of fusion and fission proteins, is frequently marked by seemingly contradictory data. The proteins Mitochondrial Fission Process 1 (MTFP1) and MTFP2 serve as prime examples, with reported expression levels and functional impacts that vary significantly across different biological contexts, disease states, and even methodological approaches. Interpreting these conflicts is not merely an academic exercise; it is essential for accurate biomarker development and therapeutic targeting. This guide posits that these apparent contradictions can be largely resolved by framing them within the context of mitochondrial membrane potential (MMP) and its role in driving metabolic specialization and cellular partitioning. The MMP is not merely a bioenergetic parameter but a dynamic signaling hub that influences and is influenced by the mitochondrial fusion-fission equilibrium [2]. This document provides a technical framework for researchers and drug development professionals to decode conflicting data, offering structured experimental protocols and analytical tools to navigate this complex landscape.
Theoretical Framework: MMP as the Interpretative Key
The Central Role of Mitochondrial Membrane Potential
The mitochondrial membrane potential (MMP), a key component of the protonmotive force, is traditionally recognized for its role in driving ATP synthesis. However, emerging research underscores its function as a dynamic regulator of mitochondrial biology and cellular signaling. The MMP facilitates metabolic specialization within mitochondrial subpopulations, enabling cells to meet diverse energetic and biosynthetic demands [2]. Critically, the MMP is intrinsically linked to mitochondrial dynamics:
- Fate Determination Post-Fission: Following mitochondrial fission, the MMP of the resulting daughter mitochondrion determines its cellular fate. Fragments that retain a higher MMP relative to baseline often re-fuse with the network, while those with a lower MMP are targeted for degradation via mitophagy [2]. This creates a binary outcome based on an MMP threshold.
- Regulation of Protein Import: The import of nuclear-encoded proteins into mitochondria, including some dynamics regulators, depends on the MMP. Regional variations in MMP could therefore create heterogeneous protein composition, influencing the functional output of fusion and fission events [2].
- Metabolic Switching: Elevated MMP can enhance the activity of enzymes like pyrroline-5-carboxylate synthase (P5CS), promoting a switch from oxidative metabolism to reductive biosynthesis. This metabolic partitioning is often supported by distinct mitochondrial subpopulations, the formation of which is influenced by dynamics processes [2].
Context-Dependency of Fusion/Fission Proteins
The expression and function of mitochondrial dynamics proteins are not static. MTFP1, for instance, exhibits pleiotropic effects across tissues. It is a central regulator of mitochondrial dynamics, bioenergetic homeostasis, and stress adaptation, with particularly critical roles in cardiac and skeletal muscle [100]. Its dysfunction is implicated in diverse pathologies, from cardiovascular diseases to cancer [100] [101]. The protein's impact is mediated through its ability to integrate metabolic status with organelle dynamics and quality control, particularly through the ROS-calcium axis, which exhibits cell-type-specific regulation [100]. This inherent pleiotropy and tissue specificity form the biological basis for the conflicting expression data often observed in the literature.
Quantitative Data Synthesis: Conflicting Expression in Context
Empirical evidence consistently reveals context-dependent expression patterns for key mitochondrial dynamics proteins. The following tables synthesize quantitative findings to illustrate these conflicts.
Table 1: Context-Dependent Expression and Diagnostic Value of MTFP1 and MTFP2
| Protein | Biological Context | Expression Trend | Statistical Significance (p-value) | Diagnostic/Prognostic Value |
|---|---|---|---|---|
| MTFP1 | Prostate Cancer (TCGA) | Significant differential expression in tumor vs. normal | p = 0.0398 [101] | AUC = 0.5904 (95% CI: 0.5128-0.6680) [101] |
| MTFP1 | Prostate Cancer (TCGA) | Not significantly associated with tumor development | p = 0.2354 (Comparison with Normal method) [101] | Not a significant survival biomarker in this analysis [101] |
| MTFP2 | Prostate Cancer (TCGA) | Highly significant upregulation in tumor tissue | p = 7.06e-06 [101] | AUC = 0.698, suggesting potential as a diagnostic biomarker [101] |
| MTFP1 | High-energy-demand tissues (e.g., heart, muscle) | High expression levels [100] | N/A | Guardian of mitochondrial health; protector against ischemia-reperfusion injury [100] |
Table 2: Functional Roles of MTFP1 in Cellular Physiology and Disease
| Functional Domain | Core Function | Associated Mechanisms | Disease Implications |
|---|---|---|---|
| N-terminal GTP-binding Domain | GTP binding/hydrolysis, driving mitochondrial outer membrane fusion [100] | Conformational changes in outer membrane proteins [100] | Dysfunction leads to mitochondrial fragmentation [100] |
| Transmembrane Domains | Anchors protein to inner mitochondrial membrane (IMM) [100] | Regulates membrane permeability and respiratory function [100] | Associated with oxidative stress and cell death [100] |
| C-terminal Dimerization Domain | Forms homodimers/heterodimers (e.g., with MTFP2) [100] | Essential for effective mitochondrial outer membrane fusion [100] | Implicated in cardiovascular disorders, myopathies, and cancer [100] |
| ROS-Calcium Axis Modulation | Integrates metabolic status with organelle quality control [100] | Cell-type-specific regulation of stress responses and cell fate [100] | Differential impacts in disease states; potential therapeutic target [100] |
Experimental Protocols for Resolving Conflicts
To systematically investigate the context-dependent nature of fusion/fission proteins, researchers require robust and reproducible methodologies. The following protocols are designed to dissect the relationship between protein expression, MMP, and functional outcomes.
Protocol 1: Simultaneous MMP and Protein Localization Imaging
This protocol details a method for correlating real-time MMP with the subcellular localization and activity of mitochondrial dynamics proteins.
Cell Staining and Live-Cell Imaging:
- Step 1: Culture cells in a glass-bottom imaging dish.
- Step 2: Load cells with the potentiometric dye Tetramethylrhodamine, Methyl Ester (TMRM) at 20-50 nM in imaging buffer for 30 minutes at 37°C. Note: Use a low dye concentration and avoid washing to ensure quantitative measurements without inducing toxicity.
- Step 3: Transfect or transduce cells with a genetically encoded fluorescent protein (e.g., GFP) tagged to the mitochondrial dynamics protein of interest (e.g., MTFP1).
- Step 4: Perform live-cell confocal microscopy. Acquire TMRM signal (excitation/emission ~548/573 nm) to monitor MMP and GFP signal (excitation/emission ~488/507 nm) to monitor protein localization.
Image Analysis and Correlation:
- Step 5: Quantify the fluorescence intensity of TMRM and the fusion/fission protein (GFP) on a per-mitochondria or regional basis using image analysis software (e.g., ImageJ/FIJI, Volocity).
- Step 6: Correlate MMP (TMRM intensity) with the accumulation of the dynamics protein. For instance, test the hypothesis that MTFP1 accumulates preferentially in mitochondrial subpopulations with a specific MMP range.
Protocol 2: Functional Validation via Genetic Perturbation and Metabolic Profiling
This protocol assesses the functional consequences of modulating fusion/fission protein expression on mitochondrial metabolism and network morphology.
Genetic Manipulation:
- Step 1: Generate stable knockdown or knockout cell lines of the target gene (e.g., MTFP1) using CRISPR/Cas9 or shRNA. Use a non-targeting scramble sequence as the control.
- Step 2: Confirm knockdown/knockout efficiency via western blotting and qPCR.
Metabolic and Morphological Phenotyping:
- Step 3: Analyze the cellular metabolic profile using a Seahorse XF Analyzer. Measure key parameters of mitochondrial function, including Basal Respiration, ATP-linked Respiration, Maximal Respiration, and Proton Leak.
- Step 4: Quantify the MMP using TMRM flow cytometry in conjunction with the Seahorse data to link bioenergetic function to membrane potential.
- Step 5: Fix cells and immunostain for the mitochondrial network (e.g., using TOMM20 antibody). Acquire confocal images and perform high-content analysis to quantify network morphology: mean branch length, number of branches, and number of individual mitochondria per cell.
Signaling Pathways and Logical Workflows
The relationship between MMP, mitochondrial dynamics, and cellular function can be conceptualized as a series of interconnected pathways. The following diagrams, generated using Graphviz DOT language, map these critical relationships and experimental workflows.
MMP and Fission in Mitochondrial Fate Decision
This diagram illustrates how mitochondrial membrane potential acts as a key determinant in the fate of a mitochondrion following a fission event.
Experimental Workflow for Contextual Data Integration
This workflow provides a logical framework for designing experiments to resolve conflicting data on fusion/fission protein expression.
The Scientist's Toolkit: Essential Research Reagents
Successfully navigating the complexities of context-dependent protein expression requires a carefully selected set of reagents and tools. The following table catalogs essential solutions for this field of research.
Table 3: Research Reagent Solutions for Mitochondrial Dynamics Studies
| Reagent/Tool Category | Specific Examples | Function & Application Note |
|---|---|---|
| Potentiometric Dyes | TMRM, TMRE, JC-1 | Quantitative measurement of MMP. TMRM is preferred for low-dose, non-quenched mode for live-cell kinetic studies [2]. |
| Genetic Encoded MMP Sensors | CEPIA-mt, mt-cpYFP | Allow targeted, compartment-specific measurement of MMP and enable correlation with other fluorescently tagged proteins [2]. |
| Mitochondrial Stains | MitoTracker Deep Red, MitoTracker Green | Label mitochondrial mass independent of membrane potential; useful for network morphology analysis. |
| Antibodies for IHC/IF | Anti-MTFP1, Anti-TOMM20, Anti-PINK1 | Validate protein expression and localization via Western Blot, Immunohistochemistry (IHC), and Immunofluorescence (IF). |
| Genetic Manipulation Tools | CRISPR/Cas9 (knockout), shRNA (knockdown), siRNA (transient knockdown) | For functional validation studies to establish causal relationships between protein expression and phenotype. |
| Bioinformatics Databases | The Cancer Genome Atlas (TCGA), STRING, Cytoscape with CytoHubba | Analyze differential gene expression, build protein-protein interaction networks, and identify hub genes (e.g., MCC algorithm) [101]. |
| Metabolic Profiling Instruments | Seahorse XF Analyzer | Directly measure key parameters of mitochondrial respiration and glycolytic function in live cells. |
Mitochondrial membrane potential (MMP) is a fundamental biophysical parameter, reflecting the charge separation across the inner mitochondrial membrane generated by the electron transport chain (ETC). It is the principal component of the protonmotive force (PMF), which drives adenosine triphosphate (ATP) synthesis [2]. Traditionally, MMP has been viewed as a static indicator of mitochondrial health. However, contemporary research reveals that MMP is a highly dynamic signaling hub that undergoes rapid fluctuations in response to cellular energy demands and stress signals [2]. These dynamics are not mere consequences of cellular status but are active regulators of critical processes, including reactive oxygen species (ROS) production, calcium handling, and mitochondrial quality control via mitophagy [2]. Capturing these dynamics, rather than relying on single timepoint measurements, is therefore essential for understanding mitochondrial function in health and disease, particularly in the context of metabolic specialization and partitioning.
The Critical Role of MMP Dynamics in Metabolic Specialization
Emerging evidence indicates that mitochondria within a single cell can specialize metabolically, and MMP dynamics are central to this partitioning. Two primary metabolic programs exist: oxidative metabolism, which relies on the ETC to generate MMP for ATP production, and reductive metabolism, which supports biosynthetic processes for cellular replication [2].
The dynamic partitioning of metabolic enzymes is influenced by changes in MMP. For instance, the activity of pyrroline-5-carboxylate synthase (P5CS), a key enzyme in proline biosynthesis, is enhanced under elevated MMP. This promotes the formation of P5CS filaments that drive reductive biosynthesis [2]. Conversely, reduced MMP inhibits this filamentation, shifting mitochondrial function toward core oxidative phosphorylation (OXPHOS) and ATP generation [2]. This MMP-dependent mechanism facilitates the emergence of specialized mitochondrial subpopulations tailored to specific metabolic demands, a phenomenon with significant implications in pathologies like cancer, where augmented substrate production supports rapid proliferation [2].
Table 1: Key Functional Roles of MMP Dynamics in Cellular Physiology
| Functional Role | Mechanism | Impact of MMP Dynamics |
|---|---|---|
| Metabolic Specialization | MMP-dependent filamentation of P5CS enzyme [2]. | High MMP promotes reductive biosynthesis; low MMP favors oxidative ATP production. |
| Mitochondrial Quality Control | Loss of MMP triggers PINK1 accumulation, recruiting Parkin to initiate mitophagy [2]. | A decrease in MMP acts as a direct signal for the selective removal of damaged mitochondria. |
| Neuronal Plasticity | Changes in MMP coordinate synaptic plasticity and dendritic spine remodeling [2]. | MMP dynamics link the metabolic state of a neuron to structural changes at synapses. |
| Protein Import | Positively charged mitochondrial pre-sequence are pulled into the matrix by the MMP [2]. | MMP regulates the import of nuclear-encoded proteins, influencing mitochondrial composition. |
Advanced Tools for Tracking MMP Dynamics
Fluorescent Probes for Live-Cell Imaging
A primary method for monitoring MMP in live cells involves the use of potentiometric fluorescent dyes. While traditional dyes exist, recent advancements have introduced more robust probes.
LDS 698, a hemicyanine solid-state laser dye, has recently been reported as a highly sensitive probe for tracking MMP. It exhibits high specificity for active mitochondria and a superior ability to detect subtle changes in MMP compared to many commercial tools [95]. Its key advantages include robustness, photostability, and non-toxicity, making it suitable for prolonged live-cell imaging across various analytical techniques like fluorescence microscopy, flow cytometry, and plate reader assays [95].
Table 2: Research Reagent Solutions for MMP Analysis
| Reagent / Tool | Function / Application | Key Features |
|---|---|---|
| LDS 698 Probe [95] | Tracking MMP dynamics in live cells via fluorescence. | High sensitivity, photostable, non-toxic, suitable for prolonged imaging. |
| MoDL (Deep Learning Algorithm) [102] | Mitochondrial image segmentation and function prediction. | Predicts functions (e.g., MMP, ROS) from morphology; trained on 100,000+ super-resolution images. |
| Super-Resolution Microscopy (e.g., SIM) [102] | High-resolution live-cell imaging of mitochondrial morphology. | ~115 nm resolution enables precise segmentation of mitochondrial contours. |
Deep Learning for Morpho-Functional Prediction
The link between mitochondrial morphology and function presents an opportunity to predict the latter by analyzing the former. A novel deep learning algorithm named MoDL (Mitochondrial deep learning) has been developed for this purpose [102]. Trained on a massive dataset of over 100,000 super-resolution images, each annotated with functional data from biochemical assays, MoDL can predict various mitochondrial functions, including MMP, respiration rate, ROS production, and ATP generation, directly from live-cell images [102]. This approach demonstrates that complex MMP dynamics and other functional parameters can be inferred from high-quality morphological data, enabling large-scale and non-invasive functional screening.
Experimental Protocols for Capturing MMP Dynamics
Protocol: Tracking MMP with LDS 698 Staining and Live-Cell Imaging
This protocol outlines the procedure for using LDS 698 to monitor MMP dynamics in live cells [95].
Key Materials:
- LDS 698 dye [95]
- Appropriate cell culture vessels (e.g., glass-bottom dishes for microscopy)
- Live-cell imaging setup with fluorescence capabilities
- Imaging medium (without phenol red, serum-free to reduce background)
Methodology:
- Cell Preparation: Plate cells in appropriate glass-bottom dishes and culture until they reach the desired confluence.
- Staining: Incubate cells with LDS 698 dye at a concentration and duration optimized for your cell type (e.g., 100-500 nM for 15-30 minutes at 37°C).
- Washing: Gently wash the cells 2-3 times with pre-warmed imaging medium to remove excess, non-specific dye.
- Image Acquisition:
- Place the dish on a pre-warmed microscope stage maintaining 37°C and 5% CO₂.
- Acquire time-lapse images using the appropriate excitation/emission filters for LDS 698.
- To capture dynamics, set a time interval that matches the biological process under investigation (e.g., every 30-60 seconds over 1-2 hours).
- Include positive controls (e.g., cells treated with an uncoupler like FCCP to depolarize membranes) and negative controls (untreated cells) in each experiment.
- Data Analysis: Quantify fluorescence intensity over time using image analysis software (e.g., ImageJ). A decrease in signal indicates MMP depolarization, while an increase indicates hyperpolarization.
Protocol: Predicting MMP via Morphology Using MoDL
This protocol describes how to use the MoDL algorithm to predict mitochondrial functions, including MMP, based on morphological features from live-cell images [102].
Key Materials:
- High-quality live-cell fluorescence images of mitochondria (super-resolution images, such as Structured Illumination Microscopy (SIM), are recommended for training and yield the best results) [102].
- MoDL software package [102].
Methodology:
- Image Acquisition: Capture high-resolution images of mitochondria in your live cells of interest. The higher the image quality, the more accurate the segmentation and subsequent prediction will be.
- Image Segmentation:
- Input your images into MoDL's first pipeline.
- The algorithm, built on an enhanced U-Net architecture with residual networks and attention modules, will perform high-precision segmentation of individual mitochondrial contours.
- The output is a mask that defines the boundaries of each mitochondrion.
- Morphological Feature Extraction: MoDL automatically extracts comprehensive morphological features from the segmented images, such as area, form factor, and branch length.
- Function Prediction:
- Feed the extracted morphological features into MoDL's second pipeline, which employs an ensemble learning strategy.
- The model, trained on a vast dataset linking morphology to biochemical assay data, will output predictions for mitochondrial functions, including MMP.
- Validation: For novel cell types or conditions, it is critical to validate MoDL's predictions against a traditional biochemical assay for MMP (e.g., using potentiometric dyes like LDS 698 or TMRM) on a subset of samples.
Signaling Pathways and Experimental Workflows
The relationship between mitochondrial dynamics, MMP, and cellular function is complex. The following diagrams, created using Graphviz, illustrate the core signaling pathway and a generalized experimental workflow.
MMP in Metabolic Fate Decisions
Workflow for Live-Cell MMP Analysis
Mitochondrial membrane potential (ΔΨm or MMP) is a fundamental component of cellular bioenergetics, serving as the primary driver for ATP synthesis. However, its role extends far beyond this canonical function. Recent research has illuminated how MMP acts as a dynamic signaling hub that regulates metabolic specialization and compartmentalization within cells [2]. The MMP facilitates the emergence of specialized mitochondrial subpopulations, each tailored to specific metabolic demands—a process of particular importance in pathological conditions such as cancer, where augmented substrate production supports rapid cellular proliferation [2].
Despite its biological significance, the assessment of MMP faces substantial standardization challenges. Widely used fluorescent dyes are prone to misinterpretation, as oversimplified interpretation of their signals can lead to significant errors in both data acquisition and conclusions [4]. This technical guide addresses the critical gaps in MMP assessment protocols, providing researchers with standardized methodologies to enhance reproducibility, particularly within the context of metabolic partitioning research.
Foundational Concepts: MMP Gradients and Compartmentalization
The inner mitochondrial membrane (IMM) is not a uniform structure but is divided into two main compartments with potentially different electrical potentials: the cristae membrane (CM), which houses the electron transport chain complexes, and the inner boundary membrane (IBM), which connects to the outer membrane [6]. The crista junction (CJ) acts as a barrier separating these compartments, regulating ion movement and potentially maintaining distinct electrical potentials across the cristae (ΔΨC) and inner boundary (ΔΨIBM) membranes [6].
This compartmentalization has profound functional implications. Recent super-resolution microscopy studies have demonstrated that mitochondrial Ca2+ elevation hyperpolarizes the CM, likely caused by Ca2+-sensitive increase of TCA cycle activity and subsequent oxidative phosphorylation in the cristae [6]. This spatial organization of MMP enables specialized functions, with regional variations in MMP influencing how mitochondrial fragments are sorted during quality control processes and directing metabolic enzymes to establish distinct mitochondrial subpopulations [2].
Figure 1: Mitochondrial Compartmentalization and MMP Gradients. The crista junction (CJ) creates distinct compartments with potentially different membrane potentials, regulating metabolite and ion exchange.
Critical Standardization Gaps in Current MMP Assessment
Methodological Variability and Dye-Related Artifacts
The assessment of MMP relies heavily on potentiometric fluorescent dyes, yet significant inconsistencies in their application undermine reproducibility across studies. Four key methodological gaps have been identified:
Dye Concentration Sensitivity: The distribution of commonly used dyes like TMRM between CM and IBM is highly concentration-dependent. At low concentrations (1.35-5.4 nM), TMRM accumulates preferentially in the cristae, while higher concentrations (13.5-81 nM) lead to saturation and increased IBM staining [6]. This creates substantial variability in measured fluorescence patterns based solely on dye concentration.
Interpretation Complexity: Fluorescence intensity is often simplistically interpreted as directly proportional to mitochondrial "health" or "activity." However, MMP has a narrow dynamic range in coupled mitochondria, and the same ΔΨm shift can be associated with divergent changes in oxidative phosphorylation [4]. An increase in fluorescence may not always indicate improved mitochondrial function.
Compartmentalization Oversight: Most conventional protocols fail to account for spatial membrane potential gradients between cristae and IBM compartments [6]. Standard widefield microscopy cannot resolve these sub-mitochondrial domains, requiring super-resolution techniques for accurate assessment.
Coupling State Ambiguity: Measurements frequently do not distinguish between coupled, leaky, and uncoupled mitochondrial states, leading to misinterpretation of MMP data in relation to actual ATP synthesis capacity [4].
Technology-Dependent Limitations
The resolution limits of conventional microscopy techniques present another significant standardization challenge. Traditional light microscopy is limited to approximately 200 nm resolution due to light diffraction, while critical mitochondrial ultrastructural features are on a much smaller scale: crista junctions range from 12-40 nm in diameter, and cristae lumen widths are approximately 25-30 nm [103]. This fundamental technical limitation means that widely available fluorescence microscopes cannot resolve the very structures that maintain MMP gradients, creating an inherent methodological gap in most studies.
Table 1: Resolution Requirements for Mitochondrial Ultrastructure Analysis
| Mitochondrial Feature | Typical Size Range | Imaging Requirement | Standard Protocol Gap |
|---|---|---|---|
| Crista Junction (CJ) | 12-40 nm | Super-resolution microscopy (STED, SIM) | Conventional fluorescence microscopy (≥200 nm) |
| Cristae Lumen | 25-30 nm | Electron tomography, cryo-EM | Cannot be resolved with standard techniques |
| Complete IMM | ~20 nm thickness | Nanoscopy (MINFLUX: ≤5 nm) | Substructural compartments not distinguished |
| MMP Gradients (ΔΨC vs ΔΨIBM) | <100 nm separation | Super-resolution with potentiometric dyes | Bulk MMP measurement without spatial resolution |
Standardized Methodologies for Reproducible MMP Assessment
Quantitative SMPG Analysis Protocol
The Spatial Membrane Potential Gradient (SMPG) analysis provides a standardized approach for assessing sub-mitochondrial potential differences using super-resolution microscopy. This method enables researchers to quantify relative differences between cristae and IBM membrane potential [6].
Experimental Workflow:
- Cell Staining: Co-stain cells with 500 nM MitoTracker Green FM (MTG) and low concentrations (1.35-5.4 nM) of TMRM
- Image Acquisition: Perform simultaneous dual-channel imaging using structured illumination microscopy (SIM)
- Image Analysis: Apply one of two standardized analysis methods:
- IBM Association Index: Automated thresholding of MTG channel to define mitochondrial boundaries, with calculation of fluorescence intensity ratio between IBM and CM regions
- ΔFWHM Method: Analysis of full width at half maximum differences between cross-section intensity profiles of MTG and TMRM
Validation Controls:
- Hyperpolarization Control: Oligomycin A (ATP synthase inhibitor) should increase fluorescence intensity
- Depolarization Control: FCCP (uncoupler) should decrease fluorescence intensity
- Inhibition Controls: Rotenone (Complex I inhibitor) and Antimycin A (Complex III inhibitor) should block histamine-induced changes in IBM association index
Figure 2: Standardized SMPG Analysis Workflow. This protocol enables quantitative assessment of spatial membrane potential gradients using super-resolution microscopy and dual staining approaches.
Real-Time Live-Cell MMP Monitoring
For longitudinal studies of MMP dynamics, the Incucyte Mitochondrial Membrane Potential Assay enables real-time detection of transient and long-term changes in MMP in live cells [104]. This platform can be multiplexed with cell labeling, apoptosis, or cytotoxicity reagents for additional measurements of cell health.
Standardized Protocol:
- Reagent Preparation: Utilize cell-permeable MMP Orange Reagent that accumulates in active mitochondria in proportion to ΔΨm
- Baseline Acquisition: Establish baseline fluorescence with automated image acquisition every 1-4 hours
- Experimental Modulation: Apply compounds of interest with appropriate controls:
- FCCP (uncoupling agent) for depolarization control
- Oligomycin A (ATP synthase inhibitor) for hyperpolarization control
- Multiplexing Capability: Combine with Caspase-3/7, Annexin V, or Cytotox dyes for simultaneous assessment of apoptosis and cytotoxicity
- Quantitative Analysis: Generate automated, quantitative fluorescence intensity measurements over time
Integrated Functional Assessment Framework
To address the limitation of MMP as a standalone parameter, an integrated assessment framework is essential for comprehensive mitochondrial functional analysis. This multi-parameter approach contextualizes MMP measurements within broader bioenergetic status.
Complementary Assays:
- Oxygen Consumption Rate (OCR): More sensitive parameter for detecting changes in OXPHOS activity in coupled mitochondria [4]
- ATP Production Measurements: FRET-based ATP biosensors to correlate MMP with energetic output [6]
- Morphometric Analysis: Mitochondrial count, aspect ratio, and form factor quantification [6]
- Calcium Imaging: Simultaneous monitoring of mitochondrial Ca2+ uptake and its effects on MMP
Table 2: Research Reagent Solutions for Standardized MMP Assessment
| Reagent/Category | Specific Examples | Function & Mechanism | Standardized Concentration Ranges |
|---|---|---|---|
| Potentiometric Dyes | TMRM, TMRE, JC-1 | Accumulate in mitochondria in potential-dependent manner; fluorescence indicates MMP | Low: 1.35-5.4 nM (gradient analysis)High: 13.5-81 nM (bulk assessment) |
| Reference Dyes | MitoTracker Green FM (MTG) | IMM reference marker independent of membrane potential changes | 500 nM (for SMPG analysis) |
| Depolarization Controls | FCCP, CCCP | Protonophores that dissipate MMP by uncoupling ETC | 1-5 µM (concentration-dependent effects) |
| Hyperpolarization Controls | Oligomycin A | ATP synthase inhibitor that increases MMP by reducing proton flux | 1-10 µM (titrate for cell type) |
| ETC Inhibitors | Rotenone (CI), Antimycin A (CIII) | Inhibit electron transport, validate MMP dependence on ETC activity | Rotenone: 100-500 nMAntimycin A: 1-5 µM |
| IMM Structure Modulators | Mic60/MICOS disruptors | Alter cristae junction integrity, test compartmentalization dependence | Compound-specific titration required |
| Multiplexing Apoptosis Dyes | Annexin V, Caspase-3/7 | Distinguish MMP changes associated with programmed cell death | Manufacturer recommended concentrations |
Implementation Guidelines and Validation Framework
Standardized Reporting Requirements
To enhance reproducibility across studies, researchers should adopt minimum reporting standards for MMP assessment:
- Dye Information: Specific dye, concentration, incubation time, and loading conditions
- Imaging Parameters: Microscope type, resolution, temporal frequency, and analysis method
- Validation Controls: Inclusion of both depolarization (FCCP) and hyperpolarization (Oligomycin) controls in each experiment
- Contextual Data: Correlation with oxygen consumption rates or ATP production measurements
- Compartmentalization Assessment: Specification of whether spatial gradients were analyzed and at what resolution
Experimental Design Considerations
Cell Type Selection: Different cell lines exhibit varying metabolic profiles that significantly impact MMP dynamics. HeLa cells represent a strongly glycolytic phenotype, while EA.hy926 cells demonstrate higher OXPHOS activity [6]. This fundamental metabolic difference should guide model system selection and data interpretation.
Temporal Dynamics: MMP is a dynamic parameter that responds rapidly to cellular stimuli. Histamine-induced mitochondrial Ca2+ elevation produces detectable changes in SMPG within minutes [6]. Experimental designs must account for these temporal dynamics through appropriate time-course analyses rather than single endpoint measurements.
Technical Replication: Given the concentration-dependent distribution of potentiometric dyes [6], technical replication should include titration across the recommended concentration ranges (1.35-81 nM for TMRM) to verify consistent findings across multiple dye concentrations.
The standardization of MMP assessment protocols represents an essential advancement for understanding mitochondrial function in metabolic partitioning and specialization. By addressing the critical gaps in current methodologies—through implementation of spatial gradient analysis, standardized controls, and multi-parameter validation—researchers can significantly enhance the reproducibility and biological relevance of MMP measurements. The protocols and frameworks presented herein provide a foundation for consistent assessment of this crucial bioenergetic parameter across diverse research applications, from basic mitochondrial biology to drug development targeting metabolic diseases.
As the field advances, emerging technologies including MINFLUX nanoscopy (with resolution of 5 nm or less) [103] and CRISPR-based biosensors for compartment-specific MMP monitoring will further refine our understanding of mitochondrial membrane potential dynamics. By adopting standardized approaches today, the research community will be better positioned to integrate these future technological innovations into a cohesive understanding of mitochondrial bioenergetics in health and disease.
From Mechanism to Medicine: Validating MMP Dysfunction in Disease Pathogenesis
Mitochondrial Membrane Potential in Metabolic Reprogramming and Chemoresistance
Mitochondrial membrane potential (MMP) serves as a fundamental regulator of cancer cell metabolic plasticity, therapeutic response, and metastatic progression. This technical review synthesizes current research demonstrating how MMP heterogeneity drives metabolic specialization in cancer populations, facilitates resistance to conventional chemotherapies, and presents novel targeting opportunities for therapeutic development. We provide comprehensive quantitative analyses of MMP across cancer models, detailed methodological frameworks for assessing MMP dynamics, and visual schematics of the underlying molecular mechanisms governing these processes, with particular emphasis on the role of mitochondrial transfer from neural components in the tumor microenvironment.
Mitochondrial membrane potential, generated by the proton gradient across the inner mitochondrial membrane during oxidative phosphorylation, represents a crucial bioenergetic parameter that extends far beyond its role in ATP production. In cancer systems, MMP serves as a dynamic indicator of metabolic fitness, reflecting the balance between glycolytic and oxidative metabolic phenotypes [105]. The maintenance of distinct MMP states across cancer cell populations enables functional specialization, with hyperpolarized mitochondria supporting anabolic processes and depolarized states triggering adaptive survival mechanisms [106] [105]. This heterogeneity creates metabolic partitions within tumors that determine chemosensitivity profiles and metastatic potential.
Recent investigations have revealed that cancer cells can acquire mitochondria from stromal components, particularly neurons, through direct transfer mechanisms that enhance MMP and oxidative capacity [107]. This non-cell-autonomous enhancement of mitochondrial function represents a paradigm shift in understanding cancer metabolic plasticity and identifies MMP as a measurable parameter of tumor aggressiveness and therapeutic vulnerability.
Quantitative Profiling of MMP in Cancer Models
Comprehensive assessment of MMP across colorectal cancer cell lines reveals significant heterogeneity in mitochondrial properties that correlate with distinct metabolic phenotypes and drug sensitivities.
Table 1: Mitochondrial Properties and Bioenergetic Profiles in Colorectal Cancer Cell Lines
| Cell Line | Relative MMP (JC-1 Red/Green Ratio) | mtDNA Copy Number | % Cells with Depolarized Mitochondria | Primary Metabolic Phenotype | Dominant ETC Complex Expression |
|---|---|---|---|---|---|
| SW480 | 2.8 ± 0.3 | 285 ± 24 | 12.3 ± 2.1 | OXPHOS-dependent | Complex II, IV |
| SW1417 | 1.9 ± 0.2 | 167 ± 18 | 28.7 ± 3.4 | Glycolytic | Complex I, III |
| LS123 | 3.2 ± 0.4 | 412 ± 35 | 8.2 ± 1.6 | OXPHOS-dependent | Complex II, IV, V |
| SW403 | 2.1 ± 0.3 | 198 ± 22 | 22.5 ± 2.8 | Mixed phenotype | Complex I, II |
| SW620 | 2.5 ± 0.3 | 254 ± 28 | 16.8 ± 2.3 | OXPHOS-dependent | Complex III, IV |
| HCT-15 | 1.7 ± 0.2 | 142 ± 16 | 34.2 ± 3.9 | Glycolytic | Complex I |
Data adapted from comprehensive mitochondrial characterization studies across CRC models [105].
Table 2: Metabolic Parameters Following Neuronal Mitochondrial Transfer in Breast Cancer Models
| Parameter | 4T1 Monoculture | 4T1-Neuron Coculture | Change (%) | p-value |
|---|---|---|---|---|
| Basal Mitochondrial Respiration | 82.3 ± 6.1 pmol/min | 124.7 ± 8.9 pmol/min | +51.5% | <0.001 |
| Maximal Respiration Capacity | 156.2 ± 11.3 pmol/min | 245.8 ± 16.4 pmol/min | +57.4% | <0.001 |
| Spare Respiratory Capacity | 73.9 ± 7.2 pmol/min | 121.1 ± 9.8 pmol/min | +63.9% | <0.001 |
| ATP-linked Respiration | 58.4 ± 4.8 pmol/min | 89.2 ± 7.1 pmol/min | +52.7% | <0.001 |
| Metastatic Incidence | 42.3 ± 5.1% | 78.6 ± 6.9% | +85.8% | <0.001 |
Data showing enhanced bioenergetic parameters following mitochondrial transfer from neurons in breast cancer models [107].
Methodological Framework: Assessing MMP in Experimental Models
JC-1 Assay for MMP Quantification
Protocol: Cells are trypsinized and resuspended in complete media containing 3 μM JC-1 dye, followed by 30-minute incubation at 37°C with 5% CO₂. Cells are acquired using a flow cytometer with FITC (green, ~529 nm) and PE (red, ~590 nm) channels. MMP is calculated as the ratio of geometric mean fluorescence intensity (PE/FITC). Cells with low PE signal are classified as having depolarized mitochondria [105].
Technical Considerations: The JC-1 dye exhibits potential-sensitive accumulation in mitochondria, indicated by fluorescence emission shift from green (~529 nm) to red (~590 nm). The red/green ratio provides a quantitative measure of MMP that is independent of mitochondrial mass, with higher ratios indicating greater mitochondrial polarization.
MitoTRACER for Tracking Mitochondrial Transfer
Innovative Methodology: The MitoTRACER system employs genetic reporters that permanently label recipient cancer cells acquiring neuronal mitochondria, enabling fate mapping of mitochondrial recipient cells and their progeny. Donor neurons are genetically modified to express eGFP-labelled mitochondria, while recipient cancer cells express mCherry, allowing precise tracking of transferred organelles and their functional impact on metastatic dissemination [107].
Application: This methodology enables quantitative assessment of mitochondrial transfer frequency and the metabolic consequences for recipient cells, including enhanced spare respiratory capacity and increased metastatic potential in primary tumors.
Mitochondrial ROS Assessment
Protocol: Cells are resuspended in serum-free media containing 0.1% BSA and 5 μM MitoSOX Red dye, followed by 30-minute incubation at 37°C with 5% CO₂. Cells are acquired using flow cytometry, with fluorescence intensity proportional to mitochondrial superoxide production [105].
Molecular Mechanisms: MMP in Metabolic Reprogramming Pathways
Diagram 1: MMP Regulation in Cancer Metabolic Plasticity
MMP-Mediated Chemoresistance Mechanisms
Diagram 2: Chemoresistance Pathways Regulated by MMP
Research Reagent Solutions for MMP Investigation
Table 3: Essential Research Reagents for MMP and Metabolic Studies
| Reagent Category | Specific Reagents | Primary Function | Application Notes |
|---|---|---|---|
| MMP Detection | JC-1, TMRE, MitoTracker Red | Quantitative MMP measurement via flow cytometry | JC-1 provides ratio-metric measurement; optimal concentration 3-5 μM |
| Mitochondrial Transfer Tracking | MitoTRACER system, eGFP-mito labeled donors | Permanent labeling of mitochondrial recipient cells | Enables fate mapping of cells acquiring exogenous mitochondria [107] |
| Mitochondrial ROS Detection | MitoSOX Red, MitoTracker Green | Superoxide-specific detection in mitochondria | MitoSOX (5 μM) specifically detects superoxide, not other ROS |
| OXPHOS Inhibition | Rotenone (Complex I), Thenoyltrifluoroacetone (Complex II), Antimycin A (Complex III) | Targeted ETC complex inhibition | Used to identify metabolic dependencies and vulnerabilities [106] |
| Metabolic Modulators | 2-Deoxy-D-glucose, Oligomycin, Carbonyl cyanide-p-trifluoromethoxyphenylhydrazone | Glycolysis and OXPHOS manipulation | FCCP used as uncoupler for maximal respiration assessment |
| mtDNA Quantification | ATP8/B2M primers, tRNALeu/LDL primers | Absolute mtDNA copy number measurement | Dual primer sets ensure accuracy; 2-ΔCt calculation method [105] |
Therapeutic Targeting of MMP in Cancer
The strategic targeting of MMP-related vulnerabilities represents an emerging frontier in cancer therapeutics. Mitochondria-targeting agents (Mitocans) demonstrate significant potential by exploiting the differential MMP between normal and malignant cells [106]. Compounds that selectively disrupt MMP homeostasis in cancer cells include:
- ETC Complex Inhibitors: Metformin (Complex I), Atovaquone (Complex III), and VLX600 (Complex I/III inhibitor) preferentially target OXPHOS-dependent cancer subpopulations.
- ROS Modulators: Mito-TEMPO (mitochondrial superoxide scavenger) effectively inhibits tumor cell migration and spontaneous metastasis in murine and human models [106].
- Uncoupling Agents: Mild mitochondrial uncouplers such as Niclosamide ethanolamine selectively reduce MMP in cancer cells without complete bioenergetic collapse.
The efficacy of these mitochondrial-targeting strategies is highly dependent on the baseline bioenergetic profile of cancer cells, with OXPHOS-dependent subtypes showing heightened sensitivity to ETC inhibition, while glycolytic subtypes require combinatorial approaches targeting compensatory pathways [105].
Mitochondrial membrane potential represents a critical nexus in cancer metabolic reprogramming, serving as both a biomarker of metabolic phenotype and a functional regulator of therapeutic response. The quantitative assessment of MMP heterogeneity within tumors provides valuable insights into metabolic partitioning and potential resistance mechanisms. Furthermore, the discovery of intercellular mitochondrial transfer as a mechanism for enhancing MMP and metastatic capacity unveils novel therapeutic targets for disrupting the nerve-cancer interface. Future therapeutic strategies should incorporate MMP profiling to identify patient-specific metabolic vulnerabilities and optimize targeting approaches for improved clinical outcomes.
Mitochondrial dysfunction constitutes a central pillar in the pathogenesis of a spectrum of neurodegenerative disorders, with the mitochondrial membrane potential (MMP) serving as a critical integrator of cellular stress and a determinant of metabolic specialization. This whitepaper delineates the mechanistic links between Mitochondrial Membrane Protein-Associated Neurodegeneration (MPAN), Alzheimer's disease (AD), and Parkinson's disease (PD), focusing on how compromised MMP and subsequent metabolic remodeling drive neuronal vulnerability. Despite distinct genetic origins and clinical presentations, these diseases converge on pathways of impaired mitochondrial quality control, aberrant protein aggregation, and bioenergetic failure. The evidence synthesized herein underscores MMP not merely as a bioenergetic parameter but as a dynamic signaling hub whose dysregulation precedes and potentiates neurodegeneration. This consolidated framework aims to orient therapeutic development toward shared nodal points in mitochondrial pathophysiology, offering a paradigm for targeting early, convergent events in these debilitating conditions.
The mitochondrial membrane potential (MMP), generated by the electron transport chain (ETC), is the principal component of the protonmotive force (PMF) and is fundamentally required for ATP synthesis, calcium buffering, and metabolic biosynthesis [2]. In neurons, which exhibit high bioenergetic demand and complex morphology, the MMP is not static but is dynamically regulated to support synaptic plasticity, dendritic remodeling, and axonal transport [2]. A growing body of evidence positions MMP as a key signaling entity that integrates cellular status and directs mitochondrial functional specialization, including the partitioning of mitochondria into subpopulations dedicated to either energy production or biosynthetic precursor generation [2]. The loss of MMP integrity serves as a primary signal for mitochondrial quality control, initiating mitophagy to remove damaged organelles [2] [108].
The dysregulation of this sophisticated system is a hallmark of neurodegenerative disease. In Alzheimer's disease (AD), impaired cerebral glucose metabolism and mitochondrial dysfunction are invariant early features, often preceding clinical onset by decades [109] [110]. In Parkinson's disease (PD), the discovery that mitochondrial toxins like MPTP inhibit ETC complex I and induce parkinsonism provided foundational evidence for mitochondrial involvement [111]. Furthermore, genetic forms of PD are frequently linked to genes like PINK1 and Parkin, which are central to MMP-sensing and mitophagy [108] [111]. MPAN, a rare disorder caused by mutations in the orphan gene C19orf12, is characterized by iron accumulation in the brain and provides a unique genetic model of the interplay between mitochondrial dysfunction, iron metabolism, and neuronal death [112] [113]. This whitepaper examines the evidence linking MMP dysfunction and subsequent metabolic collapse across MPAN, AD, and PD, framing this nexus as a critical target for therapeutic intervention.
Clinical and Pathological Hallmarks: A Comparative Analysis
The following tables provide a comparative overview of the clinical, pathological, and genetic features of MPAN, Alzheimer's disease, and Parkinson's disease, highlighting shared themes of mitochondrial involvement.
Table 1: Clinical and Genetic Profiles of MPAN, AD, and PD
| Feature | MPAN | Alzheimer's Disease (AD) | Parkinson's Disease (PD) |
|---|---|---|---|
| Primary Genetic Cause(s) | Mutations in C19orf12 (autosomal recessive/dominant) [112] | Mutations in APP, PS1, PS2 (familial); APOE4 risk allele (sporadic) [109] | Mutations in LRRK2, GBA, PINK1, Parkin, SNCA (familial & sporadic) [111] |
| Age of Onset | Childhood or early adulthood [112] | Typically late adulthood (>65 years) [109] | Typically late adulthood (>60 years); earlier in genetic forms [111] |
| Core Clinical Symptoms | Spasticity, dystonia, motor axonal neuropathy, optic atrophy, cognitive decline, psychiatric symptoms [112] | Progressive memory loss, cognitive decline, apathy, disorientation [109] [110] | Bradykinesia, resting tremor, rigidity, postural instability; non-motor symptoms (e.g., sleep disorders, depression) [111] |
| Key Neuropathological Hallmarks | Iron accumulation in globus pallidus & substantia nigra; optic atrophy [112] | Amyloid-β plaques, neurofibrillary tangles (hyperphosphorylated tau), synaptic loss [109] [110] | Loss of dopaminergic neurons in substantia nigra pars compacta; Lewy bodies (α-synuclein aggregates) [111] |
| Primary Mitochondrial Dysfunctions | Altered cellular metabolism, redox homeostasis, and lipid metabolism; impaired metabolic flexibility [114] | Impaired glucose metabolism, oxidative phosphorylation (OXPHOS) deficits, increased oxidative stress, disrupted Ca²⁺ homeostasis [109] [110] | ETC Complex I deficiency, oxidative stress, disrupted mitochondrial quality control (e.g., PINK1/Parkin pathway) [111] [115] |
Table 2: Quantitative Biomarkers of Mitochondrial Dysfunction in Neurodegeneration
| Parameter | MPAN | Alzheimer's Disease (AD) | Parkinson's Disease (PD) |
|---|---|---|---|
| Brain Glucose Metabolism | Information not specific in search results | ↓ 25-30% reduction in affected brain regions (e.g., hippocampus, cortex) measured by FDG-PET [109] | Information not explicit in search results |
| ETC Complex I Activity | Information not explicit in search results | Information not explicit in search results | ↓ Significant decrease in substantia nigra homogenates [111] |
| MMP / ATP Levels | Information not explicit in search results | ↓ Depolarized MMP and ↓ reduced ATP production linked to Aβ pathology [116] [110] | ↓ Reduced MMP and ↓ ATP levels in MPP+ cell models [115] |
| TCA Cycle Metabolites | Information not explicit in search results | ↓ Reduced levels of thiamine-dependent enzymes (PDHC, KGDHC) in TCA cycle [109] | ↓ Imbalanced α-KG/Fumarate ratio due to bottlenecks (e.g., MDH2, OGDHL) [115] |
| Oxidative Stress Markers | ↑ Altered redox homeostasis in patient fibroblasts [114] | ↑ Increased ROS and oxidative damage [110] | ↑ Increased ROS from mitochondrial oxidant stress [111] |
Molecular Mechanisms and Experimental Evidence
MMP in Alzheimer's Disease: A Determinant of Amyloid Precursor Protein Trafficking and Aβ Secretion
The relationship between mitochondrial function and amyloid precursor protein (APP) processing in AD is bidirectional. On one hand, amyloid-β (Aβ) localizes to mitochondria, inhibits key enzymes like cytochrome c oxidase (COX), and induces oxidative stress, thereby disrupting MMP [116] [110]. On the other hand, the MMP itself directly regulates the subcellular localization of APP and the subsequent secretion of Aβ. A pivotal study demonstrated that mitochondrial depolarization (e.g., via the uncoupler FCCP) routes APP to mitochondria, resulting in decreased Aβ secretion. Conversely, mitochondrial hyperpolarization (e.g., via the ATP synthase inhibitor oligomycin) routes APP away from mitochondria and increases Aβ secretion [116]. This establishes an inverse correlation between mitochondrial APP localization and Aβ secretion, positioning MMP as a critical switch in amyloidogenic processing.
Experimental Protocol: Investigating MMP-Dependent APP Localization and Aβ Secretion [116]
- Cell Models: SH-SY5Y neuroblastoma cells transfected with wild-type APP695; human iPSC-derived neurons.
- MMP Modulation: Treat cells with FCCP (5-20 µM, uncoupler) to depolarize mitochondria, or oligomycin (5-20 µM, ATP synthase inhibitor) to hyperpolarize mitochondria for 4 hours. Include a glucose starvation condition using Hank's Balanced Salt Solution (HBSS) to induce metabolic stress.
- MMP Measurement: Load cells with 200 nM Tetramethylrhodamine, ethyl ester (TMRE) for 30 minutes. Measure fluorescence intensity (Ex/Em ~549/575 nm) via plate reader or imaging, normalized to cell count (e.g., using Hoechst stain).
- Mitochondrial Isolation: Post-treatment, homogenize cells and isolate mitochondria using differential centrifugation in an MSHE buffer (225 mM mannitol, 75 mM sucrose, etc.).
- Aβ Quantification: Concentrate proteins from cell media using ice-cold acetone. Resuspend pellets in 8M urea. Measure Aβ40 and Aβ42 levels using specific human ELISA kits, normalizing to total protein content (BCA assay).
- Key Interpretation: Cells with hyperpolarized mitochondria (oligomycin-treated) show less APP in mitochondrial fractions and higher secreted Aβ in media. Cells with depolarized mitochondria (FCCP-treated, ρ0 cells) show more APP in mitochondria and lower secreted Aβ.
Diagram Title: MMP Regulates APP Localization and Aβ Secretion
MMP and Mitophagy in Parkinson's Disease: The PINK1/Parkin Pathway
In PD, the PINK1/Parkin pathway is a quintessential example of an MMP-sensing mechanism that activates mitophagy. Under healthy, polarized MMP conditions, PINK1 is imported into mitochondria and cleaved. Upon mitochondrial damage and loss of MMP (depolarization), PINK1 import is halted, leading to its accumulation on the outer mitochondrial membrane (OMM). PINK1 then phosphorylates ubiquitin and recruits the E3 ubiquitin ligase Parkin. Activated Parkin ubiquitinates numerous OMM proteins, which are recognized by autophagy receptors (e.g., OPTN, NDP52) that recruit autophagosomal membranes (LC3) for degradation [108] [2]. Mutations in PINK1 or Parkin disrupt this quality control system, leading to the accumulation of damaged mitochondria, increased oxidative stress, and ultimately, neurodegeneration of vulnerable dopaminergic neurons [108] [111].
Experimental Protocol: Assessing Mitochondrial Quality Control in a PD Cell Model [115]
- PD Model Induction: Differentiate SH-SY5Y neuroblastoma cells into a neuronal phenotype. Treat cells with 1 mM MPP+ for 24-48 hours to model PD.
- Rescue Strategies: Co-treat with ZLN005 (mitochondrial biogenesis activator) or GSK-IN-3 (mitophagy inducer) to assess functional recovery.
- Viability & Apoptosis: Measure cell viability using MTT or related assays. Assess apoptosis via TUNEL staining or caspase-3 activity.
- MMP and ATP Measurement: Load cells with TMRE to measure MMP fluorometrically. Quantify intracellular ATP levels using a luciferase-based assay.
- Mitochondrial Content: Quantify mitochondrial mass via immunoblotting or immunofluorescence for the outer membrane protein TOMM20.
- MQC Protein Analysis: Analyze key mitochondrial quality control proteins by Western blot: Biogenesis (PGC1-α, NRF2, TFAM), Dynamics (MFN2, OPA1, DRP1), Mitophagy (PINK1, PARKIN, LC3B-II/I ratio).
- Key Interpretation: MPP+ treatment reduces MMP, ATP, and expression of MQC proteins. Successful rescue with ZLN005 or GSK-IN-3 restores these parameters and improves cell viability.
Diagram Title: PINK1/Parkin Mitophagy Pathway Activated by MMP Loss
Metabolic Remodeling in MPAN and PD: From Mitochondrial Dysfunction to Epigenetic Dysregulation
Recent multi-omics analyses reveal that mitochondrial dysfunction can drive neurodegeneration through metabolic remodeling of the TCA cycle, which in turn alters the nuclear epigenetic landscape. In PD, studies of patient striatal tissues and cell models identified MDH2, OGDHL, and IDH3G as bottleneck enzymes in the TCA cycle [115]. The resulting imbalance in the ratio of TCA metabolites, specifically a decreased α-ketoglutarate (α-KG) to fumarate ratio, inhibits the activity of histone demethylases (KDMs). This leads to increased H3K4me3 levels and enhanced expression of pro-degenerative genes like SNCA (alpha-synuclein), creating a "mitochondrial-nuclear" axis of pathology [115].
A analogous process of metabolic disturbance is observed in MPAN. Patient-derived fibroblasts show significant alterations in cellular metabolism, redox homeostasis, and lipidomic profiles. These deficits are potentiated when cells are forced to rely on oxidative phosphorylation, indicating a critical loss of metabolic flexibility—the ability to switch between energy-producing pathways in response to nutrient and energy demands [114]. This failure to maintain bioenergetic homeostasis, rooted in mitochondrial dysfunction, is a shared feature across these disorders.
Experimental Protocol: Multi-Omics Analysis of TCA Cycle in PD [115]
- Transcriptomic Data Collection: Obtain bulk RNA-seq datasets from PD patient brain regions (e.g., caudate (CAU), putamen (PUT)) and single-cell RNA-seq (scRNA-seq) data from substantia nigra from public repositories.
- Differential Expression Analysis: Identify differentially expressed genes (DEGs) in PD vs. normal controls. Cross-reference with mitochondrial gene databases (e.g., MitoCarta3.0).
- Pathway & GSEA: Perform Gene Ontology (GO), KEGG pathway, and Gene Set Enrichment Analysis (GSEA) on DEGs to identify enriched pathways (e.g., TCA cycle, OXPHOS, apoptosis).
- In Vitro Validation: Establish an MPP+-induced PD model in SH-SY5Y cells. Validate key findings (e.g., reduced MDH2, OGDHL expression) via qPCR and Western blot.
- Metabolite & Epigenetic Measurement: Measure intracellular α-KG and fumarate levels using LC-MS. Assess global H3K4me3 levels by Western blot and specific SNCA promoter enrichment by ChIP-qPCR.
- Key Interpretation: Mitochondrial dysfunction in PD causes a TCA cycle bottleneck, leading to an altered α-KG/fumarate ratio that inhibits H3K4me3 demethylation, thereby increasing SNCA expression.
The Scientist's Toolkit: Key Research Reagents and Models
Table 3: Essential Research Reagents and Models for Investigating Mitochondrial Dysfunction
| Reagent / Model | Function / Application | Specific Example Use-Case |
|---|---|---|
| TMRE (Tetramethylrhodamine, Ethyl Ester) | Potentiometric dye for quantifying MMP. Accumulates in active mitochondria based on membrane potential. | Measuring FCCP/oligomycin-induced MMP changes in SH-SY5Y cells [116]. |
| MPP+ (1-methyl-4-phenylpyridinium) | Neurotoxic metabolite of MPTP; inhibits mitochondrial ETC Complex I. | Modeling PD pathogenesis in SH-SY5Y and iPSC-derived neuronal cultures [115]. |
| FCCP (Carbonyl cyanide-p-trifluoromethoxyphenylhydrazone) | Mitochondrial uncoupler; dissipates the proton gradient, causing rapid MMP loss. | Experimentally inducing mitochondrial depolarization to study APP trafficking [116]. |
| Oligomycin | ATP synthase (Complex V) inhibitor; causes mitochondrial hyperpolarization. | Experimentally inducing hyperpolarization to study its effect on Aβ secretion [116]. |
| ZLN005 | Small-molecule activator of mitochondrial biogenesis (activates PGC-1α). | Rescuing mitochondrial biogenesis defects in a PD cell model [115]. |
| GSK-IN-3 | Inducer of PINK1-Parkin mediated mitophagy. | Enhancing clearance of damaged mitochondria in models of impaired mitophagy [115]. |
| Patient-Derived Fibroblasts | Primary cells from patients (e.g., with MPAN) to study disease-specific cellular phenotypes. | Investigating metabolic flexibility, redox homeostasis, and lipid profiles under OXPHOS-promoting conditions [114]. |
| iPSC-Derived Neurons | Human neurons differentiated from induced pluripotent stem cells, enabling study of human-specific disease mechanisms in relevant cell types. | Validating findings from cell lines (e.g., MMP-APP relationship) in a human neuronal context [116]. |
Integrated Pathogenic Model and Therapeutic Outlook
The evidence converges on a model where genetic or environmental insults trigger an initial decline in mitochondrial efficiency, often manifesting as reduced MMP or ETC dysfunction. This failure compromises the organelle's ability to function as a metabolic hub, leading to:
- Bioenergetic Crisis: Reduced ATP production impairs essential neuronal functions like synaptic transmission and ionic homeostasis [109] [111].
- Oxidative Stress: Electron leak from the impaired ETC increases reactive oxygen species (ROS), damaging lipids, proteins, and DNA [111] [110].
- Failed Quality Control: MMP dissipation activates mitophagy, but genetic defects (e.g., in PINK1, Parkin, or WDR45) or general lysosomal dysfunction prevent efficient clearance, leading to the accumulation of toxic organelles [108] [111] [113].
- Metabolic Remodeling: TCA cycle dysfunction alters metabolite pools (e.g., α-KG, fumarate), which can disrupt epigenetic regulation and drive the expression of pathogenic proteins like α-synuclein [115].
- Aberrant Protein Aggregation: The compromised cellular environment promotes the misfolding and accumulation of proteins like Aβ, tau, and α-synuclein, which in turn further damage mitochondria, creating a vicious cycle of degeneration [109] [116] [111].
This integrated view highlights several promising therapeutic avenues. Strategies include: enhancing mitochondrial biogenesis (e.g., with ZLN005), promoting mitophagy (e.g., with GSK-IN-3), correcting TCA cycle metabolism (e.g., with citrate supplementation [115]), and using antioxidants to mitigate oxidative stress. For MPAN and related NBIA disorders, the intricate link between mitochondrial dysfunction and iron accumulation suggests that therapies targeting both pathways may be necessary [113]. The consistent failure of trials targeting single downstream features like Aβ plaques underscores the need for early intervention at the level of mitochondrial integrity and metabolic flexibility. Restoring the MMP and its associated signaling functions represents a unifying strategy to break the cycle of degeneration across these distinct but mechanistically linked diseases.
Cardiovascular diseases (CVDs) remain the leading cause of morbidity and mortality worldwide, surpassing traditional risk factors through complex pathophysiological mechanisms [117]. Emerging evidence positions mitochondrial dysfunction as a central driver of CVD progression, linking impaired bioenergetics, oxidative stress imbalance, and defective mitochondrial quality control to endothelial dysfunction, myocardial injury, and adverse cardiac remodeling [117]. The heart, as a high-energy-demand organ, exhibits the highest mitochondrial density among all cell types, with mitochondria occupying approximately 40% of cardiomyocyte volume [118]. These organelles are crucial for cardiac metabolism, producing over 90% of myocardial adenosine triphosphate (ATP) through oxidative phosphorylation (OXPHOS) while simultaneously regulating reactive oxygen species (ROS) production, calcium homeostasis, and apoptotic signaling [117].
Within the context of metabolic specialization and partitioning research, mitochondrial membrane potential (ΔΨm) serves as a critical indicator of mitochondrial health and functional specialization. Maintaining ΔΨm is essential for ATP synthesis, calcium buffering, and ROS regulation in cardiac tissue [119]. However, in pathological conditions such as obesity and aging, the heart undergoes significant mitochondrial dynamic remodeling characterized by imbalances in fission and fusion processes, oxidative stress, and metabolic shifts that ultimately compromise cardiac function [120] [121]. This whitepaper provides a comprehensive analysis of the molecular mechanisms through which obesity and aging disrupt mitochondrial dynamics in the heart, with particular emphasis on how these changes affect mitochondrial membrane potential and contribute to cardiac pathology. Furthermore, we evaluate emerging therapeutic strategies and experimental methodologies relevant to drug development professionals seeking to target mitochondrial dynamics for cardioprotection.
Molecular Mechanisms of Mitochondrial Dynamic Remodeling
Core Regulatory Machinery of Mitochondrial Dynamics
Mitochondrial dynamics, comprising fission and fusion processes, are governed by conserved GTPase proteins that continuously remodel the mitochondrial network. These dynamic processes are essential for maintaining mitochondrial quality, shape, and function in cardiomyocytes [118].
Mitochondrial fission is primarily mediated by Dynamin-related protein 1 (Drp1), a cytoplasmic protein that translocates to the mitochondrial outer membrane (OMM) where it oligomerizes into GTP-hydrolyzing helical polymers that drive membrane constriction and division [118]. Drp1 recruitment is facilitated by OMM receptors including Fission 1 (Fis1) and Mitochondrial Fission Factor (MFF) [118]. Fission enables the segregation of damaged mitochondrial components for degradation via mitophagy and facilitates the distribution of mitochondria during cell division [121].
Mitochondrial fusion occurs through coordinated actions of mitofusins 1 and 2 (Mfn1, Mfn2) at the OMM and Optic Atrophy 1 (OPA1) at the inner mitochondrial membrane (IMM). Mfn1 and Mfn2 form antiparallel dimers through interactions involving their Heptad Repeat Domain 1 (HR1), with GTP hydrolysis inducing conformational changes that enable OMM fusion pore formation [118]. OPA1 exists in long (L-OPA1) and short (S-OPA1) forms generated through proteolytic cleavage by OMA1 (at S1 site) and YME1L (at S2 site). Both OPA1 forms assemble into GTP-dependent complexes that facilitate IMM fusion and cristae reorganization [118]. Fusion promotes content mixing between mitochondria, complementing minor defects in mitochondrial proteins or DNA, and maintains oxidative phosphorylation efficiency [121].
The following diagram illustrates the core molecular machinery governing mitochondrial fission and fusion processes:
Mitochondrial Dysfunction in Obesity-Induced Cardiomyopathy
Obesity imposes severe stress on cardiac mitochondria through multiple mechanisms, ultimately leading to dysfunctional mitochondrial dynamic remodeling. The accumulation of adipose tissue and fatty acids contributes to the development and progression of heart pathogenic conditions [121]. Specifically, intracellular lipid accumulation leads to elevated levels of ceramides and diacylglycerol (DAG), which directly impair mitochondrial function and dynamics regulation [121]. Ceramides promote inflammation and detrimentally affect mitochondrial integrity, resulting in cardiomyocyte apoptosis and autophagy, while DAG accumulation activates protein kinase C (PKC) isoforms that induce insulin resistance and further disrupt mitochondrial function [121].
A key mechanism linking obesity to mitochondrial dysfunction involves the small GTPase RalA, which shows increased expression and activity in white adipocytes from obese mice [122]. RalA activation promotes mitochondrial fragmentation by reversing the inhibitory Ser637 phosphorylation of Drp1 through interaction with protein phosphatase PP2Aa, leading to excessive fission [122]. This process creates a vicious cycle wherein mitochondrial fragmentation reduces oxidative capacity and energy expenditure, further promoting weight gain and metabolic dysfunction.
The table below summarizes the quantitative changes in mitochondrial dynamics proteins observed in various models of obesity:
Table 1: Alterations in Mitochondrial Dynamics Proteins in Experimental Models of Obesity
| Experimental Model | Diet/Duration | Drp1 | Fis1 | Mfn1 | Mfn2 | OPA1 | Cardiac Functional Outcome | Citation |
|---|---|---|---|---|---|---|---|---|
| Rat | HFD (17-59.3% E fat), 6-24 weeks | ↑ | - | ↓ | ↓ | ↓ | Mitochondrial dysfunction | [121] |
| Mouse | HFD (45-60% E fat), 20-24 weeks | ↑ | - | ↓ | ↓ | ↓ | Metabolic & LV dysfunction | [121] |
| Pig | HFD (47.7% E fat), 6 months | ↑ | ↑ | - | - | ↓ | Cardiometabolic dysfunction | [121] |
| db/db mice (genetic obesity) | - | ↑ | - | ↓ | ↓ | ↓ | Impaired LV function | [121] |
| Mouse iWAT adipocytes | HFD (60% E fat), 16 weeks | ↑ (Drp1 Ser637 dephosphorylation) | - | - | - | - | Mitochondrial fragmentation, reduced oxidative capacity | [122] |
HFD: High-fat diet; E fat: Energy from fat; iWAT: inguinal white adipose tissue; LV: Left ventricular; ↑: Increased expression/activity; ↓: Decreased expression; -: Not reported
Obesity-induced mitochondrial fragmentation is particularly detrimental in cardiomyocytes, where the balance between fission and fusion is crucial for maintaining bioenergetic efficiency. The shift toward excessive fission leads to mitochondrial dysfunction characterized by decreased OXPHOS efficiency, reduced ATP production, increased ROS generation, and loss of mitochondrial membrane potential (ΔΨm) [121]. These changes impair cardiac contractility and promote maladaptive remodeling, ultimately contributing to the development of obese cardiomyopathy.
Age-Associated Mitochondrial Remodeling in the Heart
Cardiac aging is characterized by progressive structural and functional decline of the myocardium, with mitochondrial dysfunction serving as both a driver and amplifier of this process [120]. In the aging heart, mitochondria undergo profound morphological alterations including cristae disorganization, swelling, fragmentation, and vacuolization [120]. These structural changes are accompanied by functional deficits, particularly reduced efficiency of complexes I and IV of the electron transport chain, leading to impaired OXPHOS capacity and bioenergetic insufficiency [120].
The aging process disrupts the delicate balance of mitochondrial dynamics through multiple interconnected mechanisms:
Impaired fusion processes: Aging hearts exhibit decreased expression of fusion mediators including Mitofusin 2 (Mfn2) and Optic Atrophy 1 (Opa1), as well as alterations in cardiolipin, a lipid critical for maintaining inner mitochondrial membrane integrity [120]. These changes hinder mitochondrial elongation and cristae remodeling, thereby compromising OXPHOS and reducing ATP production.
Excessive fission activation: Mitochondrial fission becomes disproportionately activated through enhanced Drp1 translocation and upregulation of Fission 1 (Fis1), resulting in a fragmented mitochondrial network prone to depolarization and ROS overproduction [120].
Compromised quality control: Mitophagy, particularly PINK1/Parkin-mediated pathways, becomes dysfunctional with age. While aged cardiomyocytes may transiently upregulate PINK1 and Parkin under stress, overall mitophagic flux is blunted, leading to inefficient clearance of damaged mitochondria [120].
Reduced mitochondrial biogenesis: Aging is associated with downregulation of transcriptional coactivators such as PGC-1α and NRF1, resulting in a net decline in mitochondrial number and respiratory capacity [120].
Accumulation of mtDNA damage: Mitochondrial DNA is particularly vulnerable to age-associated oxidative damage due to its proximity to ROS-generating sites and lack of protective histones [120]. mtDNA mutations accumulate over time, further compromising oxidative phosphorylation and creating a vicious cycle of ROS production and mitochondrial dysfunction.
The table below summarizes key changes in mitochondrial dynamics and function observed in aging heart models:
Table 2: Mitochondrial Alterations in Experimental Models of Cardiac Aging
| Aging Model | Age/Species | Drp1 | Fis1 | Mfn1 | Mfn2 | OPA1 | Functional Consequences | Citation |
|---|---|---|---|---|---|---|---|---|
| Natural aging | 24-month mice | → | → | ↓ | ↓ | ↓ (L-OPA1/S-OPA1 ratio) | Mitochondrial fragmentation | [121] |
| Natural aging | 20-24 month rats | ↑ | ↑ | ↓ | - | ↓ | Mitochondrial dysfunction, cardiac impairment | [121] |
| Natural aging | 25-month rats | - | - | - | ↓ | - | Cardiac dysfunction | [121] |
| Natural aging | 22.4-year monkeys | - | - | ↓ (Mfn1/2 deficiency) | ↓ (Mfn1/2 deficiency) | - | Age-related cardiac dysfunction | [121] |
| D-galactose-induced aging | 150 mg/kg/day rats | ↑ | - | ↓ | ↓ | ↓ | Cardiac senescence, metabolic impairment | [121] |
| Natural aging | Aged mice & rats | - | - | - | - | - | Cristae disruption, reduced ETC activity, ↓ OXPHOS efficiency | [120] |
↑: Increased expression/activity; ↓: Decreased expression; →: No change; -: Not reported; ETC: Electron transport chain; OXPHOS: Oxidative phosphorylation
The following diagram illustrates the interconnected pathways through which aging disrupts mitochondrial homeostasis in cardiomyocytes:
Interplay Between Obesity and Aging in Cardiac Mitochondrial Pathology
While obesity and aging independently promote cardiac mitochondrial dysfunction, their combination produces synergistic detrimental effects. A study using rats demonstrated that the combination of high-fat-diet-induced obesity and D-galactose-induced aging resulted in greater vulnerability to myocardial injury through aggravated cardiac mitochondrial dysfunction and dynamic remodeling compared to either condition alone [121]. This interaction creates a pathological feedback loop wherein obesity accelerates age-related mitochondrial decline, and aging potentiates obesity-induced metabolic disturbances.
The convergence of obesity and aging on mitochondrial dysfunction involves several shared pathways:
ROS amplification: Both conditions increase mitochondrial ROS production, which damages proteins, lipids, and mtDNA, further compromising ETC function and increasing ROS generation in a self-perpetuating cycle.
Inflammasome activation: mtDNA released from damaged mitochondria acts as a damage-associated molecular pattern (DAMP), activating inflammatory pathways including cGAS-STING, NLRP3 inflammasome, and Toll-like receptor 9 signaling [117] [123]. This chronic low-grade inflammation contributes to insulin resistance, endothelial dysfunction, and cardiomyocyte injury.
Metabolic remodeling: The heart undergoes a substrate shift from fatty acid oxidation to glucose utilization in both aging and obesity. While initially compensatory, this shift reduces ATP efficiency over time and exacerbates cardiac dysfunction [117].
Calcium handling disruption: Mitochondrial calcium regulation is impaired in both conditions, affecting excitation-contraction coupling and promoting arrhythmogenesis.
Hormonal changes: Aging and obesity both affect hormonal signaling, including reduced NAD+ levels that impair sirtuin activity, further disrupting mitochondrial homeostasis [120].
Assessment Methodologies for Mitochondrial Function
Core Techniques for Evaluating Mitochondrial Parameters
Accurate assessment of mitochondrial function is essential for understanding their role in cardiac pathology and evaluating therapeutic interventions. The following methodologies represent core approaches for investigating key mitochondrial parameters:
Table 3: Essential Methodologies for Assessing Mitochondrial Function
| Parameter | Probe/Method | Principle | Application in Cardiac Studies | Technical Considerations |
|---|---|---|---|---|
| Mitochondrial Membrane Potential (ΔΨm) | Tetramethylrhodamine methyl ester (TMRM) | Potentiometric dye accumulates in mitochondria proportionally to ΔΨm | Critical for assessing mitochondrial health in cardiomyocytes; ΔΨm loss indicates pathology | Requires careful calibration of loading concentrations; confocal imaging preferred for quantitative analysis [119] |
| Reactive Oxygen Species (ROS) | MitoSOX Red | Mitochondria-targeted fluorogenic dye specifically detects superoxide | Quantifies mitochondrial oxidative stress in cardiac tissue sections and isolated cardiomyocytes | More specific than general ROS dyes; requires appropriate controls for non-mitochondrial signals [119] |
| Mitochondrial Calcium | Rhod-2 AM | Rationetric dye with affinity for mitochondrial calcium | Assesses mitochondrial calcium buffering capacity in contracting cardiomyocytes | AM ester form facilitates cellular loading; compartment-specific calibration essential [119] |
| Morphological Dynamics | Electron Microscopy | High-resolution imaging of mitochondrial ultrastructure | Reveals cristae organization, swelling, fragmentation in aged and obese hearts | Quantitative analysis requires specialized software and standardized sampling [120] |
| Oxygen Consumption | High-Resolution Respirometry | Measures oxygen consumption rates in isolated mitochondria or cells | Evaluates ETC function, coupling efficiency in cardiac mitochondrial subpopulations | Requires careful mitochondrial isolation preserving functional integrity [117] |
The Scientist's Toolkit: Research Reagent Solutions
The following table provides essential research reagents and tools for investigating mitochondrial dynamics in cardiac pathologies:
Table 4: Research Reagent Solutions for Cardiac Mitochondrial Studies
| Reagent/Category | Specific Examples | Function/Application | Experimental Context |
|---|---|---|---|
| Fluorescent Probes | TMRM, MitoSOX Red, Rhod-2 AM | Assessment of ΔΨm, mitochondrial ROS, and calcium levels | Live-cell imaging in cardiomyocytes, fluorescence microscopy [119] |
| Fission Inhibitors | Mdivi-1, P110 | Inhibit Drp1-mediated mitochondrial fission | Testing therapeutic strategies in obesity and aging models [121] |
| Fusion Promoters | - | Enhance mitochondrial fusion processes | Experimental cardioprotection approaches [121] |
| Genetic Models | Adipocyte-specific Rala KO, Mfn1/2 KO, Drp1 mutants | Investigate specific protein functions in mitochondrial dynamics | In vivo studies of obesity mechanisms, cell-autonomous effects [122] |
| Metabolic Assays | Seahorse XF Analyzer reagents | Measure OXPHOS, glycolysis, ATP production rates | Functional assessment in isolated cardiac mitochondria [117] |
| Antibodies | Anti-Drp1, anti-Mfn1/2, anti-OPA1, phospho-specific Drp1 | Protein expression and post-translational modification analysis | Western blot, immunohistochemistry of cardiac tissue [121] [118] |
| mtDNA Assessment | mtDNA copy number assays, mutation detection | Quantify mtDNA integrity and abundance | Correlation with cardiac function in aging and obesity [117] [120] |
Therapeutic Strategies and Experimental Approaches
Targeting Mitochondrial Dynamics for Cardioprotection
Emerging therapeutic strategies aim to restore mitochondrial homeostasis by targeting the imbalance between fission and fusion processes in obesity and aging. These approaches include:
Pharmacological interventions:
- Fission inhibitors: Compounds such as Mdivi-1 that inhibit Drp1 GTPase activity have demonstrated cardioprotective effects in preclinical models of obesity and aging by reducing excessive mitochondrial fragmentation [121].
- Fusion promoters: Strategies to enhance mitochondrial fusion show promise in counteracting age-related and obesity-related mitochondrial fragmentation, though development of specific pharmacological activators of Mfn1/2 and OPA1 remains challenging [121].
- Indirect modulators: Sodium-glucose cotransporter-2 (SGLT2) inhibitors like empagliflozin provide cardioprotection through indirect mitochondrial modulation, though direct mitochondria-targeting agents are limited by poor specificity [117].
Metabolic modulators:
- AMPK activators: Compounds that activate AMPK signaling can improve mitochondrial function and dynamics. For example, salidroside mitigates myocardial ischemia-reperfusion injury by activating Nrf2 and modulating the AMPK/PGC-1α/PPARα pathway [117].
- SIRT activators: Strategies to boost NAD+ levels or directly activate sirtuins can improve mitochondrial function in aging hearts by enhancing deacetylation of mitochondrial proteins [120].
Gene-based approaches:
- Mitochondrial gene-editing tools: While effective in vitro, their in vivo application is hindered by the dual barriers of cellular uptake and mitochondrial membrane penetration [117].
- AAV-mediated gene delivery: Shows promise for delivering mitochondrial protective factors to the heart, though achieving tissue specificity remains challenging.
The following diagram illustrates therapeutic strategies targeting mitochondrial dynamics:
Experimental Design Considerations for Drug Development
For researchers and drug development professionals investigating mitochondria-targeted therapies for cardiac pathologies, several experimental design considerations are critical:
Model selection: Choose animal models that appropriately recapitulate human disease pathophysiology. For obesity studies, high-fat diet feeding in rodents (typically 45-60% energy from fat for 20-24 weeks) effectively induces mitochondrial dynamic remodeling [121]. For aging research, natural aging models (20-24 month rodents) or D-galactose-induced senescence models provide complementary approaches.
Mitochondrial subpopulation analysis: Cardiac mitochondria exist as distinct subpopulations - interfibrillar (IFM), subsarcolemmal (SSM), and perinuclear (PNM) - with specialized functions [118]. Assessment of therapeutic interventions should evaluate effects on all subpopulations, as they may demonstrate differential susceptibility to pathology and treatment.
Temporal considerations: The timing of intervention is crucial, as mitochondrial dysfunction progresses through different phases in obesity and aging. Preventive versus restorative treatment strategies may target different molecular pathways.
Biomarker development: Identify and validate biomarkers for monitoring mitochondrial function in clinical trials, such as mtDNA copy number, mitochondrial membrane potential sensors, and circulating mtDNA fragments as indicators of mitochondrial damage [117] [123].
Tissue-specific delivery: Develop strategies to overcome the challenge of therapeutic specificity, including tissue-targeted delivery systems for mitochondrial agents to minimize off-target effects [117].
Mitochondrial dynamic remodeling represents a central pathological mechanism in both obesity-induced cardiomyopathy and cardiac aging. The imbalance between fission and fusion processes, characterized by excessive Drp1-mediated fission and impaired Mfn1/2 and OPA1-mediated fusion, leads to mitochondrial fragmentation, loss of membrane potential, bioenergetic deficiency, and ultimately cardiac dysfunction. The convergence of obesity and aging on mitochondrial pathways creates synergistic detrimental effects that accelerate cardiac decline.
Within the context of metabolic specialization and partitioning research, mitochondrial membrane potential serves as a critical integrator of mitochondrial health and functional state. Maintaining or restoring ΔΨm through targeted interventions that rebalance mitochondrial dynamics represents a promising therapeutic strategy for addressing the growing burden of obesity- and age-related cardiovascular diseases. Future research should focus on developing tissue-specific delivery systems for mitochondrial therapeutics, identifying biomarkers for patient stratification, and conducting well-designed clinical trials to translate promising preclinical findings into effective clinical therapies.
Sepsis, a life-threatening organ dysfunction caused by a dysregulated host response to infection, remains a leading cause of mortality worldwide despite advances in supportive care. The complexity of its pathogenesis and the limitations of current diagnostic markers have fueled the search for more specific biomarkers. Emerging research highlights the central role of mitochondrial dysfunction in sepsis pathophysiology, positioning mitochondria-derived damage-associated molecular patterns (mtDAMPs) and specific mitochondrial genes as promising biomarkers. This whitepaper examines the evidence supporting NDUFB3 and various mtDAMPs as clinical indicators, framing their utility within the broader context of mitochondrial membrane potential and metabolic specialization in sepsis. We synthesize current understanding of their roles in immune activation, organ dysfunction, and clinical outcomes, providing structured data, experimental protocols, and pathway visualizations to guide research and therapeutic development.
Sepsis is characterized by a dysregulated host response to infection that leads to life-threatening organ dysfunction [124]. With millions of cases annually and mortality rates remaining persistently high, sepsis represents a critical global health challenge [125]. The pathophysiology involves complex interactions between inflammatory pathways, immune responses, and metabolic alterations, with mitochondrial dysfunction emerging as a central mechanism driving organ failure [126] [125].
Mitochondria serve dual roles during sepsis: as crucial regulators of cellular energy metabolism and as key modulators of inflammatory responses [125] [127]. During sepsis, damage to the mitochondrial electron transport chain (ETC) in various organs impairs ATP production and oxygen utilization [127]. This dysfunction increases mitochondrial membrane permeability, releasing mitochondrial damage-associated molecular patterns (mtDAMPs) that amplify systemic inflammation and organ injury [128] [129] [127]. The assessment of mitochondrial membrane potential (ΔΨ), which reflects the functional metabolic status of mitochondria, provides critical insights into these processes [130].
Table 1: Key Mitochondrial Components in Sepsis Pathophysiology
| Mitochondrial Component | Role in Sepsis | Clinical/Research Significance |
|---|---|---|
| Electron Transport Chain (ETC) | Impaired complexes I, III, and IV reduce ATP production | Correlates with disease severity and organ dysfunction [125] |
| Mitochondrial Membrane Potential (ΔΨ) | Reflects proton electrochemical gradient and energy status | Measured via TMRM, TPP+; indicates metabolic state [130] |
| mtDAMPs | Molecular patterns released upon mitochondrial damage | Amplify inflammation; potential diagnostic biomarkers [128] [129] |
| Mitochondrial Quality Control | Mitophagy, biogenesis, and dynamics | Therapeutic target; timing crucial for intervention [126] [125] |
NDUFB3 as a Sepsis Biomarker
Biological Function and Expression Patterns
NDUFB3 (NADH:ubiquinone oxidoreductase subunit B3) is a core structural component of complex I (NADH dehydrogenase) in the mitochondrial electron transport chain. This complex plays a crucial role in initiating oxidative phosphorylation by catalyzing electron transfer from NADH to ubiquinone [131].
Research integrating bioinformatics analyses with experimental validation has identified NDUFB3 as a significant biomarker in sepsis. Analyses of multiple gene expression datasets (GSE54514, GSE65682, and GSE95233) revealed that NDUFB3 is highly expressed in sepsis patients compared to healthy controls [131]. Furthermore, studies examining both septic shock and stroke have identified NDUFB3 as one of eight core genes highly expressed in these conditions, with elevated expression correlating with poorer prognosis [132].
Diagnostic and Prognostic Value
The diagnostic performance of NDUFB3 for sepsis demonstrates considerable promise. In validation studies, NDUFB3 exhibited significant discriminatory power, contributing to a nomogram model with the formula: -2.548 × BCKDHB + -5.454 × LETMD1 + 4.507 × NDUFB3 [131]. This model effectively stratified sepsis risk, highlighting the value of NDUFB3 as a component of multi-marker panels.
Table 2: NDUFB3 as a Sepsis Biomarker: Key Findings
| Aspect | Findings | Implications |
|---|---|---|
| Expression Pattern | Significantly elevated in sepsis patients versus healthy controls [132] [131] | Potential for discriminating sepsis from non-septic inflammation |
| Prognostic Value | Higher expression associated with worse prognosis in septic shock and stroke [132] | May aid in risk stratification and treatment prioritization |
| Functional Role | Complex I component; knockdown reduces mitochondrial function in sepsis models [131] | Connects biomarker utility to pathological mechanism |
| Multi-Marker Utility | Component of diagnostic nomogram with BCKDHB and LETMD1 [131] | Enhanced diagnostic performance in combination with other markers |
Experimental Validation and Functional Assessment
The functional significance of NDUFB3 in sepsis has been validated through in vitro experiments using small interfering RNA (siRNA) technology. When NDUFB3 expression was inhibited in H9C2 cells (a cardiomyocyte cell line) under lipopolysaccharide (LPS)-simulated sepsis conditions, researchers observed significant reduction in mitochondrial function [131]. This experimental approach confirmed that NDUFB3 contributes substantially to mitochondrial quality imbalance in sepsis models, strengthening its relevance as both a biomarker and potential therapeutic target.
mtDAMPs as Circulating Inflammatory Mediators
Origins and Composition of mtDAMPs
Mitochondria are evolutionarily derived from bacterial endosymbionts, accounting for the structural similarities between mitochondrial components and bacterial pathogen-associated molecular patterns (PAMPs) [128] [129]. When mitochondrial damage occurs during sepsis, these components are released into the circulation and function as damage-associated molecular patterns (DAMPs), activating pattern recognition receptors (PRRs) and amplifying inflammatory responses [128] [129] [127].
The major mtDAMPs include:
- Mitochondrial DNA (mtDNA): Contains unmethylated CpG motifs similar to bacterial DNA [129]
- Mitochondrial formyl peptides (mtFPs): Share N-formyl methionine characteristics with bacterial peptides [129]
- ATP: Released from damaged mitochondria; functions as a DAMP at high concentrations [131]
- Cytochrome c: Component of electron transport chain; released upon membrane permeabilization [127]
- Cardiolipin: Phospholipid unique to mitochondrial inner membrane [127]
Clinical Correlations and Predictive Value
Multiple clinical studies have demonstrated the prognostic significance of circulating mtDAMPs in sepsis patients:
mtDNA levels show particularly strong clinical correlations. Plasma mtDNA concentrations are significantly elevated in sepsis patients compared to healthy controls, with one study reporting 50-fold higher levels in septic individuals [128]. Moreover, mtDNA levels correlate with disease severity, as non-survivors exhibit significantly higher plasma mtDNA than survivors [128]. The predictive value of mtDNA may even surpass conventional markers, with some evidence suggesting it could be superior to lactate levels or SOFA scores in predicting mortality after admission [131].
Other mtDAMPs also show clinical utility. ATP released from damaged mitochondria enhances macrophage bacterial killing through P2X7 and P2X4 receptors, contributing to host defense [131]. Urinary mtDNA levels have emerged as promising biomarkers for acute kidney injury in sepsis, negatively correlating with glomerular filtration rate and potentially guiding renal replacement therapy initiation [128].
Table 3: Major mtDAMPs and Their Signaling Pathways in Sepsis
| mtDAMP | Receptors/Pathways | Biological Effects | Clinical Utility |
|---|---|---|---|
| mtDNA | TLR9, cGAS-STING, NLRP3 inflammasome | Activates NF-κB, MAPK, IRF3; induces IFN-I, proinflammatory cytokines [128] [129] | Predicts mortality, organ dysfunction; 50x higher in sepsis [128] [131] |
| mtDNA (oxidized) | NLRP3 inflammasome | Preferentially activates NLRP3; promotes IL-1β, IL-18 maturation [129] | Potential severity marker; links oxidative stress to inflammation |
| ATP | P2X7, P2X4 receptors | Enhances macrophage bactericidal activity; proinflammatory at high levels [131] | Balance between host defense and inflammation amplification |
| Mitochondrial formyl peptides | Formyl peptide receptors | Neutrophil chemotaxis, inflammation amplification [129] | Contributes to neutrophil-mediated tissue damage |
Signaling Pathways Activated by mtDAMPs
TLR9 Signaling Pathway
The Toll-like receptor 9 (TLR9) pathway represents a major mechanism for mtDNA-induced inflammation. TLR9 directly binds to unmethylated CpG motifs in mtDNA, triggering intracellular signaling cascades [129]. Upon activation, TLR9 recruits the adaptor protein MyD88, leading to:
- NF-κB activation: Induces expression of proinflammatory cytokines (TNF-α, IL-1β, IL-6)
- MAPK cascade activation: Promotes AP-1 formation and cytokine expression
- IRF7 or IRF1 activation: Stimulates type I interferon production in different cell types [129]
In neutrophils, mtDNA activation through TLR9 causes Ca2+ influx and MAPK phosphorylation, mediating degranulation and migration [129]. While traditionally viewed as purely detrimental in sepsis, recent evidence suggests TLR9 may also have protective functions, such as facilitating mtDNA clearance by red blood cells to alleviate pulmonary inflammation [128].
cGAS-STING Signaling Pathway
The cyclic GMP-AMP synthase (cGAS)-stimulator of interferon genes (STING) pathway has emerged as a crucial mechanism for mtDNA recognition [128] [129]. When cytosolic mtDNA binds to cGAS, it catalyzes synthesis of 2'-3' cyclic GMP-AMP (cGAMP) from GTP and ATP. This secondary messenger then binds to STING in the endoplasmic reticulum membrane, triggering its conformational change and dimerization [129].
The activated STING complex subsequently:
- Recruits and phosphorylates TBK1, which phosphorylates IRF3, leading to type I interferon expression
- Directly binds and phosphorylates IKK complex, activating NF-κB and proinflammatory cytokine production [129]
The cGAS-STING pathway extensively influences inflammation, apoptosis, necroptosis, and pyroptosis in sepsis, with impaired autophagy potentially leading to its aberrant activation [128].
NLRP3 Inflammasome Activation
The NLRP3 inflammasome, a macromolecular complex comprising NLRP3, ASC, and caspase-1, plays a central role in mtDAMP-mediated inflammation [128] [129]. Mitochondrial dysfunction promotes mtDNA release into the cytoplasm, where oxidized mtDNA preferentially activates NLRP3 [129]. This activation leads to caspase-1-mediated processing of pro-IL-1β and pro-IL-18 into their active forms, and cleavage of gasdermin D, which forms plasma membrane pores that facilitate cytokine release and pyroptosis [129]. Recent studies further indicate that new mtDNA synthesis is a key step in NLRP3 activation [129].
Diagram 1: mtDAMP Signaling Pathways in Sepsis. Mitochondrial damage releases mtDAMPs that activate multiple pattern recognition receptors, converging on inflammatory responses that drive organ dysfunction.
Methodologies for Biomarker Investigation
Bioinformatics Approaches
Comprehensive bioinformatics analyses have been instrumental in identifying mitochondrial biomarkers for sepsis. Standard methodologies include:
Differential Gene Expression Analysis:
- Data sources: GEO datasets (e.g., GSE58294, GSE154918 for stroke and septic shock; GSE54514, GSE65682, GSE95233 for sepsis) [132] [131]
- Analysis tools: R package "limma" for background correction and normalization
- Criteria: False discovery rate (FDR) < 0.05, fold change > 1.5-2.0 [132] [131]
Weighted Gene Co-expression Network Analysis (WGCNA):
- Identifies gene modules correlated with clinical phenotypes
- Uses soft-thresholding to construct scale-free co-expression networks
- Applies topological overlap matrices to measure gene connectivity [132] [131]
Protein-Protein Interaction (PPI) Network Analysis:
- STRING database for interaction prediction (confidence >0.4)
- Cytoscape visualization and module identification
- Algorithm integration (MCC, MNC, DMNC) for hub gene identification [132]
Experimental Validation Techniques
Gene Expression Analysis:
- RNA sequencing of peripheral blood samples from sepsis patients versus healthy controls
- qPCR validation of candidate biomarkers
- Single-cell RNA sequencing to identify cell-type specific expression patterns [133]
Functional Assessment:
- siRNA-mediated gene knockdown (e.g., siNDUFB3: 5'-GAUUAUAGACAAUGGAAGATT-3') [131]
- LPS-induced sepsis models in cell cultures (e.g., H9C2 cells)
- Mitochondrial function assays:
mtDAMP Measurement:
- Quantitative PCR for circulating mtDNA levels
- ELISA-based methods for specific mtDAMPs
- Biochemical assays for ATP, cytochrome c, and cardiolipin
Diagram 2: Biomarker Identification Workflow. Integrated computational and experimental approach for identifying and validating mitochondrial biomarkers in sepsis.
The Scientist's Toolkit: Research Reagent Solutions
Table 4: Essential Research Reagents for Mitochondrial Biomarker Investigation
| Reagent/Category | Specific Examples | Research Application | Key Functions |
|---|---|---|---|
| Gene Expression Analysis | PAXgene Blood RNA System, TRIzol reagent, Agilent 2100 bioanalyzer | RNA sequencing, transcriptome profiling [133] | Sample stabilization, RNA purification, quality control |
| Computational Tools | R package "limma", WGCNA, STRING database, Cytoscape | Bioinformatics analysis, network construction [132] [131] | Differential expression, co-expression networks, PPI analysis |
| Mitochondrial Probes | TMRM, TPP+-selective electrode, JC-1, Mito-tracker | Membrane potential assessment, mitochondrial visualization [130] [131] | ΔΨ measurement, morphology analysis, mass quantification |
| Functional Assays | ATP detection kits, ROS assay kits, caspase-1 activity assays | Biochemical assessment of mitochondrial function [131] | Bioenergetic capacity, oxidative stress, inflammasome activation |
| Gene Modulation | siRNA for NDUFB3, CRISPR/Cas9 systems | Functional validation of candidate genes [131] | Target gene knockdown, mechanistic studies |
| Cell Culture Models | H9C2 cells, LPS stimulation, primary immune cells | In vitro sepsis modeling [131] | Pathway analysis, therapeutic screening |
The integration of NDUFB3 and mtDAMPs as clinical indicators represents a paradigm shift in sepsis biomarker research, moving beyond traditional inflammatory markers to targets rooted in cellular metabolic dysfunction. The evidence supporting their utility is compelling: NDUFB3 demonstrates significant overexpression in sepsis and contributes directly to mitochondrial dysfunction, while mtDAMPs provide a mechanistic link between mitochondrial injury and the hyperinflammatory state that characterizes severe sepsis.
Future research directions should focus on:
- Standardization of detection methods for consistent mtDAMP measurement across clinical settings
- Longitudinal profiling to establish dynamic changes in these biomarkers throughout sepsis progression and recovery
- Multi-omics integration combining mitochondrial biomarkers with proteomic, metabolomic, and transcriptomic data
- Therapeutic targeting of mtDAMP pathways or mitochondrial quality control mechanisms
The investigation of these mitochondrial biomarkers deepens our understanding of sepsis pathophysiology while creating opportunities for improved patient stratification, timely intervention, and targeted therapies. As research progresses, these biomarkers may form the foundation of precision medicine approaches for this complex and devastating condition.
Mitochondrial membrane potential (ΔΨm) is the electrical gradient across the inner mitochondrial membrane that serves as the principal component of the proton motive force (PMF) driving ATP synthesis [2]. Beyond this canonical energetic role, ΔΨm functions as a dynamic signaling hub that regulates reactive oxygen species production, calcium handling, and mitochondrial quality control, enabling localized and time-sensitive regulation of cellular function [2]. This regulatory capacity positions ΔΨm as a critical factor in metabolic specialization—the process by which mitochondria partition metabolic functions to meet specific cellular demands.
Investigating ΔΨm presents substantial methodological challenges, particularly in accessing relevant human tissues. This technical guide examines two fundamentally distinct experimental systems: peripheral blood mononuclear cells (PBMCs) and long-lived post-mitotic tissues, evaluating their respective utilities for mitochondrial membrane potential research within drug development and basic science contexts.
Fundamental Biological Differences
PBMCs and post-mitotic tissues represent contrasting extremes in cellular lifespan, renewal capacity, and primary functions, which fundamentally shape their mitochondrial biology.
Table 1: Fundamental Characteristics of PBMCs vs. Post-Mitotic Tissues
| Characteristic | PBMCs | Long-Lived Post-Mitotic Tissues |
|---|---|---|
| Cell Types | Lymphocytes, monocytes [134] | Neurons, cardiac myocytes, skeletal muscle fibers [135] |
| Renewal Capacity | Short-lived, continuously replenished [135] | Rarely or never replaced, can be as old as the organism [135] |
| Primary Metabolic Focus | Immune activation and response [136] | Sustained energy production for specialized functions [135] |
| Mitochondrial Turnover | High turnover rate | Slow turnover, prone to accumulation of damaged components [135] |
| Vulnerability to Aging | Minimal damage accumulation due to dilution through division [135] | High vulnerability to age-related damage accumulation [135] |
Implications for Mitochondrial Research
The differential longevity of these cell types creates distinct experimental considerations. PBMCs, as short-lived cells, do not accumulate substantial amounts of damaged structures during their lifespan, making them suitable for assessing acute interventions or recent pathological insults [135]. In contrast, long-lived post-mitotic cells progressively accumulate dysfunctional mitochondria, oxidatively modified proteins, and lysosomal garbage (lipofuscin), reflecting chronic processes and aging [135]. This fundamental difference dictates their applicability to various research timelines and questions.
Comparative Functional Parameters
Direct measurements of mitochondrial function reveal stark contrasts between these systems, reflecting their specialized physiological roles.
Table 2: Experimentally Measured Functional Parameters
| Parameter | PBMCs | Post-Mitotic Tissues | Experimental Significance |
|---|---|---|---|
| Spare Respiratory Capacity | Decreased in pediatric septic shock (1.81 vs 5.55 pmol O₂/s/10⁶ cells) [134] | Not directly measured in results | Indicator of bioenergetic reserve during stress |
| Mitochondrial Membrane Potential (ΔΨm) | Associated with organ failure-free days in sepsis [134]; Depolarized in ALS patients [137] | High in healthy state; decreased in ovarian cancer tissue [138] | Primary driver of ATP synthesis and cellular signaling [2] |
| LEAK/Maximal Respiration Ratio | Increased in early sepsis (17% vs <1%) [134] | Not available in results | Indicator of mitochondrial uncoupling |
| Mitochondrial Swelling | Not typically measured | Observed in ovarian cancer tissues [138] | Indicator of mitochondrial permeability transition |
| Response to Damage | Dynamic recalibration in response to stressors [136] | Progressive dysfunction with accumulation of defective organelles [135] | Differential predictive value for acute vs. chronic conditions |
Experimental Approaches and Methodologies
PBMC Isolation and ΔΨm Assessment
PBMC Isolation Protocol:
- Collect whole blood in heparinized tubes [137]
- Dilute blood 1:1 with balanced salt solution [134]
- Layer carefully over Ficoll-Paque PLUS density gradient (density 1.077 g/mL) [134] [137]
- Centrifuge at 400g for 40 minutes at 20-25°C [134]
- Aspirate the lymphocyte/monocyte layer at the interface [134]
- Centrifuge again at 1800g for five minutes to pellet PBMCs [134]
- Resuspend in appropriate buffer for subsequent assays [134]
Critical Considerations:
- Platelet Contamination: PBMC preparations are naturally contaminated with platelets, which contain mitochondria and mtDNA but no nuclear genome, representing a major source of bias in mitochondrial studies [136]
- Cell Subtype Variability: Mitochondrial measures are confounded by cell type distributions; different immune cell subtypes (monocytes, B cells, T cell subtypes) have distinct mitochondrial phenotypes [136]
- Sample Timing: Immune cell mitochondrial function shows dynamic variation in response to diurnal rhythms, exercise, and stress [136]
ΔΨm Measurement Techniques
Fluorescent Dye-Based Assessment: Two primary strategies are employed for semi-quantitative ΔΨm analysis in living cells:
- Steady-State Measurements - suited for comparing ΔΨm between different conditions [139]
- Dynamic Measurements - allowing temporal monitoring of ΔΨm changes in response to perturbations [139]
Commonly Used Potentiometric Dyes:
- TMRM (Tetramethylrhodamine methyl ester): Used in non-quenching/redistribution mode [139]
- Rhodamine 123: Applied in quenching mode [139]
- JC-1: Exhibits potential-dependent accumulation in mitochondria, indicated by fluorescence emission shift from green (~529 nm) to red (~590 nm) [138]
- TMRE (Tetramethylrhodamine ethyl ester): Fluorescent cationic indicator that accumulates in polarized mitochondria [137]
Experimental Workflow for ΔΨm Measurement:
Mitochondrial Respiration Assessment
Protocol for Oxygen Consumption Measurements in PBMCs:
- Resuspend intact PBMCs in Hank's balanced salt solution with 5.5 mM glucose, 1mM pyruvate, and 10 mM HEPES (pH 7.40) [134]
- Place PBMCs in oxygraph chamber at concentration of 1-2 × 10⁶ cells/mL [134]
- Record basal oxygen consumption for 10-20 minutes [134]
- Add ATP-synthase inhibitor oligomycin (1 μg/mL) to induce LEAK respiration [134]
- Titrate uncoupler CCCP (1-2 μM) until maximal respiration is achieved [134]
- Calculate spare respiratory capacity (difference between maximal and basal respiration) [134]
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Research Reagents for ΔΨm Studies
| Reagent/Category | Specific Examples | Function/Application | Considerations |
|---|---|---|---|
| PBMC Isolation | Ficoll-Paque PLUS [134] [137] | Density gradient medium for PBMC separation | Batch variability; optimal density 1.077 g/mL [134] |
| ΔΨm Dyes | TMRM, Rhodamine 123 [139], JC-1 [138], TMRE [137] | Potentiometric indicators of mitochondrial polarization | Concentration-dependent artifacts; choose quenching vs. non-quenching modes appropriately [139] |
| Respiratory Inhibitors/Modulators | Oligomycin, CCCP/FCCP, Rotenone [134] [140] | Modulate electron transport chain function to assess respiratory parameters | Titration required for optimal concentration [134] |
| Permeabilization Agents | XF Plasma Membrane Permeabilizer (PMP) [140] | Enables substrate access to mitochondria in intact cells | Requires optimization for different cell types [140] |
| Metabolic Substrates | Pyruvate, Glutamate, Malate, Succinate [140] | Specific substrates for individual electron transport chain complexes | Enables assessment of specific respiratory pathways [140] |
Research Applications and Pathological Insights
PBMCs as Accessible Biomarkers
PBMCs have demonstrated utility as biomarkers for systemic mitochondrial dysfunction across various pathologies:
- Sepsis: Pediatric septic shock patients exhibited decreased spare respiratory capacity and increased uncoupling in PBMCs, with ΔΨm associated with duration of organ injury [134]
- Neurodegenerative Disease: ALS patients showed significant mitochondrial depolarization in PBMCs, resembling findings from neural cell models [137]
- Cancer: PBMCs from breast cancer patients showed lower basal ATP and mitochondrial DNA copy numbers compared to healthy controls [140]
Post-Mitotic Tissues in Chronic Disease
Long-lived post-mitotic tissues reveal distinct patterns of mitochondrial dysfunction:
- Ovarian Cancer: Mitochondrial membrane depolarization and mitochondrial swelling were observed in ovarian tissues of cancer patients compared to normal ovaries [138]
- Aging: Long-lived post-mitotic cells show progressive accumulation of dysfunctional, enlarged mitochondria and lipofuscin-loaded lysosomes, creating a vicious cycle of deteriorating mitochondrial quality control [135]
Integration Framework for Multi-Tissue Mitochondrial Assessment
PBMCs and post-mitotic tissues offer complementary windows into mitochondrial membrane potential regulation, each with distinct advantages and limitations. PBMCs provide an accessible system for monitoring dynamic mitochondrial responses to acute insults and interventions, though their heterogeneity requires careful experimental control. Post-mitotic tissues reveal the cumulative burden of mitochondrial dysfunction in chronic disease and aging, despite challenges in sampling. The integration of findings from both systems, with careful attention to their fundamental biological differences, provides the most comprehensive approach for advancing mitochondrial medicine in metabolic specialization research. Future methodological developments should focus on standardizing measurements across tissue types and accounting for cellular heterogeneity to maximize the translational potential of mitochondrial membrane potential studies.
The journey from a therapeutic concept to a clinically approved treatment is notoriously long, expensive, and fraught with failure. Traditional drug development has heavily relied on animal models for preclinical validation, yet these models often suffer from interspecies differences that lead to poor prediction of human physiological and pathological conditions [141]. This translational gap is a primary reason approximately 60% of clinical trials fail due to lack of efficacy, while 30% fail due to toxicity [142]. In response, biomedical research is undergoing a paradigm shift toward approaches centered on bioengineered human disease models that offer higher clinical mimicry and predictability [141].
Concurrently, our understanding of fundamental biological processes has evolved, particularly regarding mitochondrial function. Beyond their canonical role as cellular powerplants, mitochondria are now recognized as dynamic signaling hubs that regulate cellular fate through processes like metabolic specialization and partitioning [2]. The mitochondrial membrane potential (MMP), a key component of the protonmotive force, sits at the center of this regulation, acting as a critical mediator that influences reactive oxygen species production, calcium handling, and quality control mechanisms [2]. This article explores how successful therapeutic validation in advanced preclinical models is increasingly reliant on understanding these mitochondrial dynamics, which enable localized and time-sensitive regulation of cellular function in health and disease.
Mitochondrial Membrane Potential: A Central Regulator in Metabolic Partitioning
Fundamental Principles and Signaling Roles
The mitochondrial membrane potential (MMP) represents an electrical gradient across the inner mitochondrial membrane, typically around -180 mV, generated by the electron transport chain during oxidative phosphorylation [2]. This charge separation serves as the primary component of the protonmotive force that drives ATP synthesis. However, contemporary research reveals that the MMP's role extends far beyond energy production, functioning as a dynamic signaling hub that integrates cellular status and directs functional outcomes.
MMP undergoes both rapid adjustments to acute changes in cellular energy demand and sustained modifications during developmental processes [2]. These dynamics influence critical cellular processes including reactive oxygen species production, calcium handling, and mitochondrial quality control. The MMP facilitates metabolic specialization by shaping the activity of specific metabolic enzymes, enabling the emergence of functionally distinct mitochondrial subpopulations tailored to specific metabolic demands [2]. For instance, elevated MMP relative to baseline enhances the activity of pyrroline-5-carboxylate synthase (P5CS), promoting the formation of filamentous assemblies that drive reductive biosynthesis for molecular precursor synthesis [2].
MMP in Quality Control and Cellular Decision-Making
Mitochondria exist not as isolated organelles but as dynamic networks that undergo continuous remodeling through fission and fusion events. The MMP plays a decisive role in the fate of mitochondrial fragments generated through fission. Daughter mitochondria retaining higher MMP often re-fuse with the network, while those with significantly reduced MMP are targeted for degradation via mitophagy [2]. This quality control mechanism is crucial for eliminating damaged mitochondrial components and maintaining network integrity.
The process is primarily driven by MMP loss, which leads to the accumulation of PINK1 on the mitochondrial surface, subsequently recruiting Parkin and LC3 to mark the organelle for degradation [143]. This binary fate decision implies the existence of MMP thresholds that direct mitochondria toward either biogenesis or clearance, representing a critical aspect of metabolic partitioning at the organellar level [2]. Furthermore, regional variations in MMP influence protein import machinery, potentially affecting local mitochondrial composition and function, and contributing to the establishment of specialized subpopulations [142].
Table 1: Key Functional Roles of Mitochondrial Membrane Potential in Cellular Physiology
| Functional Role | Mechanism | Physiological Impact |
|---|---|---|
| Energetic Regulation | Drives ATP synthesis via ATP synthase | Maintains cellular energy homeostasis |
| Metabolic Specialization | Regulates enzyme activity (e.g., P5CS filamentation) | Directs switching between oxidative and reductive metabolism |
| Quality Control | Serves as sensor for PINK1/Parkin-mediated mitophagy | Ensures removal of damaged mitochondria |
| Calcium Handling | Influences mitochondrial calcium uptake capacity | Regulates calcium-dependent signaling and buffering |
| ROS Production | Modulates electron leak from transport chain | Affects redox signaling and oxidative stress |
Success Stories in Preclinical Therapeutic Validation
Human Organoid Models for Cystic Fibrosis
Cystic fibrosis (CF), caused by mutations in the CFTR gene, has witnessed remarkable advances through the application of human colon and airway organoids derived from patient-derived stem cells [141]. These self-organizing three-dimensional structures recapitulate key aspects of the native epithelium, including functional CFTR protein expression and disease pathophysiology [141].
The forskolin-induced swelling assay has emerged as a particularly powerful functional readout for CFTR activity in these organoids. When exposed to forskolin, which activates CFTR via cAMP-mediated signaling, organoids from healthy donors swell significantly due to ion and water flux into the lumen. Conversely, CF organoids with impaired CFTR function show reduced or absent swelling [141]. This quantitative assay has been successfully standardized and validated for high-throughput drug screening, enabling researchers to test CFTR modulators that restore channel function [141]. The platform has proven instrumental in validating the efficacy of CFTR correctors and potentiators, some of which have advanced to clinical application, demonstrating the exceptional predictive value of this human-derived model system.
Table 2: Experimental Methodology for Forskolin-Induced Swelling Assay in CF Organoids
| Protocol Step | Key Parameters | Technical Considerations |
|---|---|---|
| Organoid Culture | Long-term expanding human airway or colon organoids [141] | Maintain genetic stability over multiple passages |
| Experimental Setup | Treatment with CFTR modulators followed by forskolin challenge | Include appropriate controls (DMSO, corrector/potentiator alone) |
| Image Acquisition | Bright-field microscopy at regular intervals (e.g., every 15-30 min for 1-2 h) | Maintain consistent focus and lighting conditions |
| Quantitative Analysis | Measure organoid surface area over time using automated image analysis | Normalize to baseline area; report fold-change or area under curve |
| Data Interpretation | Compare swelling response between treatment and control groups | Correlate with electrophysiological measurements (e.g., USsing chamber) |
Liver-on-Chip Technology for Predicting Drug-Induced Liver Injury
Drug-induced liver injury (DILI) remains a leading cause of drug attrition during development and post-marketing withdrawal. Conventional models, including animal studies and hepatic spheroid cultures, have demonstrated limited predictivity for human hepatotoxicity [142]. The Emulate Liver Chip, a microfluidic device lined with living human cells, has emerged as a transformative platform for DILI prediction.
This bioengineered system, derived from the Wyss Institute at Harvard University, incorporates primary human hepatocytes alongside non-parenchymal cells in a physiologically relevant architecture that recapitulates key aspects of liver tissue function [142]. The platform recently achieved a significant regulatory milestone when the FDA's Center for Drug Evaluation and Research (CDER) accepted its first letter of intent for an organ-on-a-chip technology as a drug development tool [142]. Validation studies demonstrated that the Liver Chip better predicted human-relevant drug-induced liver injury compared to animal models and hepatic spheroids, highlighting its potential to improve safety assessment in preclinical development [142].
Zebrafish Models for Duchenne Muscular Dystrophy and Melanoma
Zebrafish models have become indispensable tools for studying human diseases, offering a unique combination of genetic tractability, optical transparency, and physiological similarity to mammals. In Duchenne muscular dystrophy research, zebrafish with knockout of the dystrophin gene faithfully recapitulate key disease hallmarks, including muscle fiber necrosis, inflammation, fibrosis, and abnormally sized muscle fibers [144]. These models enable real-time observation of disease progression and high-throughput screening of therapeutic compounds that mitigate muscle degeneration [144].
In cancer research, zebrafish models have provided unprecedented insights into melanoma pathogenesis. A knock-in zebrafish model with the human BRAF mutation—a common driver in melanoma—successfully replicates tumor formation [144]. When combined with other cancer-associated genes like SETDB1, these zebrafish develop melanoma rapidly, providing a platform to identify key genetic contributors and test targeted therapies that have subsequently informed clinical trial designs [144]. The ability to observe tumor formation, progression, and metastasis in real time has accelerated our understanding of melanoma genetics and facilitated anti-cancer drug screening.
The Scientist's Toolkit: Essential Reagents and Methodologies
Research Reagent Solutions for Mitochondrial Function Analysis
Table 3: Essential Research Reagents for Mitochondrial Function and Therapeutic Validation Studies
| Reagent/Method | Function/Application | Key Insights |
|---|---|---|
| Tetramethylrhodamine Methyl Ester (TMRM) | Potentiometric fluorescent dye for measuring MMP [16] | Quantitative assessment of mitochondrial polarization status; use in live-cell imaging |
| MitoSOX | Fluorogenic dye for detecting mitochondrial superoxide [16] | Selective for mitochondrial ROS; indicates oxidative stress status |
| Rhod-2 AM | Fluorescent indicator for mitochondrial calcium [16] | Monitors Ca²⁺ dynamics within mitochondria; requires AM ester for loading |
| Forskolin-Induced Swelling Assay | Functional test for CFTR activity in organoids [141] | Gold-standard method for validating CFTR modulator efficacy in human organoids |
| Organ Chip Microfluidic Devices | Bioengineered human cell-based systems for disease modeling [142] | Physiologically relevant platforms for toxicity testing and disease modeling |
Experimental Protocol: Multiparameter Assessment of Mitochondrial Function
The simultaneous assessment of mitochondrial membrane potential, reactive oxygen species, and calcium levels provides a comprehensive picture of mitochondrial health and function. Below is a detailed methodology adapted from established protocols [16]:
Cell Preparation and Staining: Culture cells on appropriate substrates (e.g., glass coverslips, Organ Chips). For adherent cells, achieve 70-80% confluency. Prepare staining solution containing TMRM (50-100 nM), MitoSOX (5 µM), and Rhod-2 AM (2-5 µM) in pre-warmed culture medium. Protect from light during all subsequent steps.
Dye Loading: Replace culture medium with staining solution. Incubate at 37°C for 20-30 minutes. Include control samples stained with individual dyes for compensation.
Washing and De-esterification: Remove staining solution and wash cells gently with pre-warmed phosphate-buffered saline (PBS). Replace with fresh culture medium and incubate for additional 15 minutes to allow complete de-esterification of intracellular AM esters.
Image Acquisition: Acquire images using a confocal or epifluorescence microscope with appropriate filter sets. For TMRM (MMP), use excitation/emission of ~548/573 nm. For MitoSOX (ROS), use ~510/580 nm. For Rhod-2 AM (calcium), use ~552/581 nm. Maintain identical acquisition parameters across experimental groups.
Data Analysis: Quantify fluorescence intensity using image analysis software (e.g., ImageJ, CellProfiler). Normalize values to control conditions. Report changes in fluorescence intensity relative to baseline for each parameter.
Visualizing Experimental Workflows and Signaling Pathways
Mitochondrial Decision-Making in Metabolic Partitioning
Therapeutic Validation Workflow Using Advanced Models
The integration of bioengineered human disease models into therapeutic validation represents a transformative advancement in preclinical research. Success stories across diverse platforms—from organoids and Organs-on-Chips to zebrafish models—demonstrate their growing impact on drug development [141] [142] [144]. These models, when combined with a deeper understanding of fundamental regulatory mechanisms like mitochondrial membrane potential and metabolic partitioning, offer unprecedented opportunities to bridge the translational gap that has long plagued pharmaceutical development [2].
Future progress will depend on continued refinement of these systems, including enhancing their complexity through the incorporation of immune components, vascularization, and multi-tissue interactions. Furthermore, stringent model validation, regulatory guidance, and scalable production are key milestones that will facilitate broader implementation in preclinical research [141]. As these technologies mature and our understanding of mitochondrial biology deepens, we anticipate a new era of drug development where therapeutic validation more accurately predicts clinical success, ultimately bringing effective treatments to patients faster and more efficiently.
Conclusion
Mitochondrial membrane potential emerges as a central regulator of cellular metabolism, orchestrating metabolic specialization through precise partitioning of mitochondrial functions. The integration of MMP assessment with respirometry and other functional assays provides a powerful framework for understanding its role in health and disease. Current evidence strongly implicates MMP dysregulation in diverse pathologies from cancer to neurodegeneration, offering compelling therapeutic targets. Future research must focus on developing standardized methodologies for clinical translation, exploring tissue-specific MMP dynamics, and advancing MMP-targeted therapeutics. The continued elucidation of MMP-mediated metabolic partitioning will undoubtedly yield transformative insights for precision medicine and drug development.
References
- 1. Use the protonmotive force: mitochondrial uncoupling and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC6170739/]
- 2. Mitochondrial membrane potential and compartmentalized ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12454667/]
- 3. Control Over the Contribution of the Mitochondrial ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC2662253/]
- 4. Measuring mitochondrial membrane potential | The EMBO ... [https://link.springer.com/article/10.1038/s44318-025-00632-9]
- 5. Quantitative measurement of mitochondrial membrane ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC3448152/]
- 6. Implications of mitochondrial membrane potential gradients ... [https://www.nature.com/articles/s41598-024-65595-z]
- 7. Mitochondrial energy dissipation by fatty acids. ... [https://pubmed.ncbi.nlm.nih.gov/12481544/]
- 8. Quantitative analysis of mitochondrial morphology and ... [https://www.sciencedirect.com/science/article/pii/S0167488914003917]
- 9. Matrix metalloproteases: Underutilized targets for drug delivery [https://pmc.ncbi.nlm.nih.gov/articles/PMC3782085/]
- 10. MMP3 at the crossroads: Linking molecular pathways to ... [https://www.sciencedirect.com/science/article/pii/S1043661825001756]
- 11. Interactions between Notch and matrix metalloproteinases [https://journals.viamedica.pl/nowotwory_journal_of_oncology/article/view/95279/75857]
- 12. Non-canonical role of matrix metalloprotease (MMP) in ... [https://pubmed.ncbi.nlm.nih.gov/27140333/]
- 13. Non-canonical role of matrix metalloprotease (MMP) in ... [https://www.sciencedirect.com/science/article/abs/pii/S002432051630251X]
- 14. Identification and validation of matrix metalloproteinase ... [https://www.frontiersin.org/journals/oncology/articles/10.3389/fonc.2024.1471267/full]
- 15. Elevated mitochondrial membrane potential is a therapeutic ... [https://pubmed.ncbi.nlm.nih.gov/40240771/]
- 16. Analysis of mitochondrial membrane potential, ROS, and ... [https://www.sciencedirect.com/science/article/pii/S1016847825000627]
- 17. Systematic druggable genome-wide Mendelian ... [https://www.medrxiv.org/content/10.1101/2025.01.19.25320804v1.full-text]
- 18. Mitochondrial Membrane Potential - an overview [https://www.sciencedirect.com/topics/medicine-and-dentistry/mitochondrial-membrane-potential]
- 19. Mitochondrial membrane potential acts as a retrograde ... [https://www.life-science-alliance.org/content/6/12/e202302091]
- 20. High mtDNA content identifies oxidative phosphorylation ... [https://www.nature.com/articles/s41392-025-02303-x]
- 21. Advances in Mitochondrial Dysfunction and Its Role ... [https://www.mdpi.com/2073-4409/14/20/1621]
- 22. Mitochondrial Membrane Potential Fluorescent Probes ... [https://www.linkedin.com/pulse/mitochondrial-membrane-potential-fluorescent-jxswe/]
- 23. Mitochondrial membrane potential and compartmentalized ... [https://www.sciencedirect.com/science/article/pii/S2213231725003726]
- 24. Cellular ATP demand creates metabolically distinct ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11869630/]
- 25. Mitochondrial heterogeneity: subpopulations with distinct ... [https://www.nature.com/articles/s41392-025-02130-0]
- 26. Mitochondria's division of labor sheds light on how cancer ... [https://www.news-medical.net/news/20241112/Mitochondriae28099s-division-of-labor-sheds-light-on-how-cancer-cells-survive-harsh-conditions.aspx]
- 27. Structural basis of dynamic P5CS filaments [https://elifesciences.org/articles/76107]
- 28. Dynamic Arabidopsis P5CS filament facilitates substrate ... [https://www.nature.com/articles/s41477-024-01697-w]
- 29. Emergence of Specialized Mitochondrial Networks: A 'Divide ... [https://www.gethealthspan.com/research/article/specialized-mitochondrial-networks?srsltid=AfmBOoojeCFZDow3dXai5bmMTLT28QhEotu-v7So-ssHxN-onfricNHF]
- 30. Possible Role of Novel Mitochondrial Subsets in Migraine [https://www.mdpi.com/2075-1729/15/8/1273]
- 31. Cardiac Subsarcolemmal and Interfibrillar ... | PLOS One Mitochondria [https://journals.plos.org/plosone/article?id=10.1371/journal.pone.0044667]
- 32. Biochemical properties of subsarcolemmal and interfibrillar ... [https://www.sciencedirect.com/science/article/pii/S0021925819752831]
- 33. Subcellular Specialization of Mitochondrial Form and ... [https://www.frontiersin.org/journals/cell-and-developmental-biology/articles/10.3389/fcell.2021.757305/full]
- 34. Subsarcolemmal and intermyofibrillar mitochondria play ... [https://pubmed.ncbi.nlm.nih.gov/15647392/]
- 35. Mitochondria-Targeted Drugs - PMC - PubMed Central [https://pmc.ncbi.nlm.nih.gov/articles/PMC6875871/]
- 36. Pharmacological targeting of mitochondria in disease - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC2189819/]
- 37. The Interplay Between Mitochondrial Dynamics and Mitophagy [https://pmc.ncbi.nlm.nih.gov/articles/PMC3078508/]
- 38. Simultaneous impairment of mitochondrial fission and ... [https://www.nature.com/articles/srep07885]
- 39. Spontaneous Changes in Mitochondrial Membrane Potential ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC6762873/]
- 40. JC-1 Dye for Mitochondrial Membrane Potential [https://www.thermofisher.com/us/en/home/life-science/cell-analysis/cell-viability-and-regulation/apoptosis/mitochondria-function/jc-1-dye-mitochondrial-membrane-potential.html]
- 41. High-Throughput Small Molecule Screen Identifies ... [https://www.sciencedirect.com/science/article/pii/S2589004220301152]
- 42. High-Throughput Microscopy Analysis of Mitochondrial ... [https://www.mdpi.com/2073-4409/12/7/1089]
- 43. Introduction to the Mitochondrial Stress (MitoPT) Kits [https://www.antibodiesinc.com/pages/introduction-mitochondrial-stress-kits?srsltid=AfmBOop5tkAM_4D454InqwMv2WMZxd-AoEEg4Mokp_SV-oKqrXMsnvvy]
- 44. TMRM Assay Kit (Mitochondrial Membrane Potential) [https://www.abcam.com/en-us/products/assay-kits/tmrm-assay-kit-mitochondrial-membrane-potential-ab228569]
- 45. What is the difference between MitoPT TMRE and JC-1? [https://www.antibodiesinc.com/blogs/news/ict-answers-your-questions-5?srsltid=AfmBOopk3TOFd3e89wERKgJHaz3wH7KWsdmB6SJ2v1Lw4aUljBkhnFZ5]
- 46. JC-1 MitoMP Detection Kit [https://www.dojindo.com/products/MT09/]
- 47. High-throughput measurement of mitochondrial membrane ... [https://www.sciencedirect.com/science/article/abs/pii/S0006291X02025469]
- 48. Establishment of High-Throughput Screening Protocol ... [https://pubmed.ncbi.nlm.nih.gov/40877025/]
- 49. Protease assays using CyDye fluors & FRET [https://www.bmglabtech.com/en/application-notes/use-of-cydye-fluors-for-improved-fret-protease-assays-on-a-bmg-labtech-fluorescence-microplate-reader/]
- 50. Advances in high-throughput drug screening based on ... [https://www.sciencedirect.com/science/article/pii/S2090123225006897]
- 51. Overview of methods that determine mitochondrial function ... [https://www.sciencedirect.com/science/article/pii/S0026049525001696]
- 52. Artificial Intelligence in Assessing Reproductive Aging: Role of ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12386087/]
- 53. The trade-off between individual metabolic specialization ... [https://www.sciencedirect.com/science/article/pii/S2405471223003587]
- 54. A practical guide for the analysis, standardization, and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9618452/]
- 55. Using flow cytometry for mitochondrial assays - PubMed Central [https://pmc.ncbi.nlm.nih.gov/articles/PMC7289760/]
- 56. Imaging flow cytometry reveals divergent mitochondrial ... [https://www.sciencedirect.com/science/article/pii/S2589004224027238]
- 57. Protocol for measuring mitochondrial respiratory capacity from ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12339751/]
- 58. Harmonization of experimental procedures to assess ... [https://www.sciencedirect.com/science/article/pii/S0891584924005859?via%3Dihub]
- 59. Mitochondrial Membrane Potential - an overview [https://www.sciencedirect.com/topics/biochemistry-genetics-and-molecular-biology/mitochondrial-membrane-potential]
- 60. Mitochondria targeting molecular transporters [https://pmc.ncbi.nlm.nih.gov/articles/PMC8757599/]
- 61. Recent Advancements in Mitochondria-Targeted Nanoparticle ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC8911864/]
- 62. Triphenylphosphine-Based Mitochondrial Nanocarriers... Targeting [https://www.dovepress.com/triphenylphosphine-based-mitochondrial-targeting-nanocarriers-advancin-peer-reviewed-fulltext-article-CPAA]
- 63. Mitochondrial Membrane Potential - an overview [https://www.sciencedirect.com/topics/neuroscience/mitochondrial-membrane-potential]
- 64. Advancements in mitochondrial-targeted nanotherapeutics [https://www.frontiersin.org/journals/immunology/articles/10.3389/fimmu.2024.1451989/full]
- 65. Frontiers | Mitochondria - targeted strategies in tumor immunity [https://www.frontiersin.org/journals/immunology/articles/10.3389/fimmu.2025.1646138/full]
- 66. Overview of MitoQ on prevention and management of ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11934253]
- 67. Pharmacological significance of MitoQ in ameliorating ... [https://www.sciencedirect.com/science/article/pii/S2667137922000091]
- 68. MitoQ regulates autophagy by inducing a pseudo ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC5388232/]
- 69. The mitochondrially targeted antioxidant MitoQ protects ... [https://www.nature.com/articles/s41419-018-0436-x]
- 70. Therapeutic potential of the mitochondria-targeted antioxidant ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC6279977/]
- 71. MitoQ Protects Against Oxidative Stress-Induced ... [https://www.sciencedirect.com/science/article/pii/S2772976125001886]
- 72. Mitochondria-targeted antioxidant MitoQ radiosensitizes ... [https://www.nature.com/articles/s41420-024-02277-9]
- 73. Elevated mitochondrial membrane potential is a ... [https://www.nature.com/articles/s41467-025-57238-2]
- 74. evaluating its effects on clinical outcomes and oxidative ... [https://pubmed.ncbi.nlm.nih.gov/40820063/]
- 75. Overview of MitoQ on prevention and management ... [https://www.frontiersin.org/journals/cardiovascular-medicine/articles/10.3389/fcvm.2025.1506460/full]
- 76. Unlocking the Value of White Blood Cells for Heart Failure Diagnosis [https://link.springer.com/article/10.1007/s12265-020-10007-6]
- 77. Development of a multiparameter flow cytometric assay as a potential... [https://translational-medicine.biomedcentral.com/articles/10.1186/s12967-015-0604-z]
- 78. PBMCs Mitochondrial Respiration and Its Relation to Immunity ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12475958/]
- 79. Mitochondria in oxidative stress, inflammation and aging [https://www.nature.com/articles/s41392-025-02253-4]
- 80. Transcriptomic analysis in peripheral blood mononuclear cells ... [https://genesandnutrition.biomedcentral.com/articles/10.1186/s12263-025-00783-8]
- 81. Single-Cell transcriptomic profiles of peripheral blood ... [https://www.nature.com/articles/s41598-025-17078-y]
- 82. Mitochondrial dysfunction of peripheral blood mononuclear ... [https://www.sciencedirect.com/science/article/abs/pii/S1567576924004764]
- 83. “Recommendations for clinical study protocols for immune and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12537938/]
- 84. Mitochondrial heterogeneity in diseases [https://www.nature.com/articles/s41392-023-01546-w]
- 85. Mitochondrial Heterogeneity [https://www.frontiersin.org/journals/genetics/articles/10.3389/fgene.2018.00718/full]
- 86. Future of Mitochondrial Function in Neurodegeneration ... [https://www.sygnaturediscovery.com/news-and-events/blog/from-lab-to-market-the-future-of-mitochondrial-function/]
- 87. Advances in drug therapy for mitochondrial diseases - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC6995731/]
- 88. Mitochondrial dysfunction: mechanisms and advances in ... [https://www.nature.com/articles/s41392-024-01839-8]
- 89. Reducing Platelet Contamination in Blood Samples [https://www.stemcell.com/reducing-platelet-contamination.html]
- 90. Comparative analysis of methods to reduce activation ... [https://www.nature.com/articles/s41598-023-49611-2]
- 91. Analysis of mitochondrial membrane potential, ROS, and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12271852/]
- 92. Recommended Standard Method for Isolating ... [https://www.sigmaaldrich.com/US/en/technical-documents/protocol/clinical-testing-and-diagnostics-manufacturing/hematology/recommended-standard-method?srsltid=AfmBOooPUFQX27j2O6r0YWdz3Sr7-cmx0SinLs8axgWVPCnSVBdvMJZE]
- 93. Development of a new cell isolation device FlowMagicTM [https://pmc.ncbi.nlm.nih.gov/articles/PMC12543197/]
- 94. Proteomic Characterization of Human Peripheral Blood ... [https://www.mdpi.com/1422-0067/26/13/6035]
- 95. LDS 698: A highly sensitive probe for tracking ... [https://pubmed.ncbi.nlm.nih.gov/40816051/]
- 96. MitoTracker Green FM dye concentration optimization [https://www.sciencedirect.com/science/article/abs/pii/S1046202317302190]
- 97. Development of a high throughput screening assay for ... [https://pubmed.ncbi.nlm.nih.gov/12230893/]
- 98. Exploring the limitations of mitochondrial dye as a genuine ... [https://www.nature.com/articles/s42003-024-05964-6]
- 99. Photobleaching and phototoxicity of mitochondria in live ... [https://www.sciencedirect.com/science/article/pii/S2590279224000038]
- 100. Mitochondrial fission process 1 protein: a comprehensive ... [https://www.frontiersin.org/journals/cell-and-developmental-biology/articles/10.3389/fcell.2025.1646072/full]
- 101. Mitochondrial fission genes MTFP1/MTFP2 as predictive ... [https://link.springer.com/article/10.1007/s12672-025-03215-6]
- 102. Mitochondrial segmentation and function prediction in live ... [https://www.nature.com/articles/s41467-025-55825-x]
- 103. Mitochondrial Compartmentalization: Emerging Themes in ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11008732/]
- 104. Mitochondrial Membrane Potential Assay [https://www.sartorius.com/en/applications/life-science-research/live-cell-assays/mitochondrial-membrane-potential]
- 105. Mitochondrial, metabolic and bioenergetic adaptations ... [https://www.nature.com/articles/s41419-025-07596-y]
- 106. The mechanisms of action of mitochondrial targeting ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10635426/]
- 107. Nerve-to-cancer transfer of mitochondria during ... [https://www.nature.com/articles/s41586-025-09176-8]
- 108. Mapping the global research landscape of mitophagy in ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12632019/]
- 109. Mitochondria dysfunction in the pathogenesis of Alzheimer’s disease: recent advances | Molecular Neurodegeneration [https://link.springer.com/article/10.1186/s13024-020-00376-6]
- 110. Mitochondrial dysfunction in Alzheimer’s disease - ScienceDirect [https://www.sciencedirect.com/science/article/pii/S1568163725000595]
- 111. Mitochondrial dysfunction in Parkinson's disease – a key ... [https://molecularneurodegeneration.biomedcentral.com/articles/10.1186/s13024-023-00676-7]
- 112. An Update and Perspectives on Mitochondrial Membrane ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12384588/]
- 113. Links between Mutations Occurring in BPAN, PKAN, MPAN ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12641038/]
- 114. Metabolic alterations in fibroblasts of patients presenting ... [https://www.sciencedirect.com/science/article/pii/S0925443924005350]
- 115. Mitochondrial dysfunction-mediated metabolic remodeling ... [https://www.nature.com/articles/s41420-025-02651-1]
- 116. Mitochondrial Membrane Potential Influences Amyloid Precursor Protein Localization and Aβ Secretion - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC9212216/]
- 117. Advances in Mitochondrial Dysfunction and Its Role in ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12563985/]
- 118. Regulation of mitochondrial dynamics in cardiomyocytes [https://www.frontiersin.org/journals/cell-and-developmental-biology/articles/10.3389/fcell.2025.1652683/full]
- 119. Analysis of mitochondrial membrane potential, ROS, and ... [https://pubmed.ncbi.nlm.nih.gov/40516788/]
- 120. Mitochondrial Dynamics in Aging Heart - PMC - PubMed Central [https://pmc.ncbi.nlm.nih.gov/articles/PMC12650672/]
- 121. Future perspectives on the roles of mitochondrial ... [https://www.sciencedirect.com/science/article/abs/pii/S0024320524001644]
- 122. Obesity causes mitochondrial fragmentation and ... [https://www.nature.com/articles/s42255-024-00978-0]
- 123. Mitochondrial dynamics and genome alterations in cardiac ... [https://www.sciencedirect.com/science/article/pii/S004763742500020X]
- 124. Recent advances in biomarkers for detection and diagnosis of ... [https://eurjmedres.biomedcentral.com/articles/10.1186/s40001-025-03324-6]
- 125. Restoring the infected powerhouse: Mitochondrial quality ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10711241/]
- 126. Mitochondrial quality control in sepsis [https://pubmed.ncbi.nlm.nih.gov/38039825/]
- 127. Unveiling novel therapeutic frontiers in sepsis [https://www.sciencedirect.com/science/article/pii/S1567576925010689]
- 128. Mitochondria-Derived Damage-Associated Molecular Patterns ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC6530230/]
- 129. The role of mtDAMPs in the trauma-induced systemic ... [https://www.frontiersin.org/journals/immunology/articles/10.3389/fimmu.2023.1164187/full]
- 130. (ΔΨ) Fluctuations Associated with... Mitochondrial Membrane Potential [https://pubmed.ncbi.nlm.nih.gov/39546260/]
- 131. Identification and experimental validation of mitochondria ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10203506/]
- 132. Role of COX6C and NDUFB3 in septic shock and stroke - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC11627056/]
- 133. Identification of sepsis-associated mitochondrial genes ... [https://bmcmedgenomics.biomedcentral.com/articles/10.1186/s12920-024-01891-x]
- 134. Mitochondrial Dysfunction in Peripheral Blood Mononuclear ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC4286436/]
- 135. Mitochondrial Turnover and Aging of Long-Lived Postmitotic ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC2861545/]
- 136. phenotypes in purified human immune cell subtypes... Mitochondrial [https://elifesciences.org/articles/70899]
- 137. Decreased Mitochondrial Function, Biogenesis, and ... [https://link.springer.com/article/10.1007/s12035-020-02059-1]
- 138. Alterations in mitochondria isolated from peripheral blood ... [https://www.nature.com/articles/s41598-023-51009-z]
- 139. Visualization of mitochondrial in mammalian cells membrane potential [https://pubmed.ncbi.nlm.nih.gov/32183960/]
- 140. Real-time assessment of relative mitochondrial ATP synthesis... [https://cancerandmetabolism.biomedcentral.com/articles/10.1186/s40170-024-00353-3]
- 141. Human disease models in drug development [https://www.nature.com/articles/s44222-023-00063-3]
- 142. Beyond Animals: Revolutionizing Drug Discovery with Human ... [https://wyss.harvard.edu/news/beyond-animals-revolutionizing-drug-discovery-with-human-relevant-models/]
- 143. Joint Transnational Call (JTC) 2025 "Pre-clinical therapy ... [https://www.era-learn.eu/network-information/networks/rare-diseases/joint-transnational-call-jtc-2025-pre-clinical-therapy-studies-for-rare-diseases-using-small-molecules-and-biologicals-development-and-validation]
- 144. Zebrafish Models of Human Disease: Key Success Stories [https://www.zeclinics.com/blog/zebrafish-models-of-human-disease]