PARP-1 Cleavage: The Definitive Apoptosis Marker and Its Emerging Roles in Cell Death
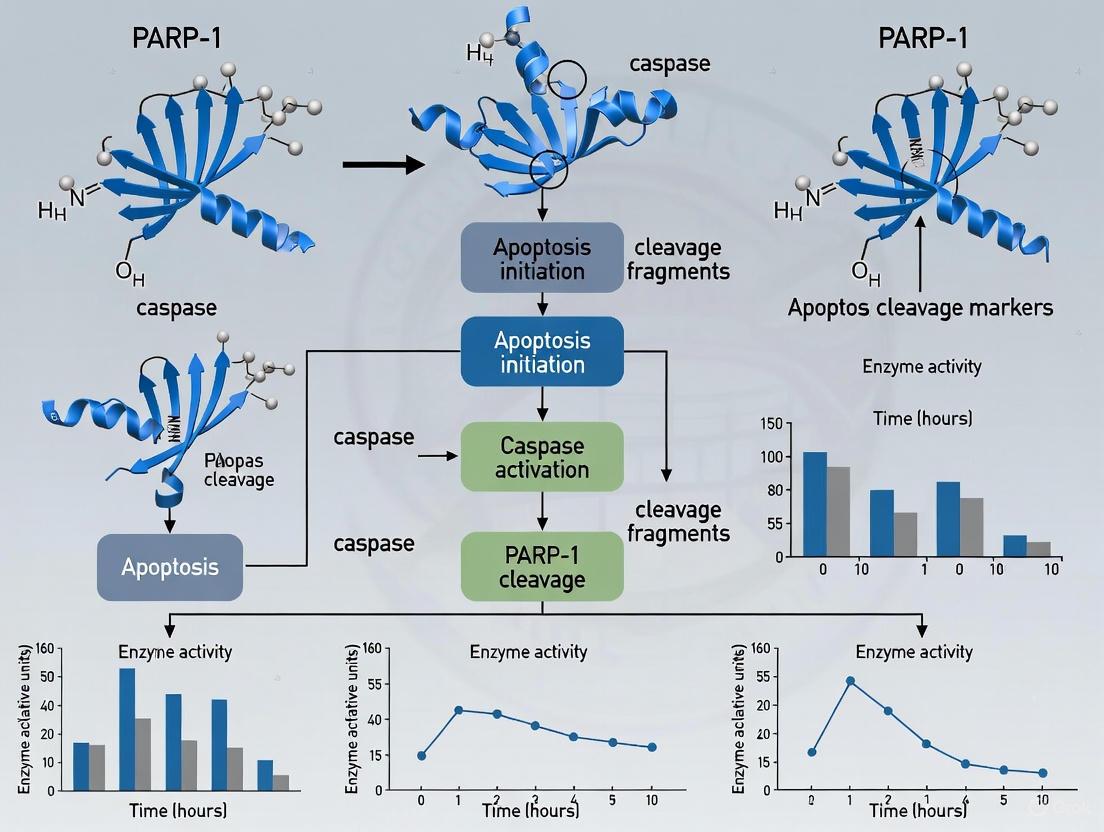
This article provides a comprehensive analysis of poly(ADP-ribose) polymerase-1 (PARP-1) cleavage as a fundamental biomarker of apoptosis.
PARP-1 Cleavage: The Definitive Apoptosis Marker and Its Emerging Roles in Cell Death
Abstract
This article provides a comprehensive analysis of poly(ADP-ribose) polymerase-1 (PARP-1) cleavage as a fundamental biomarker of apoptosis. Tailored for researchers, scientists, and drug development professionals, it explores the foundational mechanism where caspases-3 and -7 cleave PARP-1 into characteristic 24-kDa and 89-kDa fragments, inactivating DNA repair and facilitating programmed cell death. The content details methodological applications for detecting cleavage, addresses common troubleshooting scenarios, and validates PARP-1 cleavage against other cell death markers. Furthermore, it examines emerging research on the novel biological functions of the cleavage fragments, including their roles in parthanatos and innate immune signaling, discussing the implications for therapeutic development in cancer and neurodegenerative diseases.
The Biochemical Mechanism of PARP-1 Cleavage: From DNA Repair to Apoptosis Signaling
Poly (ADP-ribose) polymerase 1 (PARP1) is a crucial nuclear enzyme that serves as a primary sensor of DNA damage and plays a fundamental role in maintaining genomic integrity. As the most extensively studied member of the PARP superfamily comprising 17 proteins, PARP1 becomes activated upon binding to DNA single-strand and double-strand breaks, initiating various DNA repair pathways [1] [2]. Through its catalytic function of adding poly (ADP-ribose) (PAR) chains to target proteins, PARP1 facilitates the recruitment of DNA repair machinery to damage sites and modifies chromatin structure to enhance DNA accessibility [1]. The critical importance of PARP1 in cellular homeostasis is evidenced by its frequent upregulation in cancer and the hypersensitivity of PARP1-deficient organisms to DNA-damaging agents [1]. Beyond its established roles in DNA repair, PARP1 also participates in transcriptional regulation, and its cleavage during apoptosis serves as a definitive marker for programmed cell death [3] [4]. This technical review examines the structural features of PARP1, its multifaceted functions in DNA damage response, and the significance of its proteolytic cleavage in cell death pathways, providing a comprehensive resource for researchers and drug development professionals.
PARP-1 Domain Architecture and Structural Features
PARP1 is a 113-kDa nuclear protein consisting of multiple structured domains that confer its DNA-binding and catalytic capabilities [1] [2]. The domain organization follows a modular architecture that facilitates its transition from an autoinhibited state to an activated DNA repair enzyme.
Table 1: PARP-1 Structural Domains and Their Functions
| Domain | Position | Size | Key Functions | Structural Features |
|---|---|---|---|---|
| Zinc Finger 1 (Zn1) | N-terminal | ~ | Primary DNA break sensor | Binds DNA strand breaks, cooperates with Zn2 |
| Zinc Finger 2 (Zn2) | N-terminal | ~ | DNA break binding | Recognizes DNA damage, forms cooperative interface with Zn1 |
| Zinc Finger 3 (Zn3) | N-terminal | ~ | Inter-domain communication | Facilitates conformational changes for activation |
| BRCT (Auto-modification Domain) | Central | 22-kDa | Protein-protein interactions | Target for auto-ADP-ribosylation, contains BRCT fold |
| WGR Domain | Central | ~ | DNA binding & allosteric regulation | Bridges DNA binding to catalytic activation |
| Catalytic Domain (CAT) | C-terminal | 54-kDa | ADP-ribose polymerization | Comprises helical subdomain (HD) & ART subdomain |
The N-terminal region of PARP1 contains three zinc-binding domains (Zn1, Zn2, and Zn3) that are responsible for recognizing DNA lesions [2]. Structural studies have revealed that Zn1 and Zn2 domains cooperate to recognize damaged DNA, with both domains contacting the DNA phosphate backbone through specific residues [2]. The Zn3 domain plays a crucial role in inter-domain interactions and is essential for PARP1 enzymatic activity [3].
The central region of PARP1 contains the BRCA1 C-terminus (BRCT) fold within the auto-modification domain (AMD), which mediates protein-protein interactions and serves as the primary target for auto-ADP-ribosylation [3] [2]. Adjacent to this domain is the tryptophan-glycine-arginine (WGR) motif, which contributes to DNA binding and participates in the allosteric regulation of PARP1 catalytic activity [2].
The C-terminal region houses the catalytic (CAT) domain, which is the most conserved region across the PARP family [2]. This domain comprises the helical subdomain (HD) and the ADP-ribosyl transferase (ART) subdomain, which contains the NAD+-binding pocket responsible for poly(ADP-ribose) synthesis [2]. In the absence of DNA damage, PARP1 maintains a autoinhibited conformation, but binding to DNA lesions induces substantial conformational changes that activate the catalytic domain [1] [2].
DNA Damage Recognition and Repair Mechanisms
PARP1 functions as a critical molecular sensor for DNA damage, with particular specificity for single-strand breaks (SSBs) that arise from various genotoxic insults [2]. The enzyme rapidly recognizes DNA lesions through its zinc finger domains, which exhibit high affinity for both single-strand and double-strand DNA breaks [1] [2]. Upon DNA binding, PARP1 undergoes conformational changes that activate its catalytic function, leading to the initiation of poly(ADP-ribosyl)ation (PARylation) using NAD+ as a substrate [1].
The PARylation activity of PARP1 serves multiple essential functions in DNA damage response. PARP1 initially modifies itself through automodification, adding extensive branched polymers of ADP-ribose to glutamate residues in its auto-modification domain [1]. This automodification creates a platform for recruiting DNA repair proteins such as XRCC1, which acts as a scaffold for additional repair factors [1] [2]. The negative charges conferred by PAR chains also promote chromatin relaxation by repelling histones from DNA, thereby increasing accessibility for repair machinery [1].
PARP1 participates in several DNA repair pathways through distinct mechanisms:
- Base Excision Repair (BER): PARP1 recognizes SSBs resulting from base modifications and recruits XRCC1 and other core factors to process the repair [5].
- Double-Strand Break (DSB) Repair: PARP1 contributes to both homologous recombination (HR) and non-homologous end joining (NHEJ) pathways by regulating the recruitment of proteins such as MRE11, NBS1, BRCA1, and DNA-PKcs to damage sites [5].
- Alternative NHEJ: In the absence of Ku70/Ku80, PARP1 promotes error-prone microhomology-mediated end joining [5].
- DNA Replication: PARP1 acts as a sensor for unligated Okazaki fragments during lagging-strand synthesis and helps control replication fork velocity [5].
The following diagram illustrates PARP1's central role in coordinating the DNA damage response through multiple pathways:
Beyond its catalytic functions, PARP1 also exhibits enzyme-independent activities in gene transcription through recognition of specific DNA motifs. PARP1 binds the octamer sequence "RNNWCAAA" found in various gene promoters, which suppresses its ADP-ribosylation activity while enabling transcriptional regulation [1]. This dual functionality positions PARP1 as a critical integrator of genomic maintenance and gene expression programs.
PARP-1 Cleavage as an Apoptosis Marker
The proteolytic cleavage of PARP1 serves as a biochemical hallmark of apoptosis and has been extensively utilized as a definitive marker for programmed cell death in research and diagnostic contexts [3] [4]. During apoptosis, PARP1 becomes a primary substrate for activated caspases, resulting in characteristic fragments that provide signatures of specific protease activities [3].
Caspase-Mediated Cleavage
In caspase-dependent apoptosis, executioner caspases-3 and -7 recognize and cleave PARP1 at a specific aspartic acid residue (DEVD214↓G) located between the second zinc finger domain and the auto-modification domain [3] [6]. This proteolytic event generates two definitive fragments: a 24-kDa N-terminal fragment containing the DNA-binding domain and nuclear localization signal, and an 89-kDa C-terminal fragment comprising the auto-modification and catalytic domains [3] [6].
The biological consequences of this cleavage are significant for the apoptotic process. The 24-kDa fragment retains the DNA-binding capability and remains tightly associated with DNA strand breaks, where it acts as a trans-dominant inhibitor of intact PARP1 by blocking access to DNA damage sites [3]. This irreversible binding prevents DNA repair during apoptosis, thereby facilitating the systematic dismantling of the cell [3] [6]. Meanwhile, the 89-kDA fragment, which contains the catalytic domain, translocates from the nucleus to the cytoplasm [6].
Table 2: PARP-1 Cleavage Fragments and Their Properties
| Fragment | Size | Domains Contained | Localization After Cleavage | Functions |
|---|---|---|---|---|
| N-terminal Fragment | 24-kDa | Zinc fingers 1 & 2, NLS | Nuclear | Binds DNA irreversibly, dominant-negative inhibitor of PARP1 |
| C-terminal Fragment | 89-kDa | Zinc finger 3, BRCT, WGR, CAT | Cytosolic | Serves as PAR carrier, novel signaling functions |
Cleavage by Other Proteases
Beyond caspases, PARP1 serves as a substrate for additional proteases that are activated in alternative cell death pathways. Calpain, a calcium-activated protease, cleaves PARP1 during excitotoxicity and other pathological conditions, generating distinct fragments of 55-kDa and 62-kDa [3]. Cathepsins, which are lysosomal proteases released during autophagic cell death, also process PARP1 into specific cleavage products [1] [3]. Additionally, granzyme B from cytotoxic lymphocytes and certain matrix metalloproteinases (MMPs) can target PARP1, creating unique proteolytic signatures that serve as biomarkers for specific cell death programs in various pathological contexts [3].
The following diagram illustrates the proteolytic cleavage of PARP1 and the fate of its fragments during apoptosis:
Functional Consequences of PARP-1 Cleavage
The cleavage of PARP1 during apoptosis serves multiple critical functions in the orderly progression of programmed cell death. The 24-kDa fragment's irreversible binding to DNA breaks effectively terminates DNA repair processes, conserving cellular energy that would otherwise be expended on nuclear repair and facilitating nuclear dismantlement [3]. This represents a strategic cellular decision to abandon genomic maintenance in favor of programmed elimination.
Recent research has revealed that the 89-kDa C-terminal fragment possesses novel biological activities beyond simply inactivating PARP1. This fragment can serve as a carrier for poly(ADP-ribose) (PAR) polymers, transporting them from the nucleus to the cytoplasm during apoptosis [6]. In the cytoplasm, PAR polymers attached to the 89-kDa fragment bind to apoptosis-inducing factor (AIF), facilitating its release from mitochondria and subsequent translocation to the nucleus, where it promotes large-scale DNA fragmentation [6]. This mechanism establishes a crucial link between caspase activation and AIF-mediated DNA degradation in apoptosis.
Furthermore, the 89-kDa fragment has been shown to interact with the RNA polymerase III (Pol III) complex in the cytoplasm during innate immune responses [4]. This interaction facilitates the ADP-ribosylation of Pol III, enhancing interferon-β production and promoting apoptosis in response to cytosolic DNA, such as that from pathogenic infections [4]. This newly discovered function positions truncated PARP1 as a participant in antimicrobial defense mechanisms.
The cleavage of PARP1 also represents a strategic intervention to prevent energy depletion. Unchecked PARP1 activation consumes substantial amounts of NAD+, leading to ATP depletion and necrotic cell death [1] [5]. By inactivating PARP1, caspase-mediated cleavage maintains energy homeostasis necessary for the controlled execution of apoptosis, distinguishing it from other forms of cell death.
Experimental Analysis of PARP-1 Cleavage
Detection Methodologies
The analysis of PARP1 cleavage employs specific methodological approaches that enable researchers to identify and quantify apoptosis in experimental systems. Western blotting represents the most widely utilized technique, with antibodies targeting different PARP1 epitopes to distinguish full-length (116-kDa) from cleaved (89-kDa) fragments [6] [4]. The appearance of the 89-kDa fragment serves as a definitive indicator of caspase activation and apoptosis.
Immunofluorescence microscopy allows spatial resolution of PARP1 cleavage events, enabling researchers to visualize the translocation of the 89-kDa fragment from the nucleus to the cytoplasm and monitor AIF release from mitochondria [6]. This technique provides subcellular contextual information about the progression of apoptotic signaling.
Flow cytometry combined with Annexin V/propidium iodide staining frequently correlates with PARP1 cleavage analysis to confirm apoptosis and distinguish it from other forms of cell death [4]. This multiparametric approach strengthens experimental conclusions regarding cell death mechanisms.
Research Reagent Solutions
Table 3: Essential Research Reagents for PARP-1 Cleavage Studies
| Reagent/Category | Specific Examples | Function/Application | Experimental Notes |
|---|---|---|---|
| PARP Inhibitors | PJ34, ABT-888 (Veliparib) | Inhibit PARP catalytic activity | Control for PARP1-dependent cell death mechanisms |
| Caspase Inhibitors | zVAD-fmk | Pan-caspase inhibitor | Suppresses PARP1 cleavage; confirms caspase-dependence |
| Apoptosis Inducers | Staurosporine, Actinomycin D, Etoposide | Activate caspase cascade | Generate PARP1 cleavage fragments for study |
| Antibodies (Full-length PARP1) | Various commercial clones | Detect intact 116-kDa PARP1 | Baseline marker before cleavage |
| Antibodies (Cleaved PARP1) | Anti-89-kDa fragment specific | Identify apoptotic cells | Must distinguish from full-length PARP1 |
| PAR Polymer Detection | PAR-specific antibodies | Measure PARP1 activation | Indicator of PARP1 activity before cleavage |
| Cell Lines | PARP1-deficient 293T, HeLa | Genetic PARP1 manipulation | Control for PARP1-specific effects |
Experimental Protocols
For inducing and analyzing PARP1 cleavage, researchers typically treat cells with apoptosis inducers such as staurosporine (0.5-2 μM) or actinomycin D (0.5-5 μM) for 4-24 hours, depending on cell type and sensitivity [6] [4]. Following treatment, cells are lysed using RIPA buffer supplemented with protease inhibitors to prevent post-lysis degradation, and proteins are separated by SDS-PAGE (8-10% gels) for optimal resolution of full-length and cleaved PARP1 fragments [6].
For studies examining the innate immune response connection, transfection with poly(dA-dT) (0.5-2 μg/mL) effectively mimics pathogenic DNA and stimulates the caspase-mediated PARP1 cleavage pathway connected to RNA polymerase III activation [4]. This approach is particularly useful for investigating the non-canonical functions of truncated PARP1 in cytoplasmic signaling.
To differentiate parthanatos from apoptosis, researchers employ pharmacological inhibitors including PARP inhibitors (PJ34 at 10-50 μM) to block PARP1 catalytic activity, and caspase inhibitors (zVAD-fmk at 20-50 μM) to suppress caspase-mediated cleavage [6]. The distinct inhibitor profiles help characterize the specific cell death pathway operational in experimental models.
PARP1 represents a critical node in cellular fate decisions, balancing its functions in DNA repair against its role as a marker and mediator of programmed cell death. The structural organization of PARP1 enables its dual functionality as a DNA damage sensor and a signal transducer through poly(ADP-ribosyl)ation. The characteristic cleavage of PARP1 during apoptosis serves as a definitive biochemical marker that not only inactivates DNA repair capacity but also generates fragments with novel signaling functions, particularly in connecting caspase activation to mitochondrial-mediated DNA degradation. Ongoing research continues to elucidate the complex roles of PARP1 fragments in cellular processes beyond apoptosis, including innate immunity and transcriptional regulation. For research and drug development professionals, understanding PARP1 structure, function, and cleavage mechanisms provides valuable insights for developing targeted therapeutic strategies in cancer and other diseases characterized by dysregulated cell death pathways.
Poly(ADP-ribose) polymerase-1 (PARP-1) is a nuclear enzyme that functions as a primary sensor of DNA damage, playing crucial roles in DNA repair and maintenance of genomic integrity [7] [8]. However, during apoptosis, PARP-1 becomes a primary target for proteolytic cleavage by caspases, a process now recognized as a biochemical hallmark of programmed cell death [3]. This cleavage event serves as a critical molecular switch that fundamentally alters the cellular response to damage, redirecting energy from DNA repair to the systematic dismantling of the cell. The discovery that PARP-1 is cleaved by caspase-3 during apoptosis has established it as one of the most reliable biomarkers for distinguishing apoptosis from other forms of cell death [3]. For researchers and drug development professionals, understanding this mechanism provides not only fundamental insights into cell fate decisions but also potential therapeutic strategies for cancer, neurodegenerative disorders, and other conditions where apoptotic regulation is disrupted.
Molecular Mechanism of PARP-1 Cleavage
Caspase Recognition and Cleavage Site
PARP-1 is a modular protein consisting of multiple functional domains: three zinc finger motifs (Zn1, Zn2, Zn3) at the N-terminus responsible for DNA damage recognition, a nuclear localization signal (NLS), a BRCT domain involved in protein-protein interactions, and a C-terminal catalytic domain (CAT) that contains the NAD+ binding site [9]. During apoptosis, activated executioner caspases, primarily caspase-3 and caspase-7, recognize and cleave PARP-1 at a highly specific amino acid sequence located between the second and third zinc finger domains [3].
The cleavage occurs specifically at the DEVD214↓G215 motif, situated within the nuclear localization signal between the second zinc finger and the BRCT domain [3] [10]. This proteolytic event generates two characteristic fragments: a 24-kDa N-terminal fragment containing the first two zinc finger DNA-binding domains, and an 89-kDa C-terminal fragment containing the third zinc finger, BRCT domain, WGR domain, and the catalytic domain [3] [10]. The precise cleavage at this location effectively separates the DNA-binding capability from the catalytic activity of PARP-1, resulting in its functional inactivation.
Table 1: PARP-1 Domains and Cleavage Fragments
| Domain/Feature | Location | Function | Status After Cleavage |
|---|---|---|---|
| Zn1 & Zn2 (Zinc Fingers) | N-terminal (AA 1-214) | DNA damage recognition | 24-kDa fragment; remains nuclear |
| Nuclear Localization Signal | AA ~200-214 | Nuclear targeting | Cleaved; contains caspase site |
| Zn3 (Third Zinc Finger) | AA ~370-486 | DNA binding stabilization | 89-kDa fragment |
| BRCT Domain | AA ~387-486 | Protein-protein interactions | 89-kDa fragment |
| WGR Domain | AA ~487-586 | DNA binding & catalysis | 89-kDa fragment |
| Catalytic Domain | C-terminal (AA 587-1014) | PAR synthesis activity | 89-kDa fragment; reduced DNA binding |
Structural Consequences of Cleavage
The cleavage of PARP-1 at Asp214 produces dramatic structural and functional consequences. The 24-kDa fragment retains the strong DNA-binding affinity through its zinc finger motifs but lacks catalytic activity [3]. This fragment remains tightly bound to DNA strand breaks in the nucleus, where it functions as a trans-dominant inhibitor of intact PARP-1 and other DNA repair enzymes by physically blocking their access to DNA damage sites [3].
The 89-kDa fragment, while containing most of the PARP-1 functional domains, exhibits significantly reduced DNA binding capacity due to the loss of the first two zinc fingers [3] [10]. Importantly, this fragment loses its nuclear localization signal and consequently translocates to the cytoplasm during apoptosis [10]. Recent research has revealed that this cytoplasmic translocation is not merely a consequence of inactivation but may serve additional biological functions in amplifying cell death signaling [4] [10].
Functional Consequences: From DNA Repair to Cell Death Execution
The Energy Conservation Hypothesis
The traditional understanding of PARP-1 cleavage centers on the energy conservation hypothesis. PARP-1 activation in response to DNA damage consumes substantial amounts of NAD+, and subsequently ATP, as the cell attempts to resynthesize NAD+ to maintain this repair activity [11]. In cells undergoing severe DNA damage, persistent PARP-1 activation can lead to catastrophic ATP depletion, potentially shifting the mode of cell death from controlled apoptosis to uncontrolled necrosis [11].
By cleaving and inactivating PARP-1, caspases prevent this excessive energy consumption, thereby conserving cellular ATP pools necessary for the orderly execution of apoptosis [11] [3]. This concept is supported by research showing that cells with non-cleavable PARP-1 mutants are more sensitive to necrosis under conditions of death receptor activation [11]. The cleavage thus represents a strategic cellular decision to abandon repair efforts in favor of programmed self-destruction when damage is irreparable.
Novel Biological Functions of Cleavage Fragments
Recent evidence suggests that PARP-1 cleavage fragments may possess biological activities beyond the mere inactivation of DNA repair:
The 89-kDa fragment as a cytoplasmic PAR carrier: Research has demonstrated that the 89-kDa fragment can be poly(ADP-ribosyl)ated before or during cleavage and subsequently translocates to the cytoplasm, where it may serve as a carrier of PAR polymers [10]. This PAR-bound fragment can facilitate the release of apoptosis-inducing factor (AIF) from mitochondria, potentially amplifying the cell death signal through parthanatos, a PAR-dependent cell death pathway [10].
Cytosolic signaling functions: The truncated 89-kDa PARP-1 has been shown to interact with cytoplasmic proteins, including components of the RNA polymerase III (Pol III) complex [4]. This interaction enables ADP-ribosylation of Pol III, which facilitates interferon-β production and enhances apoptosis during innate immune responses to foreign DNA [4].
Table 2: Functional Consequences of PARP-1 Cleavage
| Functional Aspect | Consequence | Biological Significance |
|---|---|---|
| DNA Repair | Inhibition of base excision and single-strand break repair | Prevents resource allocation to futile repair |
| Energy Metabolism | Conservation of NAD+ and ATP pools | Enables energy-dependent apoptotic execution |
| Cell Fate Decision | Switch from necrotic to apoptotic death | Promotes controlled cell removal without inflammation |
| Immune Signaling | Potential enhancement of interferon response | Links apoptosis to immune activation |
| Mitochondrial Signaling | Possible facilitation of AIF release | Amplifies cell death through multiple pathways |
Experimental Analysis of PARP-1 Cleavage
Detection Methodologies
The detection of PARP-1 cleavage serves as a fundamental assay in apoptosis research, with several well-established methodological approaches:
Western Blot Analysis remains the gold standard for detecting PARP-1 cleavage. This technique typically uses antibodies that recognize either the full-length protein (116 kDa) or specific cleavage fragments (89 kDa and 24 kDa) [3]. The appearance of the 89-kDa fragment with simultaneous disappearance of the 116-kDa full-length protein provides definitive evidence of caspase-mediated cleavage [3]. For increased specificity, some protocols utilize antibodies that specifically recognize the neo-epitopes created by caspase cleavage.
Immunofluorescence Microscopy enables the spatial visualization of PARP-1 cleavage and fragment localization. Following cleavage, the 89-kDa fragment translocates to the cytoplasm, which can be detected using antibodies against the C-terminal portion of PARP-1, while the 24-kDa fragment remains nuclear [10]. This subcellular redistribution provides additional confirmation of cleavage beyond molecular weight changes.
PARP-1 Cleavage-Specific Assays include commercial kits that specifically detect the caspase-cleaved form of PARP-1, such as the PARP in vivo Apoptosis Assay, which offers higher specificity for apoptotic cells compared to general PARP-1 antibodies [12].
Research Reagent Solutions
Table 3: Essential Research Reagents for PARP-1 Cleavage Studies
| Reagent/Category | Specific Examples | Research Application | Key Features |
|---|---|---|---|
| PARP-1 Antibodies | Anti-PARP-1 (full length), Anti-cleaved PARP-1 (89 kDa), Anti-PARP-1 p24 (24 kDa) | Western blot, Immunofluorescence, IHC | Detection of specific fragments; cleavage-specific antibodies recognize neo-epitopes |
| Caspase Inhibitors | zVAD-fmk (pan-caspase), DEVD-CHO (caspase-3/7 specific) | Inhibition studies to confirm caspase-dependence | Reversible and irreversible inhibitors available |
| Apoptosis Inducers | Staurosporine, Actinomycin D, Anti-CD95, TNF-α with sensitizers | Positive controls for cleavage induction | Activate intrinsic or extrinsic apoptosis pathways |
| PARP Activity Assays | PAR ELISA, NAD+ consumption assays | Correlation of cleavage with functional inactivation | Measure loss of enzymatic activity post-cleavage |
| Cell Death Detection Kits | Annexin V/propidium iodide, TUNEL assay, Caspase-3 activity | Multiparametric cell death analysis | Confirm apoptosis alongside PARP-1 cleavage |
| PARP Inhibitors | 3-AB, PJ-34, Olaparib (clinical) | Investigation of PARP-1 function in cell death | Tool compounds and clinical agents |
Protocol for PARP-1 Cleavage Detection by Western Blot
Sample Preparation:
- Harvest cells after apoptotic stimulation and lyse in RIPA buffer supplemented with protease inhibitors.
- Quantify protein concentration using BCA assay and adjust samples to equal concentrations.
- Prepare Laemmli buffer with β-mercaptoethanol and denature at 95°C for 5 minutes.
Electrophoresis and Blotting:
- Load 20-50 μg protein per lane on 4-12% Bis-Tris gradient gels.
- Separate proteins by SDS-PAGE at 120-150V for 1-2 hours.
- Transfer to PVDF membrane using standard wet or semi-dry transfer systems.
Immunodetection:
- Block membrane with 5% non-fat milk in TBST for 1 hour.
- Incubate with primary antibody (anti-PARP-1, preferably detecting both full-length and cleaved fragments) diluted in blocking buffer overnight at 4°C.
- Wash membrane 3× with TBST for 10 minutes each.
- Incubate with HRP-conjugated secondary antibody for 1 hour at room temperature.
- Develop using enhanced chemiluminescence substrate and image.
Interpretation:
- Apoptotic samples will show the 89-kDa cleavage fragment with corresponding decrease in the 116-kDa full-length PARP-1.
- Advanced apoptosis may show complete disappearance of full-length PARP-1 with strong 89-kDa band.
- The 24-kDa fragment is less frequently detected due to poor transfer or antibody accessibility issues.
PARP-1 Cleavage in Pathophysiology and Therapeutics
Role in Disease Pathogenesis
PARP-1 cleavage serves as a significant biomarker and functional component in various pathological conditions:
Neurodegenerative Diseases: In conditions such as Alzheimer's disease, Parkinson's disease, and amyotrophic lateral sclerosis, PARP-1 cleavage is detected in vulnerable neuronal populations, indicating apoptotic activation [13] [3]. However, in these chronic conditions, the balance between PARP-1 activation and cleavage may be disrupted, contributing to disease progression through both apoptosis and parthanatos [13].
Cancer Biology: PARP-1 cleavage is routinely used as a biomarker for assessing the efficacy of chemotherapeutic agents that induce apoptosis [8]. Many anticancer drugs, including DNA-damaging agents and targeted therapies, trigger PARP-1 cleavage as part of their mechanism of action [14] [8]. Interestingly, resistance to therapy may manifest as impaired PARP-1 cleavage despite caspase activation, suggesting alternative regulatory mechanisms.
Liver Diseases: In various liver pathologies including viral hepatitis, alcoholic liver disease, and drug-induced liver injury, PARP-1 cleavage occurs during hepatocyte apoptosis [7]. The extent of cleavage correlates with disease severity and progression to fibrosis, positioning it as a potential biomarker for liver injury staging.
Therapeutic Implications and Drug Development
The central role of PARP-1 cleavage in cell fate decisions has made it a valuable readout in pharmaceutical development:
PARP Inhibitors in Cancer Therapy: PARP inhibitors (PARPi) such as olaparib, niraparib, and rucaparib have shown clinical success in BRCA-mutated cancers [8]. These inhibitors not only compromise DNA repair in HR-deficient cells but also trap PARP-1 on DNA, potentially enhancing apoptotic signaling and PARP-1 cleavage in response to replication stress [8].
Cell Death Modulators: Compounds that influence the balance between PARP-1 activation and cleavage are being explored for conditions where regulated cell death is desirable (cancer) or detrimental (neurodegeneration) [13]. For instance, caspase-resistant PARP-1 mutants have been used to study the functional consequences of preventing cleavage, demonstrating increased sensitivity to necrotic cell death under certain conditions [11].
Combination Therapies: Monitoring PARP-1 cleavage provides a pharmacodynamic biomarker for assessing the efficacy of combination therapies that target DNA repair pathways while inducing apoptosis [12] [8]. This application is particularly valuable in early-phase clinical trials for establishing proof-of-mechanism.
Caspase-mediated cleavage of PARP-1 represents a critical commitment step in the apoptotic pathway, serving as a definitive molecular switch that inactivates the DNA repair machinery and redirects cellular resources toward programmed cell death. The characteristic 89-kDa fragment generated through this process not only serves as a reliable biomarker for apoptosis but may also actively participate in cytoplasmic signaling events that amplify cell death. For researchers and drug development professionals, the detection and quantification of PARP-1 cleavage remains an essential tool for investigating cell death mechanisms, screening therapeutic compounds, and understanding disease pathogenesis across diverse conditions from cancer to neurodegeneration. As research continues to unveil novel functions for PARP-1 fragments beyond their traditional roles, this apoptotic switch continues to offer new insights into the sophisticated mechanisms governing cellular life and death decisions.
Poly(ADP-ribose) polymerase-1 (PARP-1) is a 116-kDa nuclear enzyme that functions as a primary DNA damage sensor. During the execution phase of apoptosis, caspase-3 and -7 cleave PARP-1 into signature fragments of 24-kDa and 89-kDa. This proteolytic event serves as a critical biochemical marker of apoptosis and represents a strategic functional shift from DNA repair to cellular dismantling. The 24-kDa fragment contains the DNA-binding domain, while the 89-kDa fragment retains the catalytic domain. This whitepaper provides a comprehensive technical analysis of these fragments, detailing their structural properties, functional consequences, and methodological approaches for their study in apoptotic research and drug development.
PARP-1 is a highly abundant nuclear protein that plays a dual role in cellular fate decisions. In response to mild DNA damage, it facilitates DNA repair processes, but during extensive damage, it becomes a marker for programmed cell death. The cleavage of PARP-1 by executioner caspases is considered one of the most characteristic biochemical hallmarks of apoptosis [15]. This proteolytic event serves as a definitive indicator that the cell has passed the point of recovery and has committed to apoptotic death. The discovery that PARP-1 is cleaved during apoptosis dates back to the early 1990s, and since then, its detection has become a gold standard for confirming apoptotic pathways in experimental and clinical contexts [15]. The functional consequence of this cleavage is the irreversible inactivation of PARP-1's role in DNA repair, which prevents futile repair attempts and facilitates the systematic dismantling of the cell.
Structural Domains of PARP-1 and Cleavage Sites
Domain Architecture of Full-Length PARP-1
PARP-1 is a modular protein comprising several functional domains:
- DNA-binding domain (DBD): Located at the N-terminus, this 46-kDa region contains three zinc-finger motifs (F1, F2, and F3) that recognize DNA strand breaks [16] [17]. The first two zinc fingers (F1 and F2) are structurally independent and share a highly similar fold, with F2 playing the predominant role in DNA damage recognition [16].
- Automodification domain (AMD): This central 22-kDa region contains a BRCT (BRCA1 C-terminal) motif and serves as a regulatory segment with multiple glutamic acid residues that function as acceptor sites for poly(ADP-ribose) chains [17].
- Catalytic domain: The C-terminal 54-kDa region executes the enzymatic function of PARP-1, synthesizing poly(ADP-ribose) polymers using NAD+ as a substrate [17].
The caspase cleavage site resides between the DNA-binding domain and the automodification domain, specifically in a nuclear localization signal (NLS) region [6].
Caspase Cleavage and Fragment Generation
During apoptosis, activated caspase-3 and caspase-7 cleave PARP-1 at a specific DEVD motif (amino acid sequence 214-GDEVD-218 in human PARP-1), generating two primary fragments [6] [15]:
Table 1: PARP-1 Cleavage Fragments
| Fragment | Molecular Weight | Domains Contained | Key Functions |
|---|---|---|---|
| 24-kDa N-terminal | 24 kDa | DNA-binding domain (zinc fingers F1 & F2) | DNA break binding, dominant-negative inhibition of DNA repair |
| 89-kDa C-terminal | 89 kDa | Automodification domain, Catalytic domain | Limited catalytic activity, PAR carrier function |
This cleavage event is a reliable indicator of caspase activation and apoptosis commitment. The 24-kDa fragment corresponds to the apoptotic fragment released through cleavage by caspase-3 and -7 and contains the DNA-damage-binding activity [16]. It's important to note that during necrosis, PARP-1 is processed differently, generating a distinct 50-kDa fragment through lysosomal protease activity (e.g., cathepsins B and G) rather than caspase-mediated cleavage [18].
Functional Consequences of PARP-1 Cleavage
The 24-kDa DNA-Binding Fragment: From Repair to Inhibition
The 24-kDa fragment retains the DNA-binding domain of PARP-1 but lacks catalytic activity. This fragment demonstrates several critical functions in apoptosis:
Dominant-negative inhibition of DNA repair: The 24-kDa fragment competes with full-length PARP-1 and other DNA repair proteins for binding to DNA strand breaks, thereby inhibiting DNA repair processes [19]. In cell-free DNA repair assays, this fragment effectively inhibits rejoining of DNA breaks and suppresses ADP-ribose polymer formation [19].
Transcription modulation: The 24-kDa fragment can bind to RNA and compete against the up-regulation of transcription mediated by full-length PARP-1, potentially contributing to the shutdown of cellular functions during apoptosis [19].
Irreversible DNA binding: Unlike full-length PARP-1, which undergoes automodification and dissociates from DNA, the 24-kDa fragment remains tightly bound to DNA breaks due to its lack of the automodification domain [6].
The 89-kDa Catalytic Fragment: Beyond Catalytic Inactivation
While the 89-kDa fragment retains the catalytic domain, its activity is significantly reduced compared to full-length PARP-1. However, recent research has revealed unexpected functions for this fragment:
PAR carrier function: Following caspase cleavage, the 89-kDa fragments with covalently attached PAR polymers can be translocated from the nucleus to the cytoplasm [6]. This translocation represents a crucial mechanism for communicating nuclear DNA damage to cytoplasmic compartments.
Induction of parthanatos: The 89-kDa fragment serves as a vehicle for transporting PAR to the cytoplasm, where it facilitates apoptosis-inducing factor (AIF) release from mitochondria [6]. AIF then translocates to the nucleus and induces large-scale DNA fragmentation, contributing to the parthanatos cell death pathway.
Caspase-amplification loop: In certain contexts, such as RSL3-induced ferroptosis-apoptosis crosstalk, the 89-kDa fragment directly induces caspase-mediated DNA fragmentation, creating an amplification loop that enhances apoptotic signaling [20].
Detection Methods and Experimental Approaches
Western Blot Analysis
The most common method for detecting PARP-1 cleavage is Western blotting using antibodies that recognize different PARP-1 epitopes:
- Antibody selection: Use antibodies against the N-terminal region to detect the 24-kDa fragment and antibodies against the C-terminal region to detect the 89-kDa fragment and full-length PARP-1.
- Sample preparation: Include appropriate controls (e.g., caspase inhibitors, PARP inhibitors) to confirm the specificity of cleavage.
- Interpretation: The appearance of the 89-kDa fragment and corresponding decrease in full-length PARP-1 (116-kDa) indicates caspase activation and apoptosis.
Activity-Based Assays
- PAR synthesis assays: Measure the reduction in PAR formation in apoptotic cells compared to healthy cells.
- DNA binding assays: Electrophoretic mobility shift assays (EMSA) can demonstrate the DNA-binding capability of the 24-kDa fragment [16].
- Cell-free systems: In vitro translation systems combined with caspase treatment can validate direct cleavage events.
Table 2: Quantitative Parameters of PARP-1 Cleavage
| Parameter | Full-Length PARP-1 | 24-kDa Fragment | 89-kDa Fragment |
|---|---|---|---|
| Molecular Weight | 116 kDa | 24 kDa | 89 kDa |
| DNA Binding Affinity | High (Kd ~nM) | High (Kd ~nM) [19] | None |
| Catalytic Activity | Full activity | None | Reduced activity |
| Cellular Localization | Nuclear | Nuclear | Nuclear/Cytoplasmic [6] |
| Half-life | Hours | Stable during apoptosis | Degraded during late apoptosis |
Immunofluorescence and Cellular Localization
- Subcellular localization: The 24-kDa fragment remains nuclear, while the 89-kDa fragment can translocate to the cytoplasm [6].
- TUNEL co-staining: Combine PARP cleavage detection with TUNEL assay to correlate cleavage with DNA fragmentation.
- Live-cell imaging: Use FRET-based PARP-1 biosensors to monitor real-time cleavage during apoptosis.
Technical Diagrams
PARP-1 Cleavage and Apoptotic Signaling Pathway
Diagram Title: PARP-1 Cleavage in Apoptotic Signaling
Domain Architecture and Cleavage Site
Diagram Title: PARP-1 Domain Structure and Cleavage
Research Reagent Solutions
The following table provides essential research tools for studying PARP-1 cleavage in apoptotic research:
Table 3: Essential Research Reagents for PARP-1 Cleavage Studies
| Reagent/Catalog Number | Supplier | Application | Key Features/Experimental Notes |
|---|---|---|---|
| Caspase-3 (Active) | Multiple suppliers | In vitro cleavage assays | Verify cleavage specificity using caspase inhibitors |
| Anti-PARP-1 Antibody (N-terminal) | Multiple suppliers | Western blot, immunofluorescence | Detects 24-kDa fragment and full-length PARP-1 |
| Anti-PARP-1 Antibody (C-terminal) | Multiple suppliers | Western blot, immunofluorescence | Detects 89-kDa fragment and full-length PARP-1 |
| zVAD-fmk | MedChemExpress, etc. | Caspase inhibition control | Broad-spectrum caspase inhibitor; validates caspase-dependent cleavage |
| PJ34 / ABT888 | MedChemExpress, etc. | PARP inhibition studies | PARP-1 specific inhibitors; useful for functional studies |
| Recombinant 24-kDa PARP-1 | In-house preparation | DNA repair competition assays | Retains DNA binding without catalytic activity [19] |
| Staurosporine | Multiple suppliers | Apoptosis induction | Conventional inducer of caspase-mediated PARP-1 cleavage [6] |
| RSL3 | MedChemExpress | Ferroptosis-apoptosis studies | Induces PARP-1 cleavage via ROS and caspase-3 activation [20] |
Implications for Drug Development and Therapeutic Targeting
The understanding of PARP-1 cleavage fragments has significant implications for pharmaceutical development:
PARP inhibitor resistance: RSL3, a ferroptosis inducer, promotes PARP-1 apoptotic functions through both caspase-dependent cleavage and reduced full-length PARP-1 via inhibition of METTL3-mediated m6A modification, demonstrating therapeutic potential against PARP inhibitor-resistant malignancies [20].
Radiation therapy applications: STING directly interacts with PAR produced by activated PARP-1 upon ionizing radiation, promoting apoptosis. Inhibiting PARP1 reduces cell apoptosis after radiation exposure, suggesting combinatorial approaches for radiation therapy [21].
Cell death pathway modulation: The 89-kDa fragment's role as a PAR carrier linking nuclear DNA damage to cytoplasmic AIF release provides new therapeutic targets for conditions involving parthanatos, such as neurodegenerative diseases [6].
The signature 24-kDa and 89-kDa cleavage fragments of PARP-1 represent more than just apoptotic biomarkers—they embody a fundamental switch in cellular fate from repair to death. The 24-kDa fragment acts as a dominant-negative inhibitor of DNA repair, while the 89-kDa fragment facilitates the spatial propagation of death signals through PAR translocation. Understanding the structural basis, functional consequences, and detection methodologies for these fragments provides critical insights for basic apoptosis research and the development of novel therapeutic strategies targeting cell death pathways in cancer, neurodegenerative disorders, and other pathological conditions.
Poly(ADP-ribose) polymerase-1 (PARP-1) is a critical nuclear enzyme involved in DNA damage repair and cell death signaling. Its cleavage by executioner caspases during apoptosis represents a definitive biochemical event that serves dual purposes: terminating DNA repair capacity to ensure irreversible commitment to cell death and conserving cellular energy to facilitate the apoptotic process. This whitepaper examines the molecular consequences of PARP-1 cleavage, detailing how this event functions as a molecular switch between cell death modalities and serves as a reliable apoptosis marker in research and drug development contexts. The technical guide provides comprehensive experimental methodologies, quantitative data comparisons, and visualization tools to support ongoing research into PARP-1's role in cellular fate decisions.
PARP-1 is a multifunctional nuclear enzyme composed of 1,014 amino acids with three primary domains: the N-terminal DNA-binding domain (DBD), the central automodification domain (AMD), and the C-terminal catalytic domain (CAT) [9]. The DBD contains three zinc finger structures (Zn1, Zn2, Zn3) that recognize DNA strand breaks, a nuclear localization signal (NLS), and the aspartate-glutamate-valine-aspartic acid (DEVD) motif that serves as the caspase cleavage site [9] [11]. The AMD domain contains a breast cancer type 1 (BRCT) structural domain and a tryptophan-glycine-arginine (WGR) domain that regulates catalytic activity, while the CAT domain houses the NAD+ binding site and poly(ADP-ribose) catalytic site [9] [4].
Under basal conditions, PARP-1 activity remains minimal. However, upon DNA damage recognition, its activity increases more than 500-fold [9]. PARP-1 catalyzes the cleavage of NAD+ into ADP-ribose and nicotinamide, subsequently synthesizing long, branched poly(ADP-ribose) (PAR) polymers on target proteins, including itself, at specific glutamate and serine residues (Glu 488, Glu 491, Ser 499, Ser 507, Ser 519) [9]. This PARylation acts as a signal for DNA repair machinery recruitment, facilitating base excision repair (BER) and maintaining genomic integrity [22] [23].
PARP-1 Cleavage: The Apoptotic Switch
During apoptosis, executioner caspases-3 and -7 cleave PARP-1 at the DEVD214↓G215 site located within the DBD, separating the 24-kDa N-terminal fragment (containing Zn1, Zn2, and the NLS) from the 89-kDa C-terminal fragment (containing Zn3, BRCT, WGR, and CAT domains) [11] [4] [10]. This proteolytic event serves as a well-established biochemical marker of apoptosis, with several critical functional consequences.
Table 1: PARP-1 Cleavage Fragments and Their Properties
| Fragment | Size | Domains Contained | Cellular Localization | Primary Functions |
|---|---|---|---|---|
| N-terminal | 24 kDa | Zn1, Zn2, NLS, DEVD motif | Nuclear | Dominant-negative inhibition of DNA repair; occupies DNA breaks [4] |
| C-terminal | 89 kDa | Zn3, BRCT, WGR, CAT | Cytoplasmic | Novel signaling functions; PAR carrier to cytoplasm [4] [10] |
Inhibition of DNA Repair
The cleavage of PARP-1 effectively terminates its DNA repair capabilities through two complementary mechanisms. First, the separation of the DNA-binding zinc fingers (Zn1, Zn2) in the 24-kDa fragment from the catalytic domain in the 89-kDa fragment disrupts the enzyme's ability to synthesize PAR polymers at DNA damage sites [11]. Second, the 24-kDa fragment remains nuclear localized and can act as a dominant-negative inhibitor by binding to DNA strand breaks and preventing access by other repair proteins [4]. This ensures that DNA damage remains unrepaired, committing the cell to death and preventing resource expenditure on futile repair attempts.
Conservation of Cellular Energy
PARP-1 overactivation in response to severe DNA damage can consume massive amounts of NAD+ and, subsequently, ATP in efforts to resynthesize NAD+, leading to energy depletion and necrotic cell death [22] [11]. By cleaving PARP-1 early in apoptosis, caspases prevent this energy catastrophe through several mechanisms:
- Preventing NAD+ Depletion: Termination of PARP-1's catalytic activity conserves cellular NAD+ pools [11]
- Maintaining ATP Levels: By preserving NAD+, cells maintain mitochondrial function and ATP production necessary for the energy-dependent apoptotic process [11]
- Facilitating Apoptotic Execution: ATP is required for multiple steps in apoptosis, including caspase activation, apoptotic body formation, and phagocytic recognition [11]
This cleavage event essentially functions as a molecular switch that redirects cell death from necrotic to apoptotic pathways, preventing inflammation and ensuring orderly cell disposal [11].
Figure 1: PARP-1 Cleavage as a Molecular Switch Between Cell Death Pathways. Caspase-mediated cleavage of PARP-1 during severe DNA damage conserves cellular energy and promotes apoptosis, while unregulated PARP-1 overactivation leads to energy depletion and necrosis.
Novel Functions of Cleavage Fragments
Recent research has revealed that the PARP-1 cleavage fragments possess biological activities beyond simply terminating DNA repair. The 89-kDa C-terminal fragment translocates to the cytoplasm during apoptosis, where it exhibits novel signaling functions [4] [10].
Cytoplasmic Signaling and Innate Immune Activation
The 89-kDa fragment recognizes and interacts with the RNA polymerase III (Pol III) complex in the cytosol through its BRCT domain [4]. This interaction facilitates mono-ADP-ribosylation of Pol III subunits, enhancing the transcription of foreign DNA and promoting IFN-β production during innate immune responses to pathogenic infection [4]. This mechanism connects PARP-1 cleavage to antiviral defense mechanisms during apoptosis induced by cellular stress.
AIF-Mediated Parthanatos
In alternative cell death pathways, the 89-kDa fragment can serve as a PAR carrier to the cytoplasm, where it facilitates apoptosis-inducing factor (AIF) release from mitochondria [10]. This process, known as parthanatos, represents a caspase-independent programmed cell death pathway that can be initiated by PARP-1 overactivation [10].
Table 2: Cell Death Pathways Involving PARP-1
| Pathway | Initiators | PARP-1 Role | Energy Status | Key Features |
|---|---|---|---|---|
| Apoptosis | Caspase activation, death receptors | Cleaved by caspases | ATP-dependent | Controlled, anti-inflammatory; PARP-1 cleavage conserves ATP [11] |
| Necrosis | Extreme DNA damage, physicochemical stress | Overactivated, consumes NAD+/ATP | ATP-depleted | Inflammatory; cell swelling; membrane disruption [11] |
| Parthanatos | Mild DNA damage, oxidative stress | Activated, produces PAR polymers | Variable | Caspase-independent; AIF-mediated; PAR translocation [10] |
Experimental Methods for Studying PARP-1 Cleavage
Detection and Quantification Protocols
Western Blot Analysis for PARP-1 Cleavage
- Cell Lysis: Harvest cells and lyse in RIPA buffer (50 mM Tris-HCl pH 7.4, 150 mM NaCl, 1% NP-40, 0.5% sodium deoxycholate, 0.1% SDS) supplemented with protease inhibitors (1 mM PMSF, 10 μg/mL aprotinin, 10 μg/mL leupeptin) and 1 μM caspase inhibitor to prevent post-lysis cleavage [11] [4]
- Electrophoresis: Separate 20-50 μg of protein extract on 8-10% SDS-PAGE gels for 1-2 hours at 120V
- Transfer and Blocking: Transfer to PVDF membrane, block with 5% non-fat milk in TBST for 1 hour
- Antibody Incubation: Incubate with primary antibodies specific for full-length PARP-1 (116 kDa) and the 89-kDa cleavage fragment (1:1000 dilution, 4°C overnight). Use antibodies that recognize the DEVD cleavage site for specific detection of apoptosis-generated fragments [4]
- Detection: Develop with HRP-conjugated secondary antibodies (1:5000) and ECL reagent. Quantify band intensity using densitometry software
Flow Cytometry with Annexin V/PI Staining
- Cell Preparation: Harvest 1×10^6 cells per condition, wash with cold PBS
- Staining: Resuspend in 100 μL binding buffer containing Annexin V-FITC (1:100) and propidium iodide (PI, 1 μg/mL)
- Analysis: Incubate for 15 minutes in dark, add 400 μL binding buffer, and analyze within 1 hour using flow cytometry with 488 nm excitation [4]
- Gating Strategy: Establish quadrants for viable (Annexin V-/PI-), early apoptotic (Annexin V+/PI-), late apoptotic (Annexin V+/PI+), and necrotic (Annexin V-/PI+) populations
Immunofluorescence Microscopy
- Cell Fixation: Culture cells on coverslips, fix with 4% paraformaldehyde for 15 minutes, permeabilize with 0.2% Triton X-100 for 10 minutes
- Staining: Block with 3% BSA, incubate with anti-PARP-1 (1:200) and anti-cleaved caspase-3 (1:500) antibodies overnight at 4°C
- Visualization: Use species-appropriate fluorescent secondary antibodies (1:1000), counterstain with DAPI, mount with antifade reagent
- Image Analysis: Capture images using confocal microscopy; assess nuclear fragmentation and cytoplasmic translocation of PARP-1 fragments
Induction of PARP-1 Cleavage in Experimental Systems
Chemical Inducers
- Staurosporine: 0.1-1 μM for 2-8 hours; broad-spectrum protein kinase inducer of intrinsic apoptosis [10]
- Actinomycin D: 0.5-5 μg/mL for 4-12 hours; transcription inhibitor that activates p53-dependent apoptosis [10]
- TNF-α with Cycloheximide: 10-50 ng/mL TNF-α with 1-10 μg/mL cycloheximide for 4-16 hours; induces extrinsic apoptosis in sensitive cell lines [11]
- Poly(dA-dT) Transfection: 0.5-2 μg/mL for 12-24 hours; mimics pathogenic DNA and activates cytoplasmic DNA sensing pathways [4]
Genetic Models
- PARP-1-D214N Mutant: Caspase-resistant PARP-1 with aspartate to asparagine substitution at cleavage site; enables study of cleavage-independent apoptosis [11]
- PARP-1 Knockout Cells: Cells deficient in PARP-1; useful for rescue experiments with wild-type and cleavage-resistant mutants [11] [4]
Figure 2: Experimental Workflow for PARP-1 Cleavage Studies. Comprehensive methodology for investigating PARP-1 cleavage encompasses treatment groups with various inducers and inhibitors, multiple detection techniques, and integrated analysis of cell death parameters.
Research Reagent Solutions
Table 3: Essential Research Reagents for PARP-1 Cleavage Studies
| Reagent Category | Specific Examples | Function/Application | Key Considerations |
|---|---|---|---|
| PARP-1 Antibodies | Anti-PARP-1 (full length), Anti-cleaved PARP-1 (Asp214), Anti-PARP-1 p89 | Detection of full-length and cleaved PARP-1 by WB, IF | Antibodies targeting the cleavage site (Asp214) provide apoptosis-specific detection [4] |
| Caspase Inhibitors | zVAD-fmk (pan-caspase), DEVD-CHO (caspase-3/7 specific) | Inhibit caspase activity to prevent PARP-1 cleavage | zVAD-fmk potentiates TNF-induced necrosis by blocking PARP-1 cleavage [11] |
| PARP Inhibitors | Olaparib, Niraparib, 3-aminobenzamide (3-AB) | Inhibit PARP catalytic activity; research tools and therapeutics | 3-AB suppresses PARP-1 overactivation and TNF-induced necrosis [11] [24] |
| Apoptosis Inducers | Staurosporine, Actinomycin D, TNF-α + CHX | Induce apoptosis and PARP-1 cleavage | Concentration and duration determine death modality (apoptosis vs. necrosis) [11] [10] |
| Cell Lines | PARP-1(-/-) cells, PARP-1-D214N mutants | Genetic models to study PARP-1 function | Cells expressing caspase-resistant PARP-1 show enhanced sensitivity to TNF-induced death [11] |
| Detection Kits | Annexin V-FITC/PI apoptosis detection, Caspase-3 activity assays | Quantify apoptosis and caspase activation | Annexin V/PI staining distinguishes apoptotic and necrotic populations [4] |
Therapeutic Implications and Research Applications
PARP Inhibitors in Cancer Therapy
PARP inhibitors (PARPi) have emerged as targeted therapies for homologous recombination-deficient cancers, particularly those with BRCA1/2 mutations, through synthetic lethality [25]. These inhibitors trap PARP-1 on DNA, preventing its dissociation and creating cytotoxic lesions that are lethal in DNA repair-deficient backgrounds [26] [25]. Understanding PARP-1 cleavage mechanisms has important implications for PARPi applications:
- Predictive Biomarkers: PARP-1 cleavage status may indicate apoptotic response to PARPi therapy [25]
- Resistance Mechanisms: Tumors with reduced PARP-1 expression or hypomorphic PARP-1 variants exhibit diminished PARP trapping and resistance to PARPi [25]
- Combination Strategies: PARPi combinations with DNA-damaging agents must consider scheduling to maximize efficacy while minimizing toxicity [26]
Novel Drug Development Approaches
The development of PARP-1 selective inhibitors represents an emerging strategy to overcome hematologic toxicity associated with dual PARP-1/PARP-2 inhibition [24]. Structural analyses reveal conformational differences in the active regions of PARP-1 and PARP-2 that enable selective inhibitor design, potentially widening the therapeutic window of PARP-targeted therapies [24].
Advanced clinical trials are exploring innovative combination and scheduling approaches. A phase I trial (NCT02769962) demonstrated that combining tumor-targeted topoisomerase I inhibitor delivery (CRLX101) with gapped olaparib scheduling enabled higher PARP inhibitor dosing while reducing toxicity, showing promising activity in advanced solid tumors [26].
PARP-1 cleavage represents a critical commitment point in apoptotic cell death, serving dual functions of terminating DNA repair capacity while conserving cellular energy to support the orderly execution of apoptosis. The 24-kDa and 89-kDa cleavage fragments exhibit distinct biological activities beyond their canonical roles, with emerging functions in innate immune activation and alternative cell death pathways. Methodological advances in detecting and quantifying PARP-1 cleavage provide robust tools for basic research and drug development, particularly in the context of PARP inhibitor therapies for cancer treatment. As research continues to elucidate the complex functions of PARP-1 and its cleavage products, new therapeutic opportunities will likely emerge for manipulating this key cell death switch in human disease.
Poly(ADP-ribose) polymerase-1 (PARP-1) is a nuclear enzyme renowned for its role in DNA repair and as a canonical substrate for caspase-3 during apoptosis. However, its cleavage serves as a biomarker for a far broader spectrum of proteolytic activities and cell death pathways. While caspase-mediated generation of the signature 24 kDa and 89 kDa fragments remains a hallmark of apoptosis, PARP-1 is also a preferred substrate for numerous other "suicidal proteases," including calpains, granzymes, cathepsins, and matrix metalloproteinases (MMPs). These proteases produce distinct PARP-1 cleavage fragments that serve as unique signatures for identifying specific protease activation and particular forms of cell death in pathophysiology. This review details the mechanisms and consequences of PARP-1 cleavage by these alternative proteases, framing it within the broader context of cell death research and highlighting its implications for drug development.
PARP-1 is a highly abundant nuclear protein with multifaceted roles in cellular homeostasis. Its primary function involves the detection of DNA strand breaks and the subsequent catalysis of poly(ADP-ribose) (PAR) chains onto target proteins, facilitating DNA repair [27] [16]. However, upon irreversible cell death commitment, PARP-1 becomes a primary target for proteolytic cleavage by activated proteases. The classic 89 kDa and 24 kDa fragments generated by caspases are definitive biomarkers of apoptosis [27] [11]. Yet, focusing solely on caspases overlooks a critical dimension of PARP-1 biology: its susceptibility to a diverse family of suicidal proteases. Cleavage by these proteases—calpains in necrosis, granzymes in immune-mediated killing, cathepsins in lysosomal pathways, and MMPs—results in unique fragment signatures that act as molecular fingerprints, revealing the specific death pathway activated [27]. Understanding these distinct cleavage events is paramount for accurately diagnosing cell death modes in pathological contexts and for developing targeted therapeutic strategies.
The PARP-1 Proteolysis Landscape
PARP-1 features a modular domain structure that dictates its cleavage patterns [27] [9]:
- DNA-Binding Domain (DBD): Located at the N-terminus, it contains two zinc fingers (F1 and F2) responsible for recognizing DNA damage [16].
- Automodification Domain (AMD): A central domain that acts as an acceptor for PAR chains.
- Catalytic Domain (CAT): The C-terminal domain that catalyzes PAR synthesis using NAD+ as a substrate.
The protease cleavage sites are situated within the DBD, particularly affecting the nuclear localization signal (NLS) [28]. Different proteases target specific residues within this region, leading to the generation of fragments with altered functions and localization, which ultimately influences the cell's fate.
Caspase-Independent PARP-1 Cleavage by Suicidal Proteases
Calpains
Calpains are calcium-dependent cysteine proteases activated during necrotic cell death and excitotoxicity.
- Cleavage Signature: Calpain cleavage produces a 50 kDa fragment encompassing the DBD and a separate 64 kDa fragment [27].
- Functional Consequences: Calpain activation is associated with PARP-1 overactivation, leading to severe depletion of cellular NAD+ and ATP pools. This energy collapse is a hallmark of necrotic cell death [27] [11]. The cleavage by calpain is thought to contribute to this pathological process.
- Pathophysiological Context: Calpain-mediated PARP-1 cleavage is prominent in neurological pathologies such as cerebral ischemia, traumatic brain injury, and NMDA-mediated excitotoxicity [27].
Granzymes
Granzymes are serine proteases delivered by cytotoxic T lymphocytes and natural killer (NK) cells to eliminate virus-infected and cancerous cells.
- Cleavage Signature: Granzyme A cleaves PARP-1 to generate a ~50 kDa fragment, while Granzyme B, which has a caspase-like activity, can produce the classic 89 kDa and 24 kDa apoptotic-like fragments [27].
- Functional Consequences: This cleavage disrupts the DNA repair capacity of the target cell, ensuring the successful execution of the immune-mediated cell death pathway.
- Pathophysiological Context: A key mechanism in immune surveillance and the anti-tumor response [27].
Other Proteases: Cathepsins and MMPs
- Cathepsins: These lysosomal proteases can cleave PARP-1 upon lysosomal membrane permeabilization, contributing to cell death under oxidative stress [27].
- Matrix Metalloproteinases (MMPs): Certain MMPs, such as MMP-2 and MMP-9, have been reported to cleave PARP-1, although the precise fragments and functional outcomes are an area of ongoing research [27].
Table 1: Summary of PARP-1 Cleavage by Suicidal Proteases
| Protease | Class | Cleavage Fragments | Primary Cell Death Pathway | Key Regulatory Role |
|---|---|---|---|---|
| Caspase-3/7 | Cysteine-aspartic | 24 kDa (DBD) + 89 kDa (AMD+CAT) | Apoptosis | Inactivates DNA repair, conserves ATP [27] [11] |
| Calpain | Calcium-dependent cysteine | 50 kDa + 64 kDa | Necrosis | Promotes energy depletion, necrotic cascade [27] |
| Granzyme A | Serine | ~50 kDa | Immune-mediated cytotoxicity | Disables DNA repair in target cells [27] |
| Granzyme B | Serine | 24 kDa + 89 kDa | Immune-mediated apoptosis | Mimics caspase-3 to induce apoptosis [27] |
| Cathepsin | Lysosomal cysteine | Multiple (e.g., 40-50 kDa) | Lysosomal cell death | Contributes to death under oxidative stress [27] |
| MMP | Metalloendopeptidase | Various (further characterization needed) | Associated with inflammation & cancer | Potential role in extracellular signaling [27] |
Functional Consequences of PARP-1 Cleavage Fragments
The biological outcome of PARP-1 proteolysis is dictated by the functions of the resulting fragments.
- The 24 kDa DBD Fragment: This fragment, which contains the zinc fingers F1 and F2, retains a high affinity for DNA strand breaks. It acts as a trans-dominant inhibitor of full-length PARP-1 and other DNA repair proteins by occupying DNA damage sites and blocking repair machinery access. This function is crucial in apoptosis to prevent DNA repair and ensure cell demise [27] [16].
- The 89 kDa Fragment (AMD + CAT): This fragment has a greatly reduced DNA-binding capacity. Recent research has unveiled a pro-apoptotic role where, upon caspase cleavage, it can be poly(ADP-ribosyl)ated and translocated to the cytoplasm. Here, it acts as a carrier for PAR polymers, facilitating the release of Apoptosis-Inducing Factor (AIF) from mitochondria. AIF translocation to the nucleus initiates caspase-independent DNA fragmentation, a pathway known as parthanatos [10]. This illustrates a direct mechanistic link between caspase activity and a caspase-independent death pathway.
- Differential Regulation by Cleavage Products: Studies expressing individual cleavage fragments in neurons have demonstrated their opposing roles. The expression of the 24 kDa fragment or an uncleavable PARP-1 mutant conferred protection from ischemic damage in vitro, while the 89 kDa fragment was cytotoxic. Furthermore, the 89 kDa fragment induced significantly higher NF-κB activity and expression of pro-inflammatory proteins like iNOS and COX-2, suggesting that PARP-1 cleavage products can differentially regulate inflammatory responses during cell stress [28].
Experimental Protocols for PARP-1 Cleavage Analysis
Detecting PARP-1 Cleavage Fragments by Western Blotting
Objective: To identify and distinguish specific PARP-1 cleavage fragments generated by different suicidal proteases in cell culture models.
Materials and Reagents:
- Cell Lines: Appropriate models (e.g., SH-SY5Y neuroblastoma cells, primary cortical neurons, L929 fibrosarcoma cells) [28] [11].
- Inducers: Staurosporine (apoptosis inducer), TNF-α (necrosis inducer in L929 cells), Actinomycin D, Etoposide (VP-16), RSL3 (ferroptosis/apoptosis inducer) [27] [11] [20].
- Inhibitors: zVAD-fmk (pan-caspase inhibitor), 3-aminobenzamide (3AB, PARP inhibitor), Calpain inhibitors (e.g., MDL-28170), Ferrostatin-1 (Fer-1, ferroptosis inhibitor) [11] [20].
- Antibodies: Anti-PARP-1 antibody that detects full-length (~116 kDa) and key fragments (89 kDa, 50 kDa, 24 kDa).
Methodology:
- Cell Treatment and Stimulation: Seed cells and treat with chosen inducers and/or inhibitors. For example, to distinguish caspase-dependent and -independent cleavage, pre-treat cells with zVAD-fmk before applying a death stimulus like TNF-α [11].
- Protein Extraction and Quantification: Lyse cells at specific time points post-treatment using RIPA buffer supplemented with protease inhibitors.
- Western Blotting: Separate proteins via SDS-PAGE (8-12% gradient gels are suitable), transfer to PVDF membrane, and probe with anti-PARP-1 antibody.
- Fragment Analysis: Identify the signature fragment pattern:
- 89 kDa & 24 kDa: Caspase-mediated apoptosis.
- 50 kDa & 64 kDa: Suggests calpain activity.
- ~50 kDa (Granzyme A signature): In models of immune cell killing.
Assessing the Functional Role of Cleavage Using Mutagenesis
Objective: To investigate the functional consequences of preventing PARP-1 cleavage.
Methodology:
- Generation of Uncleavable PARP-1 (PARP-1UNCL): Create a point mutation (D214N) in the caspase cleavage site (DEVD) within the PARP-1 cDNA [28] [11].
- Cell Transfection/Transduction: Stably or transiently express PARP-1UNCL, PARP-1WT (wild-type), and individual fragments (PARP-124, PARP-189) in cells. Viral vectors (e.g., AAV) can be used for primary neurons [28].
- Functional Assays:
- Viability Assays: Expose transfected cells to stressors like Oxygen/Glucose Deprivation (OGD) and measure cell death using MTT, LDH release, or Annexin V/PI staining [28].
- NF-κB Activity: Measure NF-κB transcriptional activity using luciferase reporter assays and monitor downstream targets like iNOS and COX-2 by qPCR and Western blot [28].
- Energy Metabolism: Assess NAD+ and ATP levels to determine the impact of cleavage on cellular energy status [11].
The Scientist's Toolkit: Key Research Reagents
Table 2: Essential Reagents for Studying PARP-1 Cleavage
| Reagent / Tool | Function / Specificity | Example Application |
|---|---|---|
| zVAD-fmk | Irreversible, pan-caspase inhibitor | Distinguishing caspase-dependent vs. independent PARP-1 cleavage and cell death pathways [11]. |
| 3-Aminobenzamide (3AB) | PARP catalytic activity inhibitor | Investigating the role of PAR formation in cell death and the functions of PARylated fragments [11]. |
| PARP-1UNCL Plasmid | Expresses caspase-uncleavable PARP-1 (D214N mutant) | Determining the specific biological consequences of preventing PARP-1 cleavage [28] [11]. |
| Anti-PARP-1 Antibody | Detects full-length and cleavage fragments | Western blot analysis to identify protease-specific cleavage signatures [27] [20]. |
| RSL3 | GPX4 inhibitor, induces ferroptosis/apoptosis | Studying crosstalk between ferroptosis and apoptosis, and its impact on PARP1 cleavage and depletion [20]. |
| Staurosporine / Actinomycin D | Broad-spectrum inducers of apoptosis | Triggering caspase-3 activation and the classic 89/24 kDa PARP-1 cleavage for model establishment [10]. |
| TNF-α (in L929 cells) | Inducer of PARP-dependent necrosis | Modeling necrotic cell death and studying calpain-mediated PARP-1 cleavage [11]. |
Visualizing PARP-1 Cleavage Pathways and Experimental Logic
The following diagrams illustrate the core concepts and experimental workflows discussed in this review.
Diagram 1: PARP-1 Cleavage Integrates Multiple Cell Death Pathways. Different lethal stimuli activate specific proteases, which cleave PARP-1 into signature fragments. These fragments drive distinct cell fate decisions, including apoptosis, necrosis, parthanatos, and inflammation.
Diagram 2: Experimental Workflow for PARP-1 Cleavage Research. A logical flow for designing experiments to study PARP-1 cleavage, from model establishment and perturbation to detection, functional assessment, and mechanistic investigation.
The cleavage of PARP-1 is a critical event that extends far beyond its role as a simple caspase substrate and apoptosis marker. It represents a molecular integration point for multiple cell death pathways, with distinct proteases generating unique signature fragments that dictate cellular outcomes. Recognizing the 50 kDa calpain-specific fragment or the granzyme-generated fragments, for instance, allows researchers to accurately diagnose necrosis or immune-mediated killing in complex pathological samples.
Future research and drug development efforts must account for this complexity. The role of PARP-1 fragments in regulating transcription and inflammation, particularly through pathways like NF-κB, presents a promising therapeutic avenue [28]. Furthermore, inducing specific PARP-1 cleavage patterns represents a novel strategy, as demonstrated by the ability of RSL3 to promote both caspase-dependent PARP-1 cleavage and full-length PARP-1 depletion, showing efficacy even in PARP inhibitor-resistant tumors [20]. As we continue to decipher the "PARP-1 cleavage code," we unlock new potential for precisely diagnosing and therapeutically modulating cell death in cancer, neurodegeneration, and other diseases.
Detecting PARP-1 Cleavage: Techniques, Assays, and Applications in Research and Drug Discovery
Within the broader investigation of what constitutes PARP-1 cleavage and why it serves as an apoptosis marker, the detection of its specific proteolytic fragments is paramount. During programmed cell death, caspases-3 and -7 cleave the 116-kDa PARP-1 protein into characteristic 24-kDa and 89-kDa fragments. This in-depth technical guide establishes western blot analysis as the gold standard for identifying these signature fragments. We detail optimized protocols for sample preparation, gel electrophoresis, and immunoblotting, supported by structured data on fragment size and function. Furthermore, we present essential research reagents and visualize the central signaling pathway, providing researchers and drug development professionals with a definitive framework for validating PARP-1 cleavage as a hallmark of apoptotic commitment.
Poly (ADP-ribose) polymerase-1 (PARP-1) is a 116-kDa nuclear enzyme with critical functions in DNA repair, genomic stability, and gene transcription [3] [29]. In response to minor DNA damage, PARP-1 is activated and facilitates DNA repair. However, during the irreversible commitment to apoptosis, PARP-1 becomes one of the primary substrates for executioner caspases [3] [15]. The proteolytic cleavage of PARP-1 is a definitive event in the apoptotic cascade, serving to inactivate DNA repair processes and conserve cellular energy (ATP and NAD+) for the orderly dismantling of the cell [3] [30].
The cleavage occurs at a specific aspartic acid residue (Asp214) located within a conserved nuclear localization signal sequence, catalyzed predominantly by caspases-3 and -7 [30] [6]. This reaction severs the full-length 116-kDa protein into two signature fragments: an 89-kDa C-terminal fragment (containing the automodification and catalytic domains) and a 24-kDa N-terminal fragment (containing the DNA-binding domain) [30] [6]. The detection of these specific fragments, particularly the 89-kDa species, is widely recognized as a biochemical hallmark of apoptosis [3] [15]. Western blot analysis stands as the most reliable and widely used technique for the specific and sensitive detection of these cleavage fragments, solidifying its role as a gold standard in cell death research and drug discovery.
The Biochemical Basis of PARP-1 Cleavage
Domain Architecture and Cleavage Site
PARP-1 is structurally organized into three primary functional domains [3] [6]:
- DNA-Binding Domain (DBD, 46-kDa): Located at the N-terminus, this domain contains two zinc finger motifs that recognize and bind to DNA strand breaks. This binding activates the catalytic domain.
- Automodification Domain (AMD, 22-kDa): The central domain is the target for covalent auto-poly(ADP-ribosyl)ation, which modulates the enzyme's interaction with DNA and other proteins.
- Catalytic Domain (CD, 54-kDa): Residing at the C-terminus, this domain catalyzes the transfer of ADP-ribose units from NAD+ to acceptor proteins.
The critical caspase cleavage site is located between the DBD and the AMD, specifically after Asp214 within the DEVD214G amino acid sequence [30] [6]. Cleavage at this site physically separates the DNA-binding function from the catalytic function.
Fate and Function of the Cleavage Fragments
The cleavage event produces two stable fragments with distinct cellular fates and functions, as summarized in Table 1.
Table 1: Characteristics and Functions of PARP-1 Cleavage Fragments
| Fragment | Molecular Weight | Domains Contained | Cellular Localization Post-Cleavage | Primary Function Post-Cleavage |
|---|---|---|---|---|
| 24-kDa | 24 kDa | DNA-Binding Domain (DBD) | Remains tightly bound to DNA in the nucleus [3] [6] | Acts as a trans-dominant inhibitor of DNA repair by blocking access of intact PARP-1 and other repair enzymes to DNA strand breaks [3]. |
| 89-kDa | 89 kDa | Automodification Domain (AMD) & Catalytic Domain (CD) | Liberated from the nucleus and translocated to the cytoplasm [31] [6] | Serves as a cytoplasmic carrier of poly(ADP-ribose) (PAR) polymers, potentially inducing AIF-mediated cell death (parthanatos) and facilitating cellular disassembly [31] [6]. |
The following diagram illustrates the domain structure of full-length PARP-1, the caspase cleavage event, and the fate of the resulting fragments.
Western Blot Methodology for Detecting PARP-1 Cleavage
Sample Preparation from Apoptotic Cells
Key Reagents:
- Lysis Buffer: RIPA buffer (25 mM Tris-HCl pH 7.6, 150 mM NaCl, 1% NP-40, 1% sodium deoxycholate, 0.1% SDS) supplemented immediately before use with 1 mM PMSF (or other serine protease inhibitors) and a broad-spectrum phosphatase inhibitor cocktail.
- Inducers of Apoptosis (Positive Controls): To validate your assay, treat cells with well-characterized apoptosis inducers. Etoposide (a topoisomerase II inhibitor) [32] and Staurosporine (a broad-spectrum kinase inhibitor) [6] are highly effective and widely published for triggering PARP-1 cleavage.
Protocol:
- Induce Apoptosis: Treat cells (e.g., HeLa, SH-SY5Y) with your chosen apoptotic stimulus. For example, expose HeLa cells to 1 µM Staurosporine for 3-6 hours [6].
- Harvest Cells: Collect cells by gentle scraping or trypsinization, followed by centrifugation.
- Lyse Cells: Resuspend the cell pellet in ice-cold RIPA lysis buffer (e.g., 100 µL per 1x10⁶ cells) and incubate on ice for 30 minutes with intermittent vortexing.
- Clarify Lysate: Centrifuge at 14,000 x g for 15 minutes at 4°C. Carefully transfer the supernatant (whole cell lysate) to a new tube.
- Quantify Protein: Determine protein concentration using a BCA or Bradford assay.
- Prepare Samples: Dilute lysates in Laemmli sample buffer to a final 1X concentration, heat at 95-100°C for 5 minutes, and immediately load onto a gel or store at -20°C.
Gel Electrophoresis and Immunoblotting
Key Reagents:
- Gel System: Precast or handcast 4-20% Tris-Glycine gradient gels are ideal for resolving the large molecular weight range covering full-length PARP-1 (116 kDa) and the 89-kDa fragment.
- Primary Antibody: The critical reagent is an anti-PARP-1 antibody that recognizes an epitope C-terminal to the caspase cleavage site, thus detecting both full-length PARP-1 and the 89-kDa fragment. The PARP Antibody #9542 (Cell Signaling Technology) is a well-validated example. It is a rabbit polyclonal antibody raised against a synthetic peptide corresponding to the caspase cleavage site, providing high specificity for the apoptotic signature [30].
- Secondary Antibody: HRP-conjugated anti-rabbit IgG.
- Detection System: Enhanced Chemiluminescence (ECL) substrate.
Protocol:
- Electrophoresis: Load 20-30 µg of total protein per lane alongside a pre-stained protein ladder. Run the gel at a constant voltage (e.g., 120-150V) until the dye front reaches the bottom.
- Protein Transfer: Transfer proteins from the gel to a PVDF or nitrocellulose membrane using a wet or semi-dry transfer system.
- Blocking: Incubate the membrane in a 5% non-fat dry milk or BSA solution in TBST (Tris-Buffered Saline with 0.1% Tween-20) for 1 hour at room temperature to block non-specific binding sites.
- Primary Antibody Incubation: Incubate the membrane with the primary anti-PARP antibody (e.g., PARP Antibody #9542 at a 1:1000 dilution in blocking buffer) overnight at 4°C with gentle agitation [30].
- Washing: Wash the membrane 3 times for 5-10 minutes each with TBST.
- Secondary Antibody Incubation: Incubate with the HRP-conjugated secondary antibody (typically 1:2000 to 1:10000 dilution in blocking buffer) for 1 hour at room temperature.
- Washing: Repeat the TBST wash step.
- Detection: Incubate the membrane with ECL substrate according to the manufacturer's instructions and visualize using a digital imager or X-ray film.
Expected Results and Interpretation
A successful western blot will show a clear band at 116 kDa corresponding to the full-length PARP-1 in healthy, non-apoptotic cells. In apoptotic samples, a second, strong band at 89 kDa will appear, often with a concomitant reduction in the intensity of the 116-kDa band. The 24-kDa fragment is less commonly detected in standard western blots, as the epitope for most commercial antibodies (like #9542) resides in the C-terminal 89-kDa fragment. Specific antibodies targeting the N-terminal DBD are required to visualize the 24-kDa band.
The Scientist's Toolkit: Essential Research Reagents
Successful detection and study of PARP-1 cleavage rely on a set of well-characterized reagents. The following table details essential tools for researchers in this field.
Table 2: Key Research Reagents for PARP-1 Cleavage Analysis
| Reagent / Tool | Specific Example | Function and Application in PARP-1 Research |
|---|---|---|
| Validated PARP-1 Antibody | PARP Antibody #9542 (Cell Signaling Technology) [30] | A cornerstone reagent for western blotting; specifically detects endogenous levels of full-length (116 kDa) PARP1 and the large caspase-cleaved fragment (89 kDa). |
| Caspase Inhibitor | zVAD-fmk (broad-spectrum) [6] | A cell-permeable pan-caspase inhibitor. Used as a control to confirm that fragment generation is caspase-dependent; pre-treatment should block the appearance of the 89-kDa band upon apoptotic stimulation. |
| PARP Activity Inhibitor | PJ34 [6] [28] | A potent, cell-permeable PARP enzymatic inhibitor. Used to dissect the role of PARP-1's catalytic activity in cell death pathways and to study its non-catalytic functions. |
| Apoptosis Inducer | Staurosporine, Etoposide, Actinomycin D [32] [31] [6] | Well-established chemical inducers of intrinsic apoptosis. Serve as robust positive controls in experiments designed to trigger and observe PARP-1 cleavage. |
| siRNA for PARP-1 Knockdown | PARP-1 specific siRNA (e.g., Target Sequence: 5'-ACGGTGATCGGTAGCAACAAA-3′) [28] | Used to generate PARP-1-deficient cell lines, enabling functional rescue experiments with mutant constructs (e.g., uncleavable PARP-1) to study the specific role of cleavage. |
Western blot analysis remains the unequivocal gold standard for the specific and reliable detection of PARP-1 cleavage into its 89-kDa and 24-kDa fragments. This technique provides a direct biochemical readout of caspase activation, a definitive event in the apoptotic cascade. The methodology outlined here—from robust sample preparation using validated apoptotic inducers to optimized immunoblotting with highly specific antibodies—provides a rigorous framework for researchers. The consistent detection of the 89-kDa PARP-1 fragment via western blot is more than a technical endpoint; it is a fundamental biomarker that confirms the activation of a cell's self-destruct mechanism. As research continues to unravel the complex roles of PARP-1 fragments in integrating different cell death pathways, western blotting will remain an indispensable tool for validating these findings in both basic research and the development of novel cancer therapeutics that target the apoptotic machinery.
This technical guide provides detailed methodologies for visualizing the subcellular localization of Poly (ADP-ribose) polymerase-1 (PARP-1) cleavage fragments, a hallmark of apoptosis. PARP-1 cleavage into specific fragments serves as a critical biomarker for distinguishing cell death pathways and has significant implications for basic research and drug development, particularly in oncology and neurodegenerative diseases. We present optimized immunofluorescence protocols, address technical challenges in detection, and explore the functional consequences of fragment localization, providing researchers with comprehensive tools to advance studies in cell death mechanisms.
PARP-1 is a nuclear enzyme with crucial roles in DNA repair, genomic stability, and transcriptional regulation [3]. During apoptosis, PARP-1 undergoes proteolytic cleavage at the DEVD214 site by executioner caspases-3 and -7, generating two characteristic fragments: a 24-kDa DNA-binding domain (DBD) fragment and an 89-kDa catalytic domain fragment [28] [11]. This cleavage event serves as a well-established biochemical hallmark of apoptosis, as it inactivates PARP-1's DNA repair function and facilitates the dismantling of the cell [3] [11].
The detection of these cleavage fragments, particularly through immunofluorescence and microscopy, provides researchers with a powerful tool to not only confirm apoptosis but also to gain insights into the intricate subcellular redistribution of proteins during cell death. Different cleavage fragments exhibit distinct localization patterns: the 24-kDa fragment remains nuclear due to its nuclear localization signal, while the 89-kDa fragment can translocate to the cytoplasm under specific conditions, engaging in non-canonical cell death pathways such as parthanatos [31] [10]. Understanding these dynamics through visualization techniques is essential for comprehending cell fate decisions in health and disease.
PARP-1 Cleavage Fragments: Biological Significance and Detection Challenges
Fragment Characteristics and Functions
Table: PARP-1 Cleavage Fragments and Their Properties
| Fragment Size | Domains Contained | Subcellular Localization | Primary Functions |
|---|---|---|---|
| 24 kDa | Zinc fingers 1 & 2, Nuclear Localization Signal (NLS) [28] | Remains nuclear [31] | Acts as a trans-dominant inhibitor of DNA repair; irreversibly binds DNA strand breaks [3] |
| 89 kDa | Zinc finger 3, BRCT, WGR, and Catalytic Domain [4] | Nucleus (initially); can translocate to cytoplasm [31] [10] | Serves as PAR carrier to cytoplasm; induces AIF-mediated parthanatos [10]; Can ADP-ribosylate RNA Pol III during innate immune response [4] |
Technical Challenges in Visualization
Visualizing endogenous PARP-1 fragments presents significant challenges due to the abundant nuclear presence of full-length PARP-1 (approximately 1-2 million molecules per cell), which creates substantial background "noise" that obscures detection of cleaved fragments [3] [33]. Conventional immunocytological methods often fail to distinguish fragment-specific localization because most PARP-1 antibodies recognize epitopes present in both full-length and cleaved forms. Additionally, the transient nature of fragment generation and their rapid redistribution requires optimized fixation and processing methods to capture accurate spatial and temporal localization [33].
Optimized Immunofluorescence Protocol for PARP-1 Fragment Detection
In Situ Fractionation Technique
A novel in situ fractionation technique effectively addresses the challenge of high background signal by selectively extracting unbound nuclear PARP-1 while retaining chromatin-bound fractions, including those associated with DNA damage sites [33].
Detailed Methodology:
Cell Culture and Treatment:
- Culture cells on sterilized glass coverslips until 60-70% confluent.
- Induce apoptosis using appropriate stimuli (e.g., 1 μM staurosporine for 4-6 hours; 500 nM actinomycin D for 12-16 hours) [31] [10].
- For localized DNA damage, utilize local UVC irradiation through isopore membrane filters (3 μm pores) to create subnuclear damage spots [33].
Extraction and Fixation:
- Rinse cells briefly with cytoskeleton (CSK) buffer (10 mM PIPES pH 7.0, 100 mM NaCl, 300 mM sucrose, 3 mM MgCl₂).
- Extract cells with CSK buffer containing 0.5% Triton X-100 and 0.42 M NaCl (C+T+S buffer) for 10 minutes at 4°C [33].
- Fix cells with 3.7% formaldehyde in PBS for 15 minutes at room temperature.
- Quench fixation with 100 mM glycine in PBS for 10 minutes.
Immunostaining:
- Permeabilize cells with 0.2% Triton X-100 in PBS for 10 minutes.
- Block with 5% normal serum (species-matched to secondary antibody) in PBS for 1 hour.
- Incubate with primary antibodies diluted in blocking buffer overnight at 4°C.
- Recommended antibodies:
- PARP-1 full-length/fragment detector: Mouse anti-PARP-1 antibody (e.g., BD Biosciences #556494) that recognizes both full-length and 89-kDa fragment.
- PARP-1 cleavage-specific: Rabbit anti-cleaved PARP-1 (Asp214) antibody (e.g., Cell Signaling #9544) specific to the 89-kDa fragment [33].
- DNA damage marker: Mouse anti-cyclobutane pyrimidine dimer (CPD) antibody (e.g., Cosmo Bio Co. #CAC-NM-DND-001) for UV lesions [33].
- Wash 3× with PBS for 5 minutes each.
- Incubate with species-appropriate fluorescent secondary antibodies (e.g., Alexa Fluor 488, 568, or 647) for 1 hour at room temperature.
- Counterstain nuclei with DAPI (0.1 μg/mL) for 5 minutes.
- Mount coverslips using anti-fade mounting medium.
Image Acquisition and Analysis:
- Acquire images using a high-resolution confocal microscope with 63× or 100× oil immersion objectives.
- For quantitative analysis, ensure consistent exposure settings across experimental conditions.
- Calculate background-corrected fluorescence intensity at damage sites versus non-damaged nuclear areas using ImageJ or similar software [33].
Diagram: Experimental Workflow for PARP-1 Fragment Visualization. This workflow outlines the key steps for detecting PARP-1 cleavage fragments using optimized in situ fractionation and immunofluorescence.
Critical Controls and Validation
- Cleavage-specific controls: Include cells treated with caspase inhibitor (zVAD-fmk, 20-50 μM) to prevent PARP-1 cleavage and verify antibody specificity [11].
- Localization controls: Transfect cells with GFP-tagged PARP-1 DNA binding domain (DBD) to validate 24-kDa fragment nuclear retention, or with the 89-kDa fragment to monitor its cytoplasmic translocation [33].
- Apoptosis induction controls: Use multiple apoptosis inducers (e.g., TNF-α in combination with actinomycin D) to confirm consistent fragment patterns across different death stimuli [11].
The Scientist's Toolkit: Essential Research Reagents
Table: Key Reagents for PARP-1 Cleavage and Localization Studies
| Reagent Category | Specific Examples | Function/Application | Technical Notes |
|---|---|---|---|
| Apoptosis Inducers | Staurosporine (1 μM), Actinomycin D (500 nM), TNF-α + Cycloheximide [31] [11] | Activate caspases to initiate PARP-1 cleavage | Titrate concentration to achieve gradual apoptosis for optimal visualization |
| Caspase Inhibitors | zVAD-fmk (20-50 μM) [11] | Negative control: prevents PARP-1 cleavage | Pre-treat 1-2 hours before apoptosis induction |
| PARP-1 Antibodies | Anti-PARP-1 (full-length + fragments), Anti-cleaved PARP-1 (Asp214) [33] | Detect total PARP-1 and specific 89-kDa fragment | Validate specificity using PARP-1 knockout/knockdown cells |
| Fractionation Buffers | CSK buffer + 0.5% Triton X-100 + 0.42 M NaCl (C+T+S) [33] | Selective extraction of unbound nuclear PARP-1 | Critical for reducing background signal in immunofluorescence |
| Expression Constructs | GFP-PARP-1-DBD, GFP-tPARP1 (89-kDa), PARP-1-D214N (uncleavable mutant) [28] [4] [33] | Validate fragment localization and functions | PARP-1-D214N serves as cleavage-resistant control [28] |
| Cell Lines | PARP-1-deficient fibroblasts, SH-SY5Y neuroblastoma, Primary cortical neurons [28] | Model systems for apoptosis and cleavage studies | PARP-1-/- cells enable transfection studies without background |
Advanced Applications and Functional Implications
Investigating Non-Apoptotic Fragment Functions
Beyond their role as apoptosis markers, PARP-1 fragments participate in diverse cellular processes:
Cytoplasmic 89-kDa Fragment in Parthanatos: The 89-kDa fragment can serve as a PAR carrier to the cytoplasm, where it facilitates AIF (apoptosis-inducing factor) release from mitochondria, initiating caspase-independent cell death [31] [10]. This pathway becomes particularly relevant under excessive DNA damage conditions.
Immune Signaling Regulation: Recent research reveals that the 89-kDa truncated PARP-1 (tPARP1) can mono-ADP-ribosylate RNA Polymerase III in the cytoplasm during poly(dA-dT)-stimulated apoptosis, enhancing IFN-β production and amplifying apoptotic signaling in response to pathogenic DNA [4].
Diagram: PARP-1 Cleavage Fragments in Cell Death Pathways. This diagram illustrates how PARP-1 cleavage generates fragments with distinct subcellular localizations and functions in different cell death mechanisms.
Quantitative Analysis and High-Content Screening
Advanced imaging approaches enable quantitative analysis of PARP-1 cleavage dynamics:
- Automated High-Content Analysis: Develop scripts to quantify the nuclear-to-cytoplasmic ratio of 89-kDa fragment fluorescence as a metric for apoptosis progression.
- Colocalization Studies: Calculate Manders' or Pearson's correlation coefficients between PARP-1 fragments and organelle markers (e.g., mitochondrial AIF or cytoplasmic RNA Pol III) to objectively assess fragment redistribution [4] [10].
- Time-Lapse Live-Cell Imaging: Utilize GFP-tagged PARP-1 constructs with cleavage-sensitive FRET pairs or photoactivatable tags to monitor cleavage kinetics and fragment mobility in real-time.
The visualization of PARP-1 cleavage fragments through optimized immunofluorescence and microscopy techniques provides crucial insights into cell death mechanisms and their functional consequences. The methodologies outlined in this guide—particularly the in situ fractionation approach—enable researchers to overcome historical challenges in detecting these dynamic biomarkers. As research continues to reveal novel functions for PARP-1 fragments beyond apoptosis, including their roles in parthanatos and immune signaling, these visualization techniques will remain essential tools for advancing our understanding of cell death in health and disease, and for developing targeted therapies in cancer and neurodegeneration.
PARP-1 Cleavage as a Pharmacodynamic Biomarker in Preclinical Drug Testing
Poly (ADP-ribose) polymerase-1 (PARP-1) is a nuclear enzyme with a well-established role as a DNA damage sensor and key facilitator of DNA repair processes. This 113-kDa protein functions as the primary catalyst for poly (ADP-ribosyl)ation, a critical post-translational modification that occurs in response to cellular stresses [27]. PARP-1's domain architecture includes an N-terminal DNA-binding domain (DBD) containing two zinc finger motifs, a central auto-modification domain (AMD) with a BRCT fold, and a C-terminal catalytic domain (CAT) responsible for polymerizing ADP-ribose units from NAD+ onto target proteins [27] [9].
Beyond its DNA repair functions, PARP-1 participates in diverse physiological and pathological processes including transcription regulation, immune responses, and energy metabolism [27]. However, its role as a biomarker stems from its position as a central substrate for proteolytic cleavage during programmed cell death. During apoptosis, PARP-1 undergoes specific cleavage by caspase family proteases, generating signature fragments that serve as recognizable indicators of apoptotic activation [27] [4]. This proteolytic event has become one of the most established biomarkers for detecting and quantifying apoptosis in preclinical drug development, particularly for therapeutic agents designed to trigger programmed cell death in cancer cells.
PARP-1 Cleavage as an Apoptosis Marker
Mechanism of Cleavage and Fragment Generation
The cleavage of PARP-1 during apoptosis is primarily executed by effector caspases, most notably caspase-3 and caspase-7. These proteases recognize a specific DEVD amino acid sequence (aspartate-glutamate-valine-aspartate) within PARP-1's auto-modification domain, cleaving the 113-kDa full-length protein into two characteristic fragments: a 24-kDa DNA-binding fragment and an 89-kDa catalytic fragment [27] [4] [9].
Table 1: PARP-1 Fragments Generated by Proteolytic Cleavage
| Fragment Size | Domains Contained | Cellular Localization After Cleavage | Functional Consequences |
|---|---|---|---|
| 24-kDa | Two zinc finger motifs from DBD | Retained in nucleus | Acts as trans-dominant inhibitor of PARP-1; irreversibly binds DNA breaks preventing repair |
| 89-kDa | Third zinc finger, BRCT, WGR, and CAT domains | Liberated from nucleus to cytosol | Has greatly reduced DNA binding capacity; may acquire novel functions in cytoplasm |
This cleavage event serves important biological functions during apoptosis. The 24-kDa fragment remains tightly bound to DNA strand breaks, acting as a trans-dominant inhibitor that blocks additional PARP-1 molecules from accessing damaged DNA and prevents DNA repair processes [27]. Meanwhile, the 89-kDa truncated PARP-1 (tPARP1) translocates to the cytoplasm where recent evidence suggests it may acquire novel functions, including mediating ADP-ribosylation of RNA polymerase III to facilitate innate immune responses during apoptosis [4].
Significance in Cell Death Pathways
PARP-1 cleavage represents a committed step in apoptotic execution, serving as a biochemical hallmark that distinguishes apoptosis from other forms of cell death. The proteolytic inactivation of PARP-1's DNA repair capacity ensures the irreversible progression of apoptosis while conserving cellular ATP pools that would otherwise be depleted by PARP-1 overactivation [27]. The detection of the 89-kDa PARP-1 fragment has become a gold standard biomarker for apoptosis in preclinical studies, providing a specific and reliable indicator of caspase-dependent cell death activation in response to therapeutic agents.
Figure 1: PARP-1 Cleavage Pathway During Apoptosis - This diagram illustrates the proteolytic cleavage of PARP-1 by effector caspases during apoptosis, generating signature fragments with distinct biological functions.
Detection Methodologies and Experimental Protocols
Western Blot Analysis for PARP-1 Cleavage
Western blot detection of the 89-kDa PARP-1 fragment remains the most widely used and validated method for assessing apoptosis in preclinical drug testing.
Protocol Details:
- Cell Lysis: Use RIPA buffer (25mM Tris-HCl pH 7.6, 150mM NaCl, 1% NP-40, 1% sodium deoxycholate, 0.1% SDS) supplemented with protease and phosphatase inhibitors. Incubate on ice for 30 minutes, then centrifuge at 14,000 × g for 15 minutes at 4°C to collect supernatant [34] [14].
- Protein Quantification: Perform BCA assay following manufacturer's instructions to normalize protein loading [14].
- Gel Electrophoresis: Load 20-50 μg of protein per lane on 4-12% Bis-Tris polyacrylamide gels. Execute electrophoresis at 120-150V for 1-2 hours using MOPS or MES running buffer [34].
- Membrane Transfer: Transfer proteins to PVDF membranes using wet or semi-dry transfer systems at 100V for 60-90 minutes at 4°C.
- Antibody Incubation: Block membranes with 5% non-fat milk in TBST for 1 hour. Incubate with primary antibodies against PARP-1 (specific for the 89-kDa fragment, such as anti-PARP-1 [C-terminal specific]) overnight at 4°C. Use appropriate secondary antibodies conjugated to HRP for 1 hour at room temperature [14].
- Detection: Develop blots using enhanced chemiluminescence substrate and image with CCD-based imaging systems.
Key Quality Controls:
- Include both untreated and positive control cells (e.g., staurosporine-treated) to verify apoptosis induction
- Probe for full-length PARP-1 to calculate cleavage ratio
- Assess actin or GAPDH as loading controls
- Use caspase inhibitors (Z-VAD-FMK) to confirm caspase-dependence [14]
Immunofluorescence and Immunohistochemistry Approaches
For spatial localization of PARP-1 cleavage within tissue sections or cultured cells:
Protocol Details:
- Sample Preparation: Fix cells with 4% paraformaldehyde for 15 minutes at room temperature. Permeabilize with 0.1% Triton X-100 in PBS for 10 minutes. Block with 5% normal serum for 1 hour [35].
- Antibody Staining: Incubate with primary antibodies specific for cleaved PARP-1 (89-kDa fragment) overnight at 4°C. Use fluorophore-conjugated secondary antibodies for 1 hour at room temperature protected from light [35].
- Counterstaining and Mounting: Counterstain nuclei with DAPI (1μg/mL) for 5 minutes. Mount with anti-fade mounting medium.
- Image Acquisition: Capture images using fluorescence or confocal microscopy. The 89-kDa fragment typically shows diffuse cytoplasmic localization compared to nuclear localization of full-length PARP-1 [4].
Quantification Methods:
- Calculate percentage of cleaved PARP-1 positive cells across multiple fields
- Determine fluorescence intensity ratios between cytoplasmic and nuclear compartments
- Use automated image analysis systems for high-throughput screening applications
Pharmacodynamic Assay Validation
The National Cancer Institute (NCI) has established rigorous validation parameters for PARP-1 cleavage assays as pharmacodynamic biomarkers in clinical trials [12]:
Table 2: Pharmacodynamic Assay Validation Parameters
| Validation Parameter | Requirements | Implementation in PARP-1 Cleavage Detection |
|---|---|---|
| Analytical Validation | Accuracy, precision, sensitivity, specificity | Demonstrate specific detection of 89-kDa fragment without cross-reactivity |
| Preclinical Modeling | Fit-for-purpose preclinical studies | Mirror clinical sampling procedures in xenograft models |
| Sample Feasibility | Assessment with human clinical samples | Optimize protein yield from core needle biopsies |
| Reproducibility | Inter-laboratory consistency | Establish standardized SOPs and control materials |
| Reagent Qualification | Lot-to-lot consistency | Quality control of antibody performance across suppliers |
| Data Interpretation | Clear reporting standards | Define thresholds for positive cleavage detection |
Critical Considerations for Clinical Implementation:
- Protein yield from 18-gauge needle biopsies is typically lower than xenograft models, requiring assay optimization [12]
- Baseline PARP-1 expression shows inter-individual variability that may impact cleavage quantification [35]
- Tissue fixation methods significantly impact antigen preservation and detection sensitivity [12]
Figure 2: Experimental Workflow for PARP-1 Cleavage Detection - This diagram outlines the key steps in detecting PARP-1 cleavage via Western blotting, from sample collection to data interpretation.
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Research Reagents for PARP-1 Cleavage Detection
| Reagent Category | Specific Examples | Function and Application |
|---|---|---|
| PARP-1 Antibodies | Anti-PARP-1 (C-terminal specific), Anti-cleaved PARP-1 (89-kDa fragment) | Detection of full-length and cleaved PARP-1 in Western blot, IHC, and IF |
| Caspase Inhibitors | Z-VAD-FMK (pan-caspase inhibitor) | Confirmation of caspase-dependent PARP-1 cleavage |
| Apoptosis Inducers | Staurosporine, Etoposide, RSL3 | Positive controls for apoptosis induction |
| Detection Systems | HRP-conjugated secondary antibodies, ECL substrates, Fluorescent secondaries | Signal generation and detection |
| Cell Lines | PC3 (prostate cancer), MCF7 (breast cancer), UWB1.289 (BRCA1-mutant ovarian) | Model systems for PARP-1 cleavage studies [35] [14] |
| PARP Inhibitors | Olaparib, DPQ, CVL218 | Experimental tools to study PARP inhibition effects [36] [37] |
Applications in Preclinical Drug Development
Assessment of Therapeutic Efficacy
PARP-1 cleavage serves as a robust biomarker for evaluating the efficacy of chemotherapeutic agents and targeted therapies in preclinical models. The detection of the 89-kDa fragment provides direct evidence of apoptosis induction, enabling researchers to:
- Establish dose-response relationships for apoptosis-inducing agents
- Determine optimal biological dosing in xenograft models
- Compare therapeutic efficacy across compound series in drug discovery
- Validate mechanism of action for targeted therapies
Recent studies have demonstrated the utility of PARP-1 cleavage detection in assessing novel therapeutic combinations, including PARP inhibitors with DNA-damaging agents or immune checkpoint inhibitors [38] [36].
Biomarker for PARP Inhibitor Resistance
Emerging evidence suggests that PARP-1 expression levels and cleavage patterns may serve as biomarkers for predicting response to PARP inhibitor therapy. Preclinical models have shown that:
- Loss of PARP-1 expression confers resistance to PARP inhibitors, detectable through reduced PARP-1 cleavage upon treatment [35]
- PARP-1 expression varies significantly across tumors, with implications for therapeutic response [35]
- Novel PET imaging agents targeting PARP-1 (e.g., [18F]FTT) enable non-invasive assessment of PARP-1 expression in vivo [35] [37]
Integration with Other Cell Death Markers
For comprehensive assessment of cell death mechanisms, PARP-1 cleavage should be evaluated alongside complementary biomarkers:
- γH2AX: Marker of DNA double-strand breaks, downstream of PARP inhibition [12]
- Caspase-3/7 Activation: Direct measurement of effector caspase activity
- Annexin V Staining: Detection of phosphatidylserine externalization during early apoptosis
- AIF Translocation: Marker of caspase-independent cell death pathways [34]
PARP-1 cleavage remains a cornerstone biomarker for apoptosis detection in preclinical drug development, providing a specific, reliable, and mechanistically informative indicator of caspase-dependent cell death. The well-characterized 89-kDa fragment serves as a validated pharmacodynamic marker across multiple therapeutic areas, particularly in oncology drug development.
Future directions in PARP-1 biomarker research include the development of more sensitive detection technologies, standardized assay protocols for multi-center trials, and integration with novel imaging approaches for non-invasive monitoring of treatment response. Additionally, growing understanding of the non-canonical functions of PARP-1 fragments, particularly the cytoplasmic roles of the 89-kDa tPARP1 in immune activation, may reveal new dimensions of this biomarker's significance in therapeutic contexts [4].
As targeted therapies continue to evolve, particularly in the DNA damage response field, PARP-1 cleavage will maintain its essential role as a key pharmacodynamic biomarker, enabling rigorous assessment of treatment efficacy and mechanism of action in preclinical drug development.
Applications in High-Throughput Screening for Apoptosis-Inducing Therapeutics
Poly (ADP-ribose) polymerase-1 (PARP-1) is a nuclear enzyme that functions as a primary sensor of DNA damage, playing a crucial role in DNA repair mechanisms and maintaining genomic integrity. The cleavage of PARP-1 by caspase family proteases, particularly caspase-3 and caspase-7, is a well-established biochemical hallmark of apoptosis [3] [39]. During apoptosis, caspases cleave the 116-kDa PARP-1 protein at a specific aspartic acid residue (D214) within the nuclear localization signal, generating two characteristic fragments: a 24-kDa DNA-binding fragment and an 89-kDa catalytic fragment [3] [31]. This cleavage event serves as a critical molecular switch that inactivates PARP-1's DNA repair function, facilitating the dismantling of the cell while conserving cellular ATP pools that would otherwise be depleted by excessive PARP-1 activation [11].
The detection of cleaved PARP-1 (cPARP-1) has become a gold standard in apoptosis research and drug discovery due to its high specificity for apoptotic cell death, its occurrence in both intrinsic and extrinsic apoptotic pathways, and the availability of highly specific antibodies that can distinguish the cleaved fragments from the full-length protein [40] [39]. This review explores the applications of PARP-1 cleavage detection in high-throughput screening (HTS) platforms for identifying and characterizing novel apoptosis-inducing therapeutics, with a focus on practical implementation, recent technological advances, and integration with complementary apoptotic markers.
PARP-1 Biology and Significance in Apoptosis
Structural Domains and Cleavage Mechanism
PARP-1 consists of three primary functional domains that dictate its cellular functions and fate during apoptosis:
- DNA-binding domain (DBD): Contains three zinc finger motifs that recognize and bind to DNA strand breaks [3] [9]. Zinc fingers 1 and 2 specifically recognize DNA damage, while zinc finger 3 facilitates inter-domain interactions essential for PARP-1 activation [9].
- Automodification domain (AMD): Contains a BRCT (BRCA1 C-terminus) fold that mediates protein-protein interactions and serves as the primary target for auto-poly(ADP-ribosyl)ation [3].
- Catalytic domain (CAT): Mediates the transfer of ADP-ribose units from NAD+ to target proteins, including PARP-1 itself [3] [9].
During apoptosis, caspase-3 cleaves PARP-1 between zinc fingers 1 and 2, specifically at the DEVD214↓G215 motif, separating the DNA-binding domain from the catalytic domain [3] [4]. This cleavage generates two primary fragments with distinct cellular fates: the 24-kDa fragment containing the first two zinc fingers remains tightly bound to DNA breaks, acting as a trans-dominant inhibitor of DNA repair, while the 89-kDa fragment translocates to the cytoplasm where it may acquire novel functions [3] [31].
Functional Consequences of PARP-1 Cleavage
The proteolytic inactivation of PARP-1 during apoptosis serves several critical biological functions:
- Energy conservation: Prevents NAD+ and ATP depletion that would occur through excessive PARP-1 activation, thereby preserving energy necessary for the ordered execution of apoptosis [11].
- DNA repair inhibition: The 24-kDa fragment acts as a dominant-negative inhibitor by occupying DNA break sites and blocking recruitment of functional DNA repair complexes [3].
- Novel signaling functions: Recent evidence indicates that the 89-kDa fragment (tPARP-1) can mediate mono-ADP-ribosylation of cytoplasmic targets, including RNA polymerase III, potentially amplifying apoptotic signaling through innate immune activation [4].
The following diagram illustrates the PARP-1 cleavage pathway and its role in apoptosis:
High-Throughput Screening Assays for PARP-1 Cleavage Detection
Immunoassay-Based Platforms
Immunoassays represent the most widely implemented approach for detecting PARP-1 cleavage in HTS formats, leveraging antibodies specific to the caspase-cleaved form of PARP-1.
Homogeneous Time-Resolved Fluorescence (HTRF) Assays HTRF technology enables quantification of cPARP-1 in cell lysates without wash steps, making it ideal for HTS applications. The assay employs a donor antibody tagged with a fluorophore (e.g., Europium cryptate) specific to the cleavage site of PARP-1 and an acceptor antibody conjugated with a compatible fluorophore (e.g., XL665) recognizing a different epitope on the same fragment. When the antibodies bind to cPARP-1 in close proximity, fluorescence resonance energy transfer (FRET) occurs, generating a quantifiable signal proportional to cPARP-1 levels.
Key advantages for HTS:
- Homogeneous format (no washing steps)
- High sensitivity with minimal interference from cellular autofluorescence
- Excellent Z' factors (>0.5) compatible with robust screening
- Compatibility with 384- and 1536-well plate formats
Electrochemiluminescence (ECL) Immunoassays ECL platforms (e.g., Meso Scale Discovery) utilize ruthenium-conjugated antibodies that emit light upon electrochemical stimulation, providing exceptional dynamic range (3-4 logs) and sensitivity for detecting cPARP-1 in small sample volumes (1-10 μL). These assays typically employ capture antibodies specific to the cPARP-1 fragment immobilized on carbon electrode-containing plates, with detection antibodies labeled with ruthenium for signal generation.
Live-Cell Imaging and High-Content Analysis
Advanced high-content screening (HCS) platforms enable multiparametric analysis of PARP-1 cleavage in intact cells, providing spatial and temporal resolution unattainable with bulk immunoassays.
Immunofluorescence-Based HCS Cells cultured in multi-well plates are treated with compounds, fixed, permeabilized, and stained with antibodies specific for cPARP-1 conjugated with fluorescent dyes (e.g., Alexa Fluor 488), often combined with nuclear stains (e.g., Hoechst 33342) and markers for other apoptotic features (e.g., Annexin V for phosphatidylserine exposure). Automated imaging systems acquire multiple fields per well, with sophisticated image analysis algorithms quantifying:
- Intensity and subcellular distribution of cPARP-1
- Nuclear morphology changes (condensation, fragmentation)
- Cell membrane integrity
- Mitochondrial membrane potential
Fluorescent Biosensor Approaches Genetically encoded biosensors enable real-time monitoring of PARP-1 cleavage in living cells, typically employing FRET-based constructs where PARP-1 cleavage sequences (DEVD) are flanked by donor (e.g., CFP) and acceptor (e.g., YFP) fluorescent proteins. Caspase-mediated cleavage separates the fluorophores, reducing FRET efficiency and providing a ratiometric signal of PARP-1 cleavage kinetics. Recent advances include:
- Baculovirus-mediated delivery for rapid sensor expression
- CRISPR/Cas9-mediated knock-in for endogenous expression
- Red-shifted FRET pairs (e.g., mCherry/mKate2) for reduced autofluorescence
Flow Cytometry-Based Screening
Although traditionally lower throughput than plate-based assays, recent technological advances have enabled HTS-compatible flow cytometry platforms (e.g., HyperCyt, Intellicyt iQue) that can analyze thousands of samples per day with multiparametric assessment of cPARP-1 at single-cell resolution. These systems typically employ:
- Automated sample aspiration from multi-well plates
- Integrated fluidics for high-speed analysis (100-1,000 cells/second)
- 6-12 parameter detection including cPARP-1, viability markers, and cell cycle status
Quantitative Assay Parameters and Validation
Key Performance Metrics for HTS Assays
Robust detection of PARP-1 cleavage in HTS requires careful optimization and validation of key assay parameters, as summarized in the table below.
Table 1: Key Performance Metrics for PARP-1 Cleavage HTS Assays
| Parameter | Target Value | Measurement Method | Significance in HTS |
|---|---|---|---|
| Z' Factor | >0.5 | (1 - (3σc+ + 3σc-)/|μc+ - μc-|) | Defines assay robustness and suitability for HTS |
| Signal-to-Background Ratio | >3:1 | Mean signal (treated)/Mean signal (untreated) | Determines assay window and hit detection confidence |
| Coefficient of Variation | <10% | (Standard deviation/Mean) × 100 | Measures precision and reproducibility |
| EC50 Concordance | ±0.3 log units | Comparison with reference compounds | Validates biological relevance of detected activity |
| DMSO Tolerance | Up to 1% | Signal measurement with DMSO present | Ensures compatibility with compound libraries |
Experimental Protocols for PARP-1 Cleavage Detection
Protocol 1: HTRF cPARP-1 Assay for HTS This protocol is adapted from commercially available HTRF kits (e.g., CisBio cPARP-1 HTRF Assay) and has been validated for 384-well format screening.
Materials:
- cPARP-1 HTRF Kit (anti-cPARP-1-d2 and anti-cPARP-1-Eu Cryptate antibodies)
- Cell lysis buffer (provided in kit or 50 mM HEPES, pH 7.4, 150 mM NaCl, 1% Triton X-100, 1 mM DTT)
- White, low-volume 384-well plates
- HTS-compatible liquid handling system
- HTRF-compatible plate reader
Procedure:
- Plate cells in 384-well plates at optimized density (typically 2,000-5,000 cells/well in 20 μL medium) and incubate overnight.
- Add test compounds using pintool transfer or acoustic dispensing; include controls (untreated cells for background, 1-10 μM staurosporine for maximum cleavage).
- Incubate for predetermined optimal time (typically 4-24 hours based on cell type and mechanism).
- Add 10 μL lysis buffer with HTRF antibodies and incubate for 2 hours at room temperature protected from light.
- Read HTRF signal using appropriate filters (excitation: 320 nm, emission: 615 nm and 665 nm).
- Calculate ratio: (Signal 665 nm / Signal 615 nm) × 10,000.
Protocol 2: High-Content Analysis of cPARP-1 This protocol enables multiparametric analysis of PARP-1 cleavage with subcellular resolution.
Materials:
- Black-walled, clear-bottom 384-well plates
- Cell line appropriate for assay (e.g., HeLa, HepG2, or relevant cancer models)
- Fixation buffer (4% paraformaldehyde in PBS)
- Permeabilization buffer (0.1% Triton X-100 in PBS)
- Blocking buffer (3% BSA in PBS)
- Primary antibody: anti-cPARP-1 (e.g., Abcam, Cell Signaling Technology)
- Secondary antibody: Alexa Fluor 488-conjugated anti-rabbit IgG
- Nuclear stain: Hoechst 33342 (1 μg/mL)
- Automated imaging system (e.g., ImageXpress, Operetta, Incucyte)
Procedure:
- Seed cells at optimized density (1,000-3,000 cells/well) and incubate for 24 hours.
- Treat with compounds for predetermined time (typically 6-48 hours).
- Fix cells with 4% PFA for 20 minutes at room temperature.
- Permeabilize with 0.1% Triton X-100 for 10 minutes.
- Block with 3% BSA for 1 hour.
- Incubate with primary anti-cPARP-1 antibody (1:500-1:1,000) overnight at 4°C.
- Incubate with secondary antibody (1:1,000) for 1 hour at room temperature protected from light.
- Stain nuclei with Hoechst 33342 for 10 minutes.
- Image using 10x or 20x objective, acquiring multiple fields per well.
- Analyze images using integrated intensity and morphology algorithms.
Integration with Complementary Apoptosis Markers
While PARP-1 cleavage serves as a specific apoptosis marker, comprehensive HTS campaigns benefit from multiplexing with complementary apoptotic markers to enhance confidence in hit identification and provide mechanistic insights.
Table 2: Multiplexed Apoptosis Markers for HTS
| Marker | Detection Method | Biological Significance | Advantages for Multiplexing |
|---|---|---|---|
| Caspase-3/7 Activity | Fluorogenic substrates (e.g., DEVD-AMC) | Direct measurement of executioner caspase activation | Early apoptosis marker; compatible with live-cell formats |
| Mitochondrial Membrane Potential | JC-1, TMRM, or MitoTracker dyes | Indicator of intrinsic apoptotic pathway activation | Distinguishes intrinsic vs. extrinsic apoptosis pathways |
| Phosphatidylserine Externalization | Annexin V-FITC/PI staining | Early plasma membrane alteration in apoptosis | Distinguishes early vs. late apoptosis; flow cytometry compatible |
| DNA Fragmentation TUNEL assay | Direct detection of DNA strand breaks | Late apoptotic marker | High specificity; complements PARP-1 cleavage detection |
| Nuclear Morphology | High-content imaging with DNA stains | Characteristic apoptotic nuclear changes | Provides spatial context; distinguishes apoptosis from necrosis |
The following workflow diagram illustrates a comprehensive HTS approach integrating PARP-1 cleavage detection with complementary apoptosis markers:
The Scientist's Toolkit: Essential Research Reagents
Successful implementation of PARP-1 cleavage detection in HTS requires access to high-quality, well-validated reagents and tools. The following table summarizes essential research solutions for apoptosis screening campaigns.
Table 3: Research Reagent Solutions for PARP-1 Cleavage Detection
| Reagent Category | Specific Examples | Application Notes | Supplier Examples |
|---|---|---|---|
| cPARP-1 Antibodies | Anti-cleaved PARP-1 (Asp214) monoclonal; Anti-PARP-1 p25 fragment polyclonal | Validate specificity for cleaved vs. full-length PARP-1; optimize dilution for each application | Cell Signaling Technology, Abcam, Santa Cruz Biotechnology |
| HTRF Kits | cPARP-1 (Asp214) HTRF Assay Kit | Optimized for 384- and 1536-well formats; includes lysis buffer and premixed antibodies | CisBio, Revvity |
| ECL Kits | PARP-1 Cleavage Assay Kit | High dynamic range; compatible with small sample volumes | Meso Scale Discovery |
| Live-Cell Caspase Kits | CellEvent Caspase-3/7 Green Detection Reagent | Compatible with live-cell imaging; minimal cytotoxicity | Thermo Fisher Scientific |
| Flow Cytometry Kits Alexa Fluor 488 Anti-PARP-1 Cleaved Form Specific Antibody | Compatible with intracellular staining protocols; optimized for flow cytometry | BD Biosciences, BioLegend | |
| Positive Controls | Staurosporine, Camptothecin, Etoposide | Induce robust apoptosis and PARP-1 cleavage; establish assay window | Sigma-Aldrich, Tocris |
| Cell Lines | HeLa, HepG2, Jurkat, PC-3 | Well-characterized apoptotic responses; suitable for HTS | ATCC, Sigma-Aldrich |
Recent Advances and Future Perspectives
Emerging Technologies in PARP-1 Detection
Single-Cell Western Blotting Microfluidic platforms enabling single-cell resolution western blotting (e.g., Milo from ProteinSimple) permit simultaneous detection of PARP-1 cleavage and other apoptotic markers in hundreds of individual cells, revealing heterogeneous responses to therapeutic compounds that would be masked in population-averaged measurements.
Multiplexed Bead-Based Assays Luminex xMAP technology and related platforms enable simultaneous quantification of cPARP-1 alongside dozens of other apoptotic and signaling proteins from minimal sample volumes (10-25 μL), providing comprehensive pathway activation signatures for mechanism-of-action studies.
CRISPR/Cas9-Mediated Endogenous Tagging Precise knock-in of fluorescent tags (e.g., GFP, HALO) into the endogenous PARP-1 locus enables monitoring of PARP-1 cleavage without overexpression artifacts, with recent demonstrations showing superior performance compared to traditional overexpression approaches [41].
Application in Targeted Therapy Development
PARP-1 cleavage detection has found particular utility in screening for compounds that synergize with PARP inhibitors (PARPi) in BRCA-deficient cancers, with recent studies employing HTS approaches to identify novel sensitizers that enhance PARPi efficacy through induction of apoptotic cell death [41]. Additionally, the discovery of non-apoptotic functions for truncated PARP-1 fragments, including roles in innate immune activation through RNA polymerase III ADP-ribosylation, has opened new avenues for therapeutic intervention that may be leveraged in future screening campaigns [4].
The detection of PARP-1 cleavage remains a cornerstone in high-throughput screening for apoptosis-inducing therapeutics, offering high specificity, well-established detection methodologies, and relevance across diverse therapeutic areas including oncology, neurodegenerative diseases, and cardiovascular disorders. Integration of PARP-1 cleavage detection with complementary apoptotic markers in multiplexed HTS platforms provides a powerful approach for identifying novel therapeutic candidates with robust apoptotic activity, while emerging technologies in single-cell analysis and endogenous tagging promise to further enhance the depth and quality of information obtained from screening campaigns. As our understanding of PARP-1 biology continues to evolve, including recent discoveries of novel functions for cleavage fragments, the applications of PARP-1 cleavage detection in drug discovery will continue to expand and refine our ability to identify effective apoptosis-modulating therapeutics.
Flow Cytometry Approaches for Single-Cell Analysis of PARP-1 Cleavage
Poly(ADP-ribose) polymerase-1 (PARP-1) is a 116 kDa nuclear enzyme that plays a critical role in DNA repair and the maintenance of genomic integrity. During the early stages of apoptosis, caspase-3 and -7 cleave PARP-1 at a specific aspartic acid residue (Asp214), generating 24 kDa and 89 kDa fragments. This cleavage event serves as a well-established biochemical marker for apoptosis, as it inactivates PARP-1's DNA repair function and facilitates cellular disassembly. Flow cytometry enables robust, single-cell analysis of this key apoptotic event, allowing researchers to quantify the percentage of cells undergoing apoptosis within a heterogeneous population. This technical guide details optimized flow cytometry methodologies for detecting PARP-1 cleavage, complete with structured experimental data, essential reagents, and visual workflows to support research and drug development in cell death mechanisms.
PARP-1 is a ubiquitously expressed nuclear enzyme that functions as a molecular sensor for DNA strand breaks. Its primary role involves facilitating DNA repair through the synthesis of poly(ADP-ribose) (PAR) chains on target proteins. During the execution phase of apoptosis, activated effector caspases (primarily caspase-3) cleave PARP-1 at the conserved Asp214-Gly215 sequence [42] [43]. This proteolytic event separates the N-terminal DNA-binding domain (24 kDa) from the C-terminal catalytic domain (89 kDa), resulting in the inactivation of the enzyme's DNA repair capacity [15] [44]. The detection of this 89 kDa fragment has become a gold standard method for identifying apoptotic cells, as it represents a committed step in the dismantling of the cell.
The biological significance of this cleavage is twofold. First, it prevents PARP-1 from exhausting cellular energy reserves (NAD+ and ATP) in a futile attempt to repair extensively damaged DNA, thereby facilitating the apoptotic process [15]. Second, emerging evidence suggests that the cleavage fragments themselves may have active roles in modulating cell death and inflammatory responses [45]. For instance, the 89 kDa fragment can be translocated to the cytoplasm in vesicles, where it may participate in non-nuclear signaling events [46]. The specificity of PARP-1 cleavage by caspases makes it an exceptionally reliable marker for distinguishing apoptosis from other forms of cell death.
Flow Cytometry Methodologies for Detecting Cleaved PARP
Direct Staining Protocol for Fixed and Permeabilized Cells
The detection of intracellular cleaved PARP (cPARP) by flow cytometry requires antibodies specific to the neo-epitope exposed after caspase cleavage at Asp214. The following protocol, adapted from commercial reagent providers and research publications, ensures specific detection of the 89 kDa fragment [42] [47].
Sample Preparation:
- Induction of Apoptosis: Treat cells (e.g., Jurkat T-cells) with a known apoptosis inducer. Camptothecin (4-6 µM for 4 hours) is commonly used as it inhibits topoisomerase I, causing DNA damage and triggering apoptosis [42]. Alternatively, staurosporine (10 µM) or anti-Fas antibodies can be used [40] [47].
- Cell Harvesting and Washing: Collect approximately 1 × 10^6 cells per sample. Wash cells twice with cold phosphate-buffered saline (PBS) to remove serum proteins.
Fixation and Permeabilization:
- Fixation: Resuspend cell pellets in Cytofix/Cytoperm solution (BD Biosciences) and incubate for 20 minutes on ice. This step preserves intracellular structures and epitopes.
- Permeabilization and Washing: Wash cells twice with 1× Perm/Wash Buffer (BD Biosciences) to remove fixative and create pores in the membrane for antibody access.
Antibody Staining:
- Staining: Resuspend fixed and permeabilized cells in Perm/Wash Buffer at a concentration of 1 × 10^7 cells/mL. Aliquot 100 µL (1 × 10^6 cells) per test tube. Add the recommended volume of fluorochrome-conjugated anti-cleaved PARP antibody (e.g., 20 µL of PE Mouse Anti-Cleaved PARP [Asp214] or 5 µL of the eBioscience HLNC4 clone) [42] [47].
- Incubation: Incubate cells with antibody for 30 minutes at room temperature, protected from light.
- Final Wash and Resuspension: Wash cells once with Perm/Wash Buffer to remove unbound antibody. Resuspend the final cell pellet in 0.5 mL of Perm/Wash Buffer or a standard flow cytometry staining buffer for acquisition.
Controls: Always include an untreated control sample and an isotype-matched control antibody to establish background fluorescence and define positive populations.
Multiparameter Panel Design for Apoptosis Detection
A significant advantage of flow cytometry is the ability to perform multiparameter analysis. Cleaved PARP detection can be combined with other markers to gain a more comprehensive understanding of cell death dynamics. A typical panel might include:
- Viability Dye: A fixable viability dye (e.g., Fixable Viability Dye eFluor 780) to exclude necrotic cells from the analysis [47].
- Early Apoptosis Marker: Annexin V to detect phosphatidylserine externalization [47].
- Other Caspase Targets: Antibodies for other caspase cleavage targets, such as active caspase-3, which can be detected simultaneously using different fluorochromes [48].
When designing a panel, ensure fluorochrome conjugates are compatible with your flow cytometer's laser and filter configuration. For instance, PE-conjugated anti-cleaved PARP (Ex Max 496/566 nm, Em Max 576 nm) is excited by blue (488 nm), green (532 nm), or yellow-green (561 nm) lasers and detected with a ~575/26 nm bandpass filter [42].
Quantitative Data from Model Systems
Flow cytometry allows for the precise quantification of the percentage of cells positive for cleaved PARP under various experimental conditions. The following table consolidates key quantitative findings from published studies utilizing flow cytometry for cPARP detection.
Table 1: Quantification of Cleaved PARP-Positive Cells in Various Experimental Models
| Cell Type/Model | Treatment | Incubation Time | % cPARP+ Cells (Mean ± SD or SEM) | Citation |
|---|---|---|---|---|
| Ram Spermatozoa | Untreated (0 h control) | 0 hours | 21.4 ± 3.3% | [40] |
| Staurosporine (10 µM) or Betulinic Acid (200 µM) | 0 hours | 44.3 ± 1.4% | [40] | |
| Untreated (4 h control) | 4 hours | 44.3 ± 2.4% | [40] | |
| Staurosporine (10 µM) or Betulinic Acid (200 µM) | 4 hours | 53.3 ± 1.4% | [40] | |
| Ram Spermatozoa (Density Gradient Interface) | Untreated | Not specified | 36.2 ± 2.0% | [40] |
| Ram Spermatozoa (Density Gradient Pellet) | Untreated | Not specified | 28.5 ± 1.2% | [40] |
| Human Jurkat T-Cells | Camptothecin (topoisomerase inhibitor) | 4 hours | Significant increase (demonstrated via histogram shift) | [42] |
| Human Jurkat T-Cells | Anti-human CD95 (Fas) antibody | 20 hours | Significant increase (demonstrated via histogram shift) | [47] |
The data in Table 1 highlights several key points. First, the baseline level of cPARP can vary between cell types and even between subpopulations, as seen in the different fractions of ram spermatozoa [40]. Second, apoptosis inducers like staurosporine and camptothecin cause a significant and measurable increase in the percentage of cPARP-positive cells [40] [42]. Finally, the duration of treatment is a critical factor, as evidenced by the increase in cPARP in untreated ram spermatozoa from 0 to 4 hours, suggesting spontaneous apoptosis in culture over time [40].
The Scientist's Toolkit: Essential Reagents
Successful detection of PARP-1 cleavage relies on specific, high-quality reagents. The following table lists critical antibodies and kits commonly used in flow cytometry, as cited in the literature and commercial sources.
Table 2: Essential Research Reagents for Cleaved PARP Flow Cytometry
| Reagent Name | Specificity / Clone | Conjugate | Key Feature | Application / Function | Citation |
|---|---|---|---|---|---|
| PE Mouse Anti-Cleaved PARP (Asp214) | Human cleaved PARP (89 kDa fragment); Clone F21-852 | R-PE | Does not recognize full-length PARP-1. | Intracellular staining for flow cytometry to specifically detect apoptotic cells. | [42] |
| Cleaved PARP (Asp214) (D64E10) Rabbit mAb | Human cleaved PARP (89 kDa fragment); Clone D64E10 | Pacific Blue | Specific for the large fragment (89 kDa) produced by caspase cleavage. | Direct intracellular staining for flow cytometry; compatible with blue laser (405 nm). | [43] |
| PARP1 (cleaved Asp214) mAb (HLNC4) | Human cleaved PARP (85 kDa fragment); Clone HLNC4 | PE | Specifically recognizes the 85 kDa fragment, not full-length PARP1. | Intracellular staining followed by flow cytometric analysis. | [47] |
| PARP (46D11) Rabbit mAb | Total PARP-1 (full-length and 89 kDa fragment); Clone 46D11 | Unconjugated | Detects both full-length PARP and the cleaved 89 kDa fragment. | Western blotting to confirm cleavage; not for flow (included for validation context). | [44] |
| Cytofix/Cytoperm Kit | N/A | N/A | Provides optimized fixative and permeabilization solution. | A kit for cell fixation and permeabilization, a critical step for intracellular staining. | [42] |
When selecting a reagent, consider the species reactivity, the specificity for the cleaved form, the fluorochrome conjugate, and its compatibility with other dyes in a multicolor panel. Antibodies like F21-852 and D64E10 are particularly valuable because their specificity for the cleaved fragment minimizes background signal from healthy cells expressing full-length PARP-1 [42] [43].
Visualizing the Workflow and Signaling Pathway
Molecular Pathway of PARP-1 Cleavage in Apoptosis
The following diagram illustrates the key molecular events leading to PARP-1 cleavage and its role in the apoptotic pathway, integrating canonical and emerging mechanisms.
Experimental Workflow for Flow Cytometry
The diagram below outlines the step-by-step experimental procedure for detecting cleaved PARP via flow cytometry, from cell preparation to data analysis.
Flow cytometry provides a powerful and quantitative platform for the single-cell analysis of PARP-1 cleavage, a definitive marker of apoptosis. The methodologies outlined in this guide, from robust staining protocols to multiparameter panel design, empower researchers to accurately detect and quantify this critical cellular event. The consistent availability of highly specific antibodies against the caspase-cleaved fragment of PARP-1 further standardizes this approach across laboratories. As research continues to uncover the complex roles of PARP-1 and its fragments in cell death, inflammation, and disease, flow cytometry will remain an indispensable tool for validating findings, screening novel therapeutics, and advancing our fundamental understanding of apoptotic pathways in health and disease.
Resolving Experimental Challenges: Optimization and Interpretation of PARP-1 Cleavage Data
Poly(ADP-ribose) polymerase-1 (PARP-1) cleavage is a well-established biochemical hallmark of apoptosis, serving as a critical indicator of caspase-3 and -7 activation in programmed cell death. This technical guide examines the principal methodological challenges in accurately detecting PARP-1 cleavage, focusing on three major pitfalls: incomplete cleavage, non-specific antibody reactivity, and degradation artifacts. Within the context of apoptosis research, proper interpretation of PARP-1 cleavage fragments—the 24 kDa DNA-binding fragment and the 89 kDa catalytic fragment—is essential for understanding cell death mechanisms in cancer therapy and neurodegenerative diseases. This whitepaper provides detailed protocols for mitigating these experimental challenges, standardized methodologies for consistent results, and visual guides for pathway analysis and experimental workflows, enabling researchers to improve assay precision in both basic research and drug development applications.
Poly(ADP-ribose) polymerase-1 (PARP-1) is a 116 kDa nuclear enzyme that plays a fundamental role in DNA damage repair and maintenance of genomic stability [49] [50]. During the early stages of apoptosis, PARP-1 serves as a primary substrate for executioner caspases-3 and -7, which cleave the protein at a specific DEVD²¹⁴G motif located between its DNA-binding domain and automodification domain [28]. This proteolytic event generates two characteristic fragments: a 24 kDa N-terminal fragment containing the DNA-binding domain and a 89 kDa C-terminal fragment containing the automodification and catalytic domains [49] [6].
The cleavage of PARP-1 is considered a hallmark of apoptosis for several reasons. First, it represents a definitive biochemical marker of caspase activation, one of the most specific proteolytic events in the apoptotic cascade [15]. Second, the functional consequence of this cleavage—inactivation of PARP-1's DNA repair function—facilitates the apoptotic process by conserving cellular ATP pools that would otherwise be depleted by excessive PARP-1 activation and prevents the repair of apoptotic DNA fragmentation [49] [51]. Third, the appearance of the 89 kDa fragment in particular serves as a reliable indicator of commitment to apoptotic cell death [6].
Beyond its role as a simple marker, research has revealed that the PARP-1 cleavage fragments may have active and complex roles in cell death pathways. The 89 kDa fragment has been shown to translocate to the cytoplasm and facilitate apoptosis-inducing factor (AIF)-mediated DNA fragmentation, creating a bridge between caspase-dependent apoptosis and parthanatos, a caspase-independent programmed cell death pathway [6]. Meanwhile, the 24 kDa fragment remains bound to DNA breaks, acting as a trans-dominant inhibitor of any remaining full-length PARP-1 and further suppressing DNA repair [49] [28].
In cancer research and therapeutic development, PARP-1 cleavage monitoring provides crucial information about drug mechanisms and efficacy, particularly for agents designed to induce apoptotic cell death in tumors [20] [50]. The accurate detection and interpretation of PARP-1 cleavage patterns is therefore essential for proper assessment of treatment response and understanding resistance mechanisms.
PARP-1 Cleavage Fundamentals and Significance
Molecular Mechanism of Cleavage
PARP-1 cleavage occurs through a highly specific proteolytic mechanism during apoptosis. Executioner caspases-3 and -7 recognize and cleave the DEVD²¹⁴G amino acid sequence in PARP-1, which is situated within a nuclear localization signal near the DNA-binding domain [6] [28]. This cleavage site strategically separates the functional domains of the protein, with the 24 kDa fragment (amino acids 1-214) containing two zinc finger motifs that facilitate DNA binding, and the 89 kDa fragment (amino acids 215-1014) encompassing the automodification domain (AMD) and catalytic domain (CD) responsible for poly(ADP-ribose) synthesis [49].
The structural consequences of this cleavage are profound. The 24 kDa fragment retains the nuclear localization signal and exhibits high affinity for DNA strand breaks, where it remains tightly bound during apoptosis [49] [28]. Interestingly, this fragment acts as a trans-dominant inhibitor of intact PARP-1 by occupying DNA damage sites and blocking access by full-length PARP-1 molecules [49]. Meanwhile, the 89 kDa fragment, which loses the nuclear localization signal, translocates to the cytoplasm where it can participate in additional cell death signaling pathways [6].
Functional Consequences in Apoptosis
The biological significance of PARP-1 cleavage extends beyond merely inactivating the DNA repair function. While initial theories suggested that cleavage primarily conserved cellular energy by preventing NAD+ and ATP depletion [51], contemporary research reveals more complex roles:
- DNA Repair Disruption: Cleavage terminates ongoing DNA repair processes, facilitating the preservation of DNA damage signals that promote apoptotic execution [49].
- Fragment-Mediated Signaling: The 89 kDa fragment can serve as a carrier for poly(ADP-ribose) (PAR) polymers to the cytoplasm, where they participate in AIF release from mitochondria, bridging caspase-dependent and independent death pathways [6].
- Transcriptional Regulation: PARP-1 cleavage fragments differentially influence NF-κB activity, with the 89 kDa fragment enhancing pro-inflammatory responses that may contextualize cell death within tissue environments [28].
Table 1: PARP-1 Cleavage Fragments and Their Characteristics
| Fragment | Size | Domains Contained | Cellular Localization After Cleavage | Primary Functions |
|---|---|---|---|---|
| Full-length PARP-1 | 116 kDa | DBD, AMD, CD | Nuclear | DNA damage sensing, repair initiation |
| N-terminal Fragment | 24 kDa | DNA-binding domain (DBD) with two zinc fingers | Remains nuclear, bound to DNA | Trans-dominant inhibitor of PARP-1; blocks DNA repair |
| C-terminal Fragment | 89 kDa | Automodification domain (AMD), Catalytic domain (CD) | Translocates to cytoplasm | May carry PAR to cytoplasm; facilitates AIF release |
This sophisticated mechanism explains why PARP-1 cleavage has become such a valuable biomarker not only for detecting apoptosis but also for understanding the functional progression of cell death in experimental systems and therapeutic contexts.
Core Pitfalls and Methodological Challenges
Incomplete Cleavage
Incomplete PARP-1 cleavage represents a significant interpretative challenge that can lead to false negative conclusions or underestimation of apoptotic activity. This phenomenon occurs when a mixture of full-length PARP-1 and its cleavage fragments is detected, creating ambiguity in data interpretation.
Causes and Contexts:
- Suboptimal Caspase Activation: Treatments that induce weak or transient caspase activation may result in partial rather than complete PARP-1 cleavage [15]. This is particularly common with low-dose chemotherapeutic agents or short exposure times.
- Cell Type-Specific Variations: Certain cell lines exhibit inherent resistance to complete PARP-1 cleavage due to expression of endogenous caspase inhibitors like XIAP or differential expression of caspase-3/-7 [28].
- Alternative Protease Activities: Calpains, cathepsins, granzymes, and matrix metalloproteinases can generate PARP-1 fragments of different sizes that may interfere with clear interpretation of caspase-specific cleavage [49].
Detection Challenges: The simultaneous presence of 116 kDa (full-length), 89 kDa, and 24 kDa bands on western blots requires careful quantification and normalization. Researchers often underestimate the significance of faint cleavage fragments in the presence of strong full-length signals, potentially missing biologically relevant apoptotic activity.
Non-Specific Antibodies
Antibody quality represents one of the most variable factors in PARP-1 cleavage detection, with non-specific reactivity leading to both false positives and false negatives.
Common Antibody-Related Issues:
- Cross-Reactivity with PARP Family Members: The PARP superfamily includes 17 members, many of which share structural homology with PARP-1 [49]. Poorly validated antibodies may detect PARP-2 (62 kDa) or tankyrases, creating confusing banding patterns.
- Differential Affinity for Fragments: Some antibodies recognize the full-length protein effectively but have reduced affinity for cleavage fragments, particularly the 24 kDa fragment, leading to underestimation of cleavage efficiency [28].
- Lot-to-Lot Variability: Commercial antibodies often demonstrate significant batch variations that can alter assay performance without researcher awareness.
Validation Gaps: Many laboratories rely solely on molecular weight markers without including appropriate controls to verify antibody specificity. The 89 kDa fragment is particularly challenging to distinguish from other cellular proteins of similar size, while the 24 kDa fragment can be confused with non-specific degradation products or unrelated proteins.
Degradation Artifacts
Proteolytic degradation during sample preparation represents a critical technical challenge that can generate bands mimicking authentic cleavage fragments.
Sources of Artifactual Degradation:
- Improper Sample Handling: Extended processing times, repeated freeze-thaw cycles, or inadequate inhibition of non-apoptotic proteases during cell lysis can produce non-specific degradation [15].
- Cellular Stress Conditions: Experimental conditions that induce significant cellular stress (e.g., nutrient deprivation, hypoxia) may activate non-apoptotic proteases that generate PARP-1 fragments of sizes similar to caspase cleavage products [49].
- Post-Lysis Proteolysis: Incomplete inhibition of calpains, cathepsins, or other abundant cellular proteases during protein extraction can continue to degrade PARP-1 after cell lysis.
Differentiating Authentic Cleavage: Genuine apoptotic cleavage produces precise 89 kDa and 24 kDa fragments in a consistent ratio, while non-specific degradation typically creates a smear or multiple bands of varying sizes. The 24 kDa fragment is particularly stable due to its compact structure and DNA-binding capacity, while non-specific degradation often produces smaller fragments [49].
Table 2: Troubleshooting PARP-1 Cleavage Detection Pitfalls
| Pitfall | Key Indicators | Validation Methods | Impact on Data Interpretation |
|---|---|---|---|
| Incomplete Cleavage | Simultaneous presence of full-length and fragment bands; variation between replicates | Caspase activity assays; time-course experiments | Underestimation of apoptotic extent; incorrect timing conclusions |
| Non-Specific Antibodies | Bands at unexpected molecular weights; multiple extra bands | Knockdown controls; comparison with multiple antibodies | False positive identification of cleavage; misidentification of fragments |
| Degradation Artifacts | Smearing bands; fragments smaller than 24 kDa; lot-to-lot variability | Sample processing controls; protease inhibitor testing | False positive apoptosis detection; overestimation of cell death |
These methodological challenges underscore the necessity for rigorous controls and validation procedures when utilizing PARP-1 cleavage as an apoptosis marker in both research and drug development settings.
Experimental Protocols for Reliable Detection
Standardized Western Blot Protocol
Sample Preparation:
- Harvest cells at appropriate density (80-90% confluence recommended) using gentle scraping rather than trypsinization to prevent mechanical stress.
- Lyse cells in pre-chilled RIPA buffer supplemented with comprehensive protease inhibitors: 1 mM PMSF, 10 μg/mL aprotinin, 10 μg/mL leupeptin, 1 mM sodium orthovanadate, and 10 mM sodium fluoride.
- Critical Step: Include 10 μM caspase-3 inhibitor DEVD-CHO in control samples to distinguish caspase-specific cleavage from non-specific degradation [6] [28].
- Incubate lysates on ice for 30 minutes with occasional vortexing, followed by centrifugation at 14,000 × g for 15 minutes at 4°C.
- Determine protein concentration using BCA assay and adjust samples to equal concentrations with lysis buffer.
Electrophoresis and Transfer:
- Load 20-50 μg total protein per lane on 4-12% Bis-Tris gradient gels for optimal resolution of both high and low molecular weight fragments.
- Include molecular weight markers and positive controls (etoposide-treated Jurkat cell extracts) on every gel.
- Transfer to PVDF membrane using wet transfer system at 100V for 70 minutes at 4°C with continuous cooling.
Immunodetection:
- Block membranes with 5% non-fat dry milk in TBST for 1 hour at room temperature.
- Incubate with primary antibodies against PARP-1 overnight at 4°C with gentle agitation. Use validated antibodies recognizing both N-terminal (for 24 kDa fragment) and C-terminal epitopes (for 89 kDa fragment) [28].
- Antibody Validation: Include PARP-1 knockout cell lysates or siRNA knockdown controls to confirm specificity.
- Perform appropriate secondary antibody incubation and develop with enhanced chemiluminescence substrate.
- Always reprobe membranes with loading control antibodies (e.g., GAPDH, histone H3) to normalize for protein loading variations.
Controls and Validation Experiments
Essential Control Conditions:
- Induced Apoptosis Positive Control: Treat cells with 50 μM etoposide or 1 μM staurosporine for 6-16 hours to generate robust PARP-1 cleavage [6].
- Caspase Inhibition Control: Pre-treat cells with 20 μM pan-caspase inhibitor Z-VAD-FMK for 2 hours before apoptosis induction to confirm caspase-dependent cleavage.
- PARP-1 Specificity Control: Include samples from PARP-1 deficient cells or use siRNA-mediated knockdown to verify antibody specificity.
Quantification and Analysis:
- Capture multiple exposures of western blots to ensure linear detection of both strong and weak signals.
- Quantify band intensities using image analysis software and calculate cleavage ratio as: (89 kDa intensity) / (89 kDa + 116 kDa intensities) × 100.
- Normalize data to loading controls and express as fold-change relative to untreated controls.
- Perform statistical analysis on at least three independent biological replicates.
Complementary Assays for Verification
Caspase Activity Measurement:
- Complement PARP-1 western analysis with fluorometric caspase-3/7 activity assays using DEVD-AFC or DEVD-AMC substrates.
- Monitor cleavage kinetics in real-time to establish temporal relationship between caspase activation and PARP-1 cleavage.
Cell Death Detection:
- Correlate PARP-1 cleavage data with Annexin V/propidium iodide staining by flow cytometry.
- Assess morphological changes associated with apoptosis (nuclear condensation, membrane blebbing) to contextualize biochemical findings.
This comprehensive methodological approach ensures reliable detection and accurate interpretation of PARP-1 cleavage while controlling for the major pitfalls discussed in this guide.
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for PARP-1 Cleavage Research
| Reagent/Category | Specific Examples | Function/Application | Technical Considerations |
|---|---|---|---|
| PARP-1 Antibodies | Anti-PARP-1 (C-terminal specific), Anti-PARP-1 (N-terminal specific), Cleaved PARP-1 (Asp214) antibodies | Detection of full-length and cleavage fragments; confirmation of specific cleavage event | Validate using PARP-1 knockout cells; use multiple epitopes for confirmation |
| Caspase Inhibitors | Z-VAD-FMK (pan-caspase), DEVD-CHO (caspase-3/7 specific), Q-VD-OPh (broad-spectrum) | Distinguish caspase-dependent cleavage; confirm mechanism | Use cell-permeable forms for pretreatment; include in lysis buffer for some applications |
| Apoptosis Inducers | Staurosporine, Etoposide, Actinomycin D, TNF-α with cycloheximide | Generate positive controls for PARP-1 cleavage | Titrate for cell type-specific response; establish time courses |
| Protease Inhibitor Cocktails | PMSF, Aprotinin, Leupeptin, Pepstatin A, EDTA | Prevent non-specific protein degradation during sample processing | Use fresh preparations; include calpain inhibitors (e.g., E64d) for sensitive cells |
| Cell Lines | Jurkat (lymphocyte), SH-SY5Y (neuroblastoma), HeLa (cervical carcinoma), PARP-1 -/- fibroblasts | Model systems for apoptosis studies; controls for antibody specificity | Select lines with robust apoptotic response; use low-passage stocks |
| Detection Systems | Chemiluminescent substrates, Fluorescent secondary antibodies, Near-infrared dyes | Visualization and quantification of cleavage fragments | Choose based on sensitivity needs; fluorescent methods offer wider linear range |
Signaling Pathways and Experimental Workflows
PARP-1 Cleavage in Apoptosis Signaling Pathway
PARP-1 Cleavage Detection Workflow
The detection of PARP-1 cleavage remains a cornerstone method for apoptosis assessment in biomedical research, but its utility depends entirely on rigorous methodological execution and appropriate interpretation. The pitfalls of incomplete cleavage, non-specific antibodies, and degradation artifacts represent significant challenges that require systematic approaches to overcome. By implementing the standardized protocols, validation methods, and control strategies outlined in this technical guide, researchers can significantly improve the reliability of their apoptosis data. The integration of PARP-1 cleavage analysis with complementary assays provides a comprehensive understanding of cell death mechanisms that is essential for both basic research and the development of novel therapeutic agents targeting apoptotic pathways. As research continues to reveal new dimensions of PARP-1 biology beyond its role in DNA repair, the accurate assessment of its cleavage status will remain critical for advancing our understanding of cell death in health and disease.
Optimizing Cell Lysis and Sample Preparation to Preserve Fragments
Within the landscape of programmed cell death research, the proteolytic cleavage of Poly(ADP-ribose) polymerase-1 (PARP-1) serves as a critical biochemical marker for distinguishing apoptosis from other forms of cell death. The generation of specific signature fragments, notably the 24-kDa and 89-kDa peptides, is a definitive event catalyzed by apoptotic proteases like caspases-3 and -7. Preserving these fragments during cell lysis and sample preparation is paramount for accurate experimental interpretation. This technical guide provides in-depth methodologies and optimized protocols designed to maintain the integrity of these cleavage fragments, thereby ensuring data reliability in studies of apoptosis, parthanatos, and the efficacy of PARP-targeted therapeutics.
PARP-1 is a ubiquitous nuclear enzyme with a well-established role as a molecular sensor for DNA damage. Its primary function involves facilitating DNA repair through the synthesis of poly(ADP-ribose) (PAR) chains on target proteins. During the controlled process of apoptosis, executioner caspases, primarily caspase-3 and -7, cleave the full-length 116-kDa PARP-1 protein at a specific aspartic acid residue (Asp214) [3] [11]. This proteolytic event results in the separation of the N-terminal DNA-binding domain (24-kDa fragment) from the C-terminal catalytic domain (89-kDa fragment) [3] [15].
The biological consequence of this cleavage is the effective inactivation of PARP-1's DNA repair capacity, which facilitates the apoptotic dismantling of the cell [11]. The 24-kDa fragment, which contains two zinc-finger motifs, remains tightly bound to DNA strand breaks and can act as a trans-dominant inhibitor of further PARP-1 activity, thereby conserving cellular ATP pools for the apoptotic process [3]. Consequently, the detection of these specific fragments has become a gold-standard biomarker for identifying apoptotic cells in diverse research contexts, from fundamental biology to drug development in oncology and neurodegeneration.
PARP-1 Cleavage Fragments: Signatures and Significance
The specific cleavage of PARP-1 by different proteases yields signature fragments that act as molecular fingerprints for the activation of distinct cell death pathways. The table below summarizes the key fragments and their biological significance.
Table 1: Characteristic PARP-1 Cleavage Fragments and Their Properties
| Fragment Size | Originating Protease | Domains Contained | Cellular Localization & Function |
|---|---|---|---|
| 89-kDa | Caspase-3 and -7 [3] [11] | Auto-modification Domain (AMD) and Catalytic Domain (CD) [3] | Liberated into cytosol; can act as a PAR carrier to induce AIF-mediated parthanatos [31] [10] |
| 24-kDa | Caspase-3 and -7 [3] [11] | DNA-Binding Domain (DBD) with two zinc-finger motifs [3] | Retained in nucleus; irreversibly binds damaged DNA, inhibiting repair and conserving ATP [3] |
It is crucial to note that while caspase-mediated cleavage is a hallmark of apoptosis, other "suicidal" proteases, including calpains, cathepsins, granzymes, and matrix metalloproteinases (MMPs), can also process PARP-1, generating fragments of different molecular weights [3]. This underscores the necessity of using precise detection methods, such as western blotting with well-characterized antibodies, to distinguish between these fragments and accurately assign the cell death pathway being activated.
The Role of the 89-kDa Fragment in Parthanatos
Recent research has elucidated a more complex role for the caspase-generated 89-kDa fragment. It can be poly(ADP-ribosyl)ated and subsequently translocate to the cytoplasm, where the covalently attached PAR polymers serve as a docking platform for Apoptosis-Inducing Factor (AIF) [31] [10]. This binding facilitates the release of AIF from mitochondria and its translocation to the nucleus, triggering a caspase-independent, regulated cell death known as parthanatos. This pathway illustrates a critical intersection between apoptotic and necrotic signaling and highlights the functional importance of the PARP-1 cleavage fragments beyond simply inactivating DNA repair [31].
Figure 1: PARP-1 Cleavage in Cell Death Pathways. Caspase-3/7 cleavage of PARP-1 generates fragments that contribute to both apoptosis and parthanatos.
Optimized Cell Lysis and Preparation for Fragment Preservation
Accurate analysis of PARP-1 cleavage is entirely dependent on sample preparation that instantly halts all proteolytic activity and preserves the native protein state.
Critical Considerations for Lysis Buffer
A well-formulated lysis buffer is the first and most critical defense against artifactual protein degradation.
- Protease Inhibitors: A broad-spectrum cocktail is non-negotiable. This must include potent caspase inhibitors (e.g., Z-VAD-FMK) to prevent ongoing apoptosis during handling, as well as inhibitors for calpains, cathepsins, and metalloproteinases, especially if studying non-apoptotic death [3] [11].
- Phosphatase Inhibitors: To maintain the native phosphorylation state of proteins, which can influence antibody binding and protein function.
- PARP Activity Modulators: Including PARP inhibitors (e.g., Olaparib, Veliparib) in the lysis buffer can prevent auto-poly(ADP-ribosyl)ation, which might alter protein mobility on gels and mask epitopes [26] [36].
- Chelating Agents: Use EDTA or EGTA to chelate metal ions required for the activity of metalloproteases. However, note that PARP-1's DNA-binding zinc-finger domains are sensitive to chelators, which can disrupt the protein's structure. A balance must be struck, and sample handling should be quick.
- Detergent: Use a non-ionic detergent (e.g., NP-40, Triton X-100) at a concentration of 0.5-1% to efficiently solubilize nuclear proteins without denaturing them.
Recommended Lysis Buffer Formulation
- 25 mM Tris-HCl (pH 7.4)
- 150 mM NaCl
- 1% NP-40 (or Igepal CA-630)
- 1 mM EDTA (use with caution for zinc-finger integrity)
- 1 mM EGTA
- 10% Glycerol
- Add fresh before use:
- 1 mM DTT or β-mercaptoethanol
- 1 mM PMSF
- Complete EDTA-free Protease Inhibitor Cocktail Tablet
- Phosphatase Inhibitor Cocktail (e.g., PhosSTOP)
- 10 µM Z-VAD-FMK (caspase inhibitor)
- 1 µM PARP inhibitor (e.g., Olaparib)
Step-by-Step Cell Lysis Protocol
- Pre-chill: Pre-cool all centrifuges, rotors, and tubes to 4°C. Prepare lysis buffer and keep it on ice.
- Harvest Cells: For adherent cells, gently scrape them into cold PBS. For suspension cells, collect via centrifugation. Wash cells once with ice-cold PBS.
- Lysate Preparation: Resuspend the cell pellet in cold lysis buffer (e.g., 100 µL per 1x10⁶ cells). Vortex briefly to mix.
- Incubation: Incubate on ice for 15-30 minutes with occasional gentle vortexing.
- Clarification: Centrifuge at 12,000-16,000 x g for 15 minutes at 4°C to pellet insoluble debris and genomic DNA.
- Collection: Immediately transfer the supernatant (whole cell lysate) to a fresh pre-chilled tube.
- Protein Quantification: Perform a rapid protein assay (e.g., BCA assay).
- Sample Denaturation: Mix lysate with an equal volume of 2X Laemmli sample buffer, boil at 95-100°C for 5-10 minutes, and then immediately place on ice. Aliquot and store at -80°C if not used immediately. Avoid repeated freeze-thaw cycles.
Table 2: Troubleshooting Common Issues in PARP-1 Fragment Detection
| Problem | Potential Cause | Solution |
|---|---|---|
| No cleavage fragments detected | Insufficient apoptosis; inefficient cell lysis | Include a positive control (e.g., Staurosporine-treated cells) [31]; confirm lysis efficiency. |
| High background or smearing | Sample degradation; incomplete inhibition | Use fresher protease inhibitors; shorten lysis time on ice; work faster and keep samples cold. |
| Inconsistent results between replicates | Variable lysis efficiency; uneven boiling | Ensure uniform cell resuspension; use a heat block for consistent boiling of samples. |
| Failure to detect 24-kDa fragment | Fragment is tightly bound to chromatin | Briefly sonicate the lysate after incubation or use a benchtop cup-horn sonicator to shear DNA. |
The Scientist's Toolkit: Essential Research Reagents
The following table lists key reagents essential for conducting research on PARP-1 cleavage.
Table 3: Key Reagents for PARP-1 Cleavage Research
| Reagent / Material | Function / Application | Example(s) |
|---|---|---|
| Caspase Inhibitors | Halts ongoing apoptotic cleavage during sample prep to preserve fragment ratios. | Z-VAD-FMK (pan-caspase inhibitor) [11] |
| PARP Inhibitors | Used in lysis buffer to prevent auto-modification; also as experimental therapeutics. | Olaparib, Veliparib, Talazoparib, CVL218 [26] [36] [52] |
| Apoptosis Inducers | Positive control for inducing PARP-1 cleavage in experimental systems. | Staurosporine, Actinomycin D, Etoposide [31] [11] |
| Anti-PARP-1 Antibodies | Detection of full-length and cleavage fragments via Western Blot or immunofluorescence. | Antibodies specific to the N-terminal (detects 24-kDa) or C-terminal (detects 89-kDa) regions. |
| Protease Inhibitor Cocktails | Broad-spectrum inhibition of non-caspase proteases to prevent non-specific degradation. | Commercial cocktails (e.g., from Roche, Thermo Fisher) |
| Phosphatase Inhibitor Cocktails | Preserves phosphorylation status of signaling proteins upstream of caspase activation. | Commercial cocktails (e.g., PhosSTOP) |
Experimental Workflow for PARP-1 Cleavage Analysis
A robust experimental workflow, from treatment to detection, is vital for reliable data. The following diagram outlines the key steps, with special attention to points critical for preserving fragile protein fragments and modifications.
Figure 2: Workflow for PARP-1 Cleavage Analysis. The workflow highlights critical steps performed at low temperatures and with inhibitors to preserve fragment integrity.
Detailed Methodologies for Key Experiments
Western Blot Analysis for PARP-1 Cleavage:
- Gel Electrophoresis: Use a 4-12% or 4-15% gradient SDS-PAGE gel for optimal separation of the full-length (116-kDa) and cleaved (89-kDa) PARP-1. Include a pre-stained protein ladder.
- Transfer: Perform standard wet or semi-dry transfer to a PVDF membrane.
- Blocking: Block the membrane with 5% non-fat dry milk in TBST for 1 hour at room temperature.
- Antibody Incubation:
- Primary Antibody: Use a well-validated anti-PARP-1 antibody that recognizes the C-terminal catalytic domain (to detect full-length and the 89-kDa fragment) or the N-terminal DNA-binding domain (to detect the 24-kDa fragment). Incubate overnight at 4°C with gentle agitation.
- Secondary Antibody: Incubate with an appropriate HRP-conjugated secondary antibody for 1 hour at room temperature.
- Detection: Use a high-sensitivity chemiluminescent substrate and image with a digital imager capable of capturing a wide dynamic range.
Data Interpretation: The classic apoptotic signature is the disappearance of the 116-kDa band with the concomitant appearance of the 89-kDa band. The 24-kDa fragment is often more difficult to detect due to its small size and potential loss during transfer. Quantification of the ratio of the 89-kDa fragment to the full-length PARP-1 provides a semi-quantitative measure of the extent of apoptosis in the sample.
The cleavage of PARP-1 remains one of the most reliable and informative biomarkers for apoptosis. However, the biological complexity of its fragments, particularly the emerging role of the 89-kDa fragment in parthanatos, demands rigorous technical approaches. By implementing the optimized lysis strategies and detailed protocols outlined in this guide—emphasizing speed, constant cooling, and comprehensive protease inhibition—researchers can faithfully preserve these critical proteolytic signatures. This methodological rigor is fundamental for advancing our understanding of cell death in health, disease, and in response to novel therapeutic agents like PARP inhibitors.
Poly(ADP-ribose) polymerase-1 (PARP-1) is a nuclear enzyme that plays a pivotal role in DNA damage repair and serves as a critical marker for programmed cell death. As a 116-kDa protein, PARP-1 becomes activated upon binding to DNA lesions and catalyzes the transfer of ADP-ribose units from NAD+ to target proteins, facilitating DNA repair processes [6] [53]. The cleavage of PARP-1 by caspases during apoptosis represents a fundamental event in cell death pathways, serving to conserve cellular energy and prevent DNA repair during cellular dismantling [3] [11]. This technical guide provides researchers with a comprehensive framework for accurately identifying PARP-1 cleavage events and distinguishing apoptotic cleavage from other proteolytic fragments that occur in necrosis and alternative cell death pathways.
The significance of PARP-1 cleavage as an apoptosis marker extends across multiple research domains, from basic cancer biology to drug development. The characteristic cleavage of PARP-1 by caspase-3 and -7 into specific 24-kDa and 89-kDa fragments has become a gold standard for detecting apoptotic cells in experimental models [3] [31]. However, emerging evidence reveals that PARP-1 is also a substrate for other proteases, including calpains, cathepsins, granzymes, and matrix metalloproteinases, producing distinct fragments that signify alternative cell death pathways [3]. This complexity necessitates rigorous experimental approaches to correctly interpret PARP-1 cleavage patterns within the context of a broader cell death landscape.
PARP-1 Domain Architecture and Cleavage Sites
PARP-1 possesses a modular structure consisting of three primary domains that dictate its function and proteolytic fate. The N-terminal DNA-binding domain (DBD) contains three zinc finger motifs (Zn1, Zn2, Zn3) that recognize DNA strand breaks, along with a nuclear localization signal (NLS) [9] [53]. The central automodification domain (AMD) contains a BRCT domain that facilitates protein-protein interactions and serves as the primary site for PARP-1 auto-ADP-ribosylation [3] [9]. The C-terminal catalytic domain (CAT) houses the NAD+ binding site and is responsible for synthesizing poly(ADP-ribose) chains [9].
The domain structure of PARP-1 and its cleavage sites by different proteases can be visualized in the following diagram:
Figure 1: PARP-1 domain structure and proteolytic cleavage sites. The 116-kDa PARP-1 protein contains three primary domains that give rise to specific fragments upon protease cleavage. Caspase-3/7 cleaves at the DEVD↓G site, generating 24-kDa and 89-kDa fragments with distinct cellular localizations and functions.
The caspase cleavage site resides between the DNA-binding domain and automodification domain, specifically at the DEVD214↓G215 sequence [3] [11]. Cleavage at this site separates the DNA-binding domain from the catalytic portion of the enzyme, generating the characteristic 24-kDa N-terminal fragment containing the zinc finger motifs and a 89-kDa C-terminal fragment comprising the automodification and catalytic domains [3] [6]. This specific cleavage event serves as a biochemical hallmark of apoptosis, distinguishing it from other forms of cell death that involve different PARP-1 proteolytic patterns.
Comparative Analysis of PARP-1 Cleavage Patterns
Apoptotic Versus Necrotic PARP-1 Cleavage
The differentiation between apoptotic and necrotic PARP-1 cleavage patterns is essential for accurate interpretation of cell death mechanisms. The following table summarizes the key characteristics that distinguish these processes:
Table 1: Differentiation between Apoptotic and Necrotic PARP-1 Cleavage Patterns
| Characteristic | Apoptotic Cleavage | Necrotic Cleavage |
|---|---|---|
| Primary Proteases | Caspase-3 and Caspase-7 [3] [11] | Calpains, Cathepsins [3] |
| Characteristic Fragments | 24-kDa and 89-kDa [3] [31] | Multiple atypical fragments (varying sizes) [3] |
| Energy Dependence | ATP-dependent [11] | ATP-independent [11] |
| PARP-1 Activity Pre-cleavage | Limited activation [11] | Massive overactivation [11] |
| Energy Consequences | Preserves ATP by inactivating PARP-1 [11] | Depletes ATP via PARP-1 overactivation [11] |
| Nuclear Translocation of AIF | Induced by 89-kDa fragment [6] | Direct PAR-mediated release [6] |
| DNA Fragmentation Pattern | Ordered, nucleosomal-sized fragments [3] | Random, diffuse fragmentation [3] |
PARP-1 Cleavage by Alternative Proteases
Beyond the classical apoptotic and necrotic pathways, PARP-1 serves as a substrate for several other proteases that generate distinctive cleavage fragments. These alternative cleavage events occur in specific pathological contexts and can be identified by their characteristic fragment sizes:
Table 2: PARP-1 Cleavage by Alternative Proteases and Their Pathological Contexts
| Protease | Cleavage Fragments | Associated Cell Death Pathways | Pathological Contexts |
|---|---|---|---|
| Granzyme A | ~50-kDa and ~60-kDa fragments [3] | Lymphocyte-mediated cytotoxicity [3] | Immune response, viral infection [3] |
| Matrix Metalloproteinases (MMPs) | 35-kDa and 55-kDa fragments [3] | Inflammation-associated cell death [3] | Neurodegeneration, cerebral ischemia [3] |
| Cathepsins | 35-kDa, 40-kDa, and 50-kDa fragments [3] | Lysosome-mediated cell death [3] | Trauma, excitotoxicity [3] |
| Calpains | 40-kDa and 55-kDa fragments [3] | Calcium-mediated cell death [3] | Neurodegeneration, cerebral ischemia [3] |
The diversity of PARP-1 cleavage fragments underscores its role as a molecular integrator of cell death signaling. Each protease generates signature fragments that serve as biomarkers for specific cell death programs, enabling researchers to decipher the predominant death pathway activated in particular experimental or pathological conditions [3].
Experimental Protocols for PARP-1 Cleavage Analysis
Western Blot Methodology for PARP-1 Cleavage Detection
Sample Preparation:
- Harvest cells and wash with ice-cold PBS
- Lyse cells in RIPA buffer (50 mM Tris-HCl pH 7.4, 150 mM NaCl, 1% NP-40, 0.5% sodium deoxycholate, 0.1% SDS) supplemented with protease inhibitors (e.g., PMSF, aprotinin, leupeptin) and caspase inhibitors (when analyzing non-apoptotic cleavage) [20]
- Centrifuge at 14,000 × g for 15 minutes at 4°C
- Determine protein concentration using BCA assay [20]
Gel Electrophoresis and Blotting:
- Separate 20-50 μg of total protein on 4-12% Bis-Tris gradient gels [20]
- Transfer to PVDF membranes using standard western blotting protocols
- Block membranes with 5% non-fat milk in TBST for 1 hour
Antibody Detection:
- Incubate with primary antibodies against PARP-1:
- Use appropriate HRP-conjugated secondary antibodies [20]
- Develop with enhanced chemiluminescence substrate and image
Controls:
- Include apoptosis inducers (e.g., staurosporine 1 μM, 6 hours) as positive controls for caspase-mediated cleavage [6]
- Use caspase inhibitors (z-VAD-fmk, 20-50 μM) to confirm caspase-dependent cleavage [20] [11]
- Include PARP-1 shRNA or PARP-1-deficient cells as negative controls [6]
Immunofluorescence Protocol for Subcellular Localization
Cell Staining:
- Culture cells on glass coverslips and treat with experimental conditions
- Fix with 4% paraformaldehyde for 15 minutes
- Permeabilize with 0.2% Triton X-100 in PBS for 10 minutes
- Block with 5% BSA in PBS for 1 hour
- Incubate with primary antibodies against PARP-1 (1:200-1:500) and AIF (1:200) overnight at 4°C [6]
- Use species-appropriate fluorescent secondary antibodies (1:1000) for 1 hour at room temperature
- Counterstain with DAPI (0.5 μg/mL) for 5 minutes
- Mount with anti-fade mounting medium
Analysis:
- Assess nuclear translocation of AIF as an indicator of parthanatos [6]
- Determine subcellular localization of PARP-1 fragments:
- Use confocal microscopy for high-resolution imaging
Quantitative Assessment of Cell Death Pathways
Annexin V/PI Staining:
- Harvest cells and wash with cold PBS
- Resuspend in binding buffer (10 mM HEPES, 140 mM NaCl, 2.5 mM CaCl₂, pH 7.4)
- Stain with Annexin V-FITC (1:20) and propidium iodide (0.5 μg/mL) for 15 minutes in the dark [4]
- Analyze by flow cytometry within 1 hour
- Interpret results:
- Annexin V+/PI-: Early apoptosis
- Annexin V+/PI+: Late apoptosis/secondary necrosis
- Annexin V-/PI+: Primary necrosis [4]
Caspase Activity Assay:
- Lyse cells in caspase assay buffer
- Measure cleavage of caspase-specific substrates:
- Caspase-3: DEVD-pNA (405 nm) [20]
- Caspase-7: DEVD-pNA (405 nm)
- Caspase-9: LEHD-pNA (405 nm)
- Express activity as fold-change over untreated controls
The relationship between PARP-1 cleavage and different cell death pathways can be visualized as follows:
Figure 2: PARP-1 cleavage pathways in different cell death mechanisms. The intensity and nature of DNA damage determine the activation of distinct cell death pathways involving PARP-1. Apoptosis features caspase-mediated cleavage, parthanatos involves PARP-1 overactivation and AIF translocation, while necrosis involves calpain-mediated cleavage and energy depletion.
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Research Reagents for PARP-1 Cleavage Studies
| Reagent Category | Specific Examples | Function/Application | Experimental Use |
|---|---|---|---|
| PARP Inhibitors | PJ34, ABT-888 (Veliparib), Olaparib [20] [6] | Inhibit PARP catalytic activity | Differentiate PAR-dependent and independent cell death [6] |
| Caspase Inhibitors | z-VAD-fmk (pan-caspase), z-DEVD-fmk (caspase-3 specific) [20] [11] | Inhibit caspase activity | Confirm caspase-dependent PARP-1 cleavage [11] |
| Apoptosis Inducers | Staurosporine, Actinomycin D [6] | Activate intrinsic apoptosis pathway | Positive controls for apoptotic PARP-1 cleavage [6] |
| Necrosis Inducers | H₂O₂, MNNG (N-methyl-N'-nitro-N-nitrosoguanidine) [6] [11] | Induce oxidative stress and DNA damage | Positive controls for necrotic cell death [11] |
| Primary Antibodies | Anti-PARP-1 (C-terminal), Anti-PARP-1 (N-terminal), Anti-cleaved PARP-1 (Asp214) [20] [6] | Detect full-length and cleaved PARP-1 | Western blot, immunofluorescence [20] |
| Cell Death Assays | Annexin V/PI staining, JC-1 mitochondrial membrane potential, LDH release [20] [4] | Quantify and characterize cell death | Correlate PARP-1 cleavage with cell death type [4] |
| PARP-1 Deficient Cells | PARP-1⁻⁄⁻ fibroblasts, PARP-1 shRNA knockdown [6] | Genetic PARP-1 ablation | Control for PARP-1 specific effects [6] |
Advanced Technical Considerations
Interpreting Complex Cleavage Patterns
Researchers often encounter complex PARP-1 cleavage patterns that require careful interpretation. Several technical considerations are essential for accurate analysis:
Multiple Protease Activation: In many pathological contexts, particularly neurodegeneration and ischemia, multiple proteases become activated simultaneously, generating complex PARP-1 fragment patterns [3]. To decipher these patterns, researchers should employ protease-specific inhibitors in combination (e.g., z-VAD-fmk for caspases, calpeptin for calpains, E-64 for cathepsins) to identify the contribution of each protease to the overall cleavage pattern [3].
Fragment Stability and Detection: The 24-kDa PARP-1 fragment tends to remain tightly bound to damaged DNA, potentially affecting its extraction and detection efficiency [3]. Researchers should optimize extraction protocols, potentially including benzonase treatment to release DNA-bound fragments. Additionally, the 89-kDa fragment may undergo post-translational modifications, including poly(ADP-ribosyl)ation, which can alter its electrophoretic mobility [6].
Cell-Type Specific Considerations: Different cell types exhibit variations in PARP-1 expression levels and may utilize distinct cell death pathways preferentially. Primary neurons, for example, are particularly prone to PARP-1-mediated parthanatos, while lymphocytes predominantly undergo caspase-dependent apoptosis [3]. These cell-type specific differences should inform experimental design and data interpretation.
Functional Consequences of PARP-1 Cleavage Fragments
Beyond their utility as biomarkers, PARP-1 cleavage fragments possess distinct biological activities that influence cell death progression:
The 24-kDa Fragment: This DNA-binding fragment remains nuclear and acts as a trans-dominant inhibitor of DNA repair by occupying DNA strand breaks and blocking access by intact PARP-1 and other repair factors [3]. This function ensures the irreversibility of the cell death commitment.
The 89-kDa Fragment: Recent research has revealed novel functions for this fragment beyond its initial characterization as an inactive cleavage product. The 89-kDa fragment translocates to the cytoplasm where it can serve as a carrier for poly(ADP-ribose) (PAR) polymers, facilitating AIF release from mitochondria [6]. Additionally, it can interact with and mono-ADP-ribosylate RNA polymerase III, potentially contributing to innate immune responses during apoptosis [4].
Cross-Talk Between Cell Death Pathways: Evidence indicates significant cross-talk between apoptotic and parthanatos pathways. Caspase-generated 89-kDa PARP-1 fragments can carry PAR polymers to the cytoplasm, initiating AIF-mediated DNA fragmentation that resembles parthanatos [6]. This intersection highlights the complexity of cell death execution and underscores the importance of comprehensive PARP-1 cleavage analysis.
The interpretation of PARP-1 cleavage results requires a sophisticated understanding of multiple cell death pathways and their associated proteolytic events. The characteristic 24-kDa/89-kDa cleavage pattern remains a definitive marker for caspase-dependent apoptosis, but researchers must remain vigilant for atypical cleavage fragments that indicate alternative cell death mechanisms. Through the integrated application of pharmacological inhibitors, careful antibody selection, and appropriate cellular assays, researchers can accurately differentiate between apoptotic, necrotic, and alternative PARP-1 cleavage events.
The continuing discovery of novel functions for PARP-1 cleavage fragments, particularly the non-canonical activities of the 89-kDa fragment, expands our understanding of cell death signaling networks. These advances highlight PARP-1's role not merely as a passive marker of cell death, but as an active participant in multiple cell fate decisions. As research in this field progresses, the interpretation of PARP-1 cleavage patterns will continue to provide valuable insights into physiological and pathological cell death processes, with significant implications for therapeutic development across multiple disease contexts.
Poly(ADP-ribose) polymerase-1 (PARP-1) is a nuclear enzyme that functions as a DNA damage sensor and participates in base excision repair. During apoptosis, PARP-1 is cleaved by activated caspase-3 and caspase-7 at the DEVD214↓G site within its nuclear localization signal, generating characteristic 24-kDa and 89-kDa fragments [6] [3]. This proteolytic cleavage serves as a well-established biochemical marker of apoptosis, as it inactivates PARP-1's DNA repair function and facilitates the systematic dismantling of the cell [3] [10]. The 24-kDa fragment contains the DNA-binding domain and remains tightly bound to DNA breaks, acting as a trans-dominant inhibitor of DNA repair, while the 89-kDa fragment, containing the automodification and catalytic domains, can be translocated to the cytoplasm [6] [28]. Accurate quantification of these cleavage fragments provides crucial information about the commitment to and progression of apoptotic cell death in experimental and clinical contexts.
PARP-1 Cleavage Fragments and Their Biological Significance
The specific cleavage fragments of PARP-1 serve as signatures of protease activity in programmed cell death. The table below summarizes the key fragments and their roles in apoptosis.
Table 1: Characteristic PARP-1 Cleavage Fragments in Apoptosis
| Fragment Size | Domains Contained | Subcellular Localization After Cleavage | Functional Consequences |
|---|---|---|---|
| 24 kDa | DNA-binding domain (DBD) with two zinc fingers | Remains nuclear, irreversibly bound to DNA breaks | Acts as trans-dominant inhibitor of DNA repair; conserves cellular ATP |
| 89 kDa | Automodification domain (AMD) and catalytic domain (CD) | Translocates to cytoplasm; can carry PAR polymers | Binds AIF via PAR; facilitates AIF-mediated DNA fragmentation |
The generation of these specific fragments represents a commitment point in cell death execution. The 89-kDa fragment has been shown to serve as a carrier for poly(ADP-ribose) (PAR) polymers to the cytoplasm, where it facilitates apoptosis-inducing factor (AIF) release from mitochondria and subsequent nuclear translocation, amplifying the cell death signal [6] [10]. This caspase-mediated interaction creates a bridge between classical apoptosis and parthanatos, a PARP-1-dependent form of programmed cell death [10].
Experimental Workflow for PARP-1 Cleavage Analysis
The standard methodological pipeline for detecting and quantifying PARP-1 cleavage involves sample preparation, separation, detection, and quantitative analysis as summarized below.
Table 2: Key Methodological Steps for PARP-1 Cleavage Quantification
| Experimental Phase | Key Procedures | Technical Considerations |
|---|---|---|
| Cell Treatment & Lysis | Apoptosis induction (e.g., staurosporine, actinomycin D); RIPA buffer lysis with protease inhibitors | Include positive controls (e.g., H₂O₂-treated cells); normalize cell numbers across conditions |
| Protein Separation | SDS-PAGE (8-12% gels) | Use appropriate molecular weight markers to identify full-length (116 kDa) and fragments (89 kDa, 24 kDa) |
| Immunodetection | Western transfer; blocking; incubation with primary and secondary antibodies | Optimize antibody concentrations; validate with PARP-1 knockout/depleted controls |
| Image Acquisition | Chemiluminescent/fluorescent detection; CCD-based imaging | Ensure non-saturating exposure; capture multiple exposures for linear range |
| Densitometric Analysis | Band quantification; background subtraction | Normalize to loading controls; calculate cleavage ratios |
Detailed Protocol: Western Blot Detection of PARP-1 Cleavage
Based on established methodologies in the field [54] [28], the following protocol provides reliable detection of PARP-1 cleavage fragments:
Sample Preparation: Harvest cells after apoptotic stimulation. For staurosporine-induced apoptosis, treat HeLa cells with 0.5-1 μM for 4-6 hours [6]. Lys cells in RIPA buffer (50 mM Tris-HCl, pH 7.5, 150 mM NaCl, 1% NP-40, 0.5% sodium deoxycholate, 0.1% SDS) supplemented with complete protease inhibitor cocktail. Include 1 mM PMSF to prevent non-specific proteolysis.
Protein Quantification and Separation: Determine protein concentration using BCA assay. Load 20-40 μg of total protein per lane on 4-20% gradient or 10% SDS-PAGE gels. Include pre-stained molecular weight markers and appropriate controls (untreated, apoptosis-induced, and caspase inhibitor-treated cells).
Western Blotting: Transfer proteins to PVDF membranes using wet or semi-dry transfer systems. Block membranes with 5% non-fat dry milk in TBST (Tris-buffered saline with 0.1% Tween-20) for 1 hour at room temperature.
Antibody Incubation: Incubate with primary antibodies specific for PARP-1. Recommended antibodies include:
- Anti-PARP-1 antibody (Santa Cruz Biotechnology, SC-74469) for full-length and fragments
- Anti-cleaved PARP-1 antibody (Santa Cruz Biotechnology, SC-194C1439) specific for apoptotic fragments [54] Dilute antibodies in blocking solution according to manufacturer recommendations. Incubate overnight at 4°C with gentle agitation.
Detection: After TBST washes, incubate with appropriate HRP-conjugated secondary antibodies (e.g., donkey anti-rabbit at 0.88 mg/mL) [54]. Develop with enhanced chemiluminescence substrate and image using a CCD-based system capable of quantitative analysis.
Densitometry and Normalization Strategies
Densitometric Analysis Principles
Densitometry converts band intensity into quantitative data that can be statistically analyzed. The fundamental steps include:
Background Subtraction: Apply rolling ball or local background subtraction to eliminate uneven background signal.
Band Detection and Volume Quantification: Define bands corresponding to full-length PARP-1 (116 kDa) and the 89-kDa fragment. Most analysis software (ImageJ, Image Lab, Image Studio) provides tools for automatic band detection with manual correction capability.
Normalization Approaches:
- Loading Controls: Normalize PARP-1 signals to housekeeping proteins (e.g., GAPDH, β-actin, β-tubulin)
- Total Protein Normalization: Use total protein staining (e.g., Coomassie, REVERT) as a more reliable normalization method
- Cleavage Index Calculation: Express results as the ratio of cleaved fragment to full-length PARP-1 or as percentage of total PARP-1 signal
Quantitative Data Interpretation
The following dot language script illustrates the signaling pathway and quantification strategy for PARP-1 cleavage analysis:
PARP-1 Cleavage Quantification Workflow
Statistical Considerations and Data Presentation
For robust quantification, include these statistical measures:
- Replication: Perform minimum of three independent biological replicates
- Data Transformation: Express cleavage as percentage of total PARP-1 or as fold-change relative to control
- Statistical Testing: Apply appropriate tests (t-test, ANOVA with post-hoc) based on experimental design
- Error Representation: Use standard error of mean (SEM) or standard deviation (SD) in graphical representations
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Research Reagents for PARP-1 Cleavage Analysis
| Reagent Category | Specific Examples | Function/Application |
|---|---|---|
| Apoptosis Inducers | Staurosporine (0.5-1 μM), Actinomycin D (0.5-1 μg/mL), H₂O₂ (1 mM) | Positive controls for PARP-1 cleavage induction [6] [54] |
| Caspase Inhibitors | zVAD-fmk (20-50 μM) | Pan-caspase inhibitor to confirm caspase-dependent cleavage [6] |
| PARP-1 Antibodies | Anti-PARP-1 (Santa Cruz SC-74469), Anti-cleaved PARP-1 (Santa Cruz SC-194C1439) | Detection of full-length and cleaved fragments [54] |
| Secondary Antibodies | Donkey anti-rabbit FITC-conjugated (Santa Cruz SC-2024), HRP-conjugated antibodies | Fluorescent or chemiluminescent detection [54] |
| PARP Inhibitors | PJ34, ABT-888, AG14361, Olaparib | Investigate PARP-1 function and inhibition effects [6] [55] [14] |
| Loading Controls | Anti-GAPDH, Anti-β-actin, Anti-β-tubulin | Normalization for protein loading variations |
| Detection Reagents | Enhanced chemiluminescence substrates, Fluorescent secondary antibodies | Signal generation for quantification |
Advanced Applications and Methodological Considerations
Alternative Detection Methods
While Western blotting remains the gold standard for PARP-1 cleavage quantification, several complementary approaches provide additional information:
Immunofluorescence Detection: Using cleaved PARP-1 antibodies (1 mg/mL concentration) followed by appropriate secondary antibodies (e.g., donkey anti-rabbit FITC-conjugated at 0.88 mg/mL) enables subcellular localization assessment of cleavage fragments [54]. The 89-kDa fragment translocates to the cytoplasm while the 24-kDa fragment remains nuclear [6].
Flow Cytometry: Antibodies against cleaved PARP-1 can be combined with other apoptosis markers (Annexin V, caspase activation) for multiparameter analysis at single-cell resolution.
Troubleshooting Common Challenges
Several technical challenges may affect quantification accuracy:
Multiple Cleavage Fragments: In addition to the canonical 89-kDa fragment, some systems report 55-kDa, 40-kDa, or 35-kDa fragments generated by other proteases (calpains, cathepsins, granzymes, MMPs) [3]. These should be distinguished from caspase-dependent fragments.
Cell Type-Specific Considerations: Neuronal cells and cancer cell lines may exhibit different basal PARP-1 expression levels and cleavage kinetics [28]. Primary neurons show PARP-1 cleavage fragments regulating cellular viability and inflammatory responses during ischemic challenge [28].
Incomplete Cleavage: During early apoptosis, partial cleavage may yield intermediate patterns. Use caspase inhibitors to confirm specificity.
The quantification strategies outlined here provide a robust framework for accurate assessment of PARP-1 cleavage as a specific apoptosis marker. Proper implementation of densitometry and normalization approaches ensures reliable data interpretation across experimental systems, facilitating research in cell death mechanisms and therapeutic development.
Poly(ADP-ribose) polymerase-1 (PARP-1) is a nuclear enzyme that functions as a primary DNA damage sensor and plays a crucial role in base excision repair. The cleavage of PARP-1 by caspases during apoptosis has become a well-established biochemical marker for programmed cell death, serving as a key indicator in cellular stress response pathways. This 113-kDa nuclear enzyme is specifically proteolyzed by caspase-family proteases to generate signature fragments of 89 kDa and 24 kDa, which represents a definitive commitment to the apoptotic execution phase. However, the interpretation of PARP-1 cleavage patterns demands careful consideration of contextual factors including cell type variations, the nature of the death stimulus, and temporal aspects of the cleavage process. This review examines the complex interplay of these factors that researchers must consider when utilizing PARP-1 cleavage as an apoptosis marker in experimental and drug discovery settings.
PARP-1 Structure and Domains
PARP-1 comprises three primary functional domains that dictate its activity and cleavage behavior:
- DNA-binding domain (DBD): N-terminal domain containing two zinc finger motifs that facilitate binding to DNA strand breaks
- Automodification domain (AMD): Central region containing a BRCT fold involved in protein-protein interactions
- Catalytic domain (CAT): C-terminal region responsible for poly(ADP-ribose) synthesis using NAD+ as substrate
The caspase cleavage site (DEVD214) is situated within the DBD, specifically between the second zinc finger motif and the nuclear localization signal, making it accessible to specific proteases during apoptosis.
Protease-Specific Cleavage Patterns
PARP-1 serves as a substrate for multiple proteases, with the specific cleavage pattern providing crucial information about the cell death pathway activated.
Caspase-Mediated Cleavage in Apoptosis
During apoptosis, executioner caspases (primarily caspase-3 and -7) recognize the DEVD214/G215 site in PARP-1, generating characteristic 89-kDa (containing AMD and CAT domains) and 24-kDa (DBD fragment) cleavage products. The 24-kDa fragment retains the DNA-binding capacity but exhibits dominant-negative inhibition of full-length PARP-1, potentially conserving cellular ATP pools during apoptotic execution.
Table 1: Protease-Specific PARP-1 Cleavage Patterns
| Protease | Cleavage Fragments | Cell Death Context | Functional Consequences |
|---|---|---|---|
| Caspase-3/7 | 89 kDa + 24 kDa | Apoptosis | Inactivation of PARP-1, inhibition of DNA repair, energy conservation |
| Calpain | 55-62 kDa fragments | Ca2+-mediated death | Alternative regulation of PARP-1 activity |
| Cathepsins (B/G) | 50 kDa fragment | Necrosis | Lysosomal-mediated cleavage |
| Granzyme A | Unknown | Immune-mediated killing | Alternative cleavage pattern |
| MMPs | Various fragments | Tissue remodeling | Context-dependent outcomes |
Alternative Cleavage in Non-Apoptotic Contexts
Beyond caspase-mediated cleavage, PARP-1 undergoes proteolysis by other proteases in distinct cell death contexts. During necrosis, lysosomal proteases (cathepsins B and G) generate a predominant 50-kDa fragment, which is not inhibited by broad-spectrum caspase inhibitors like zVAD-fmk. This alternative cleavage pattern provides a biochemical signature to differentiate apoptotic from necrotic cell death.
Cell Type-Specific Considerations
The regulation and consequences of PARP-1 cleavage exhibit significant variation across different cell types, influenced by developmental origin, metabolic characteristics, and specialized functions.
Neuronal Cells
In neuronal systems, PARP-1 cleavage products demonstrate unique regulatory functions. The 24-kDa fragment confers protection against oxygen/glucose deprivation damage, while the 89-kDa fragment exhibits cytotoxic properties. These cell type-specific outcomes highlight the importance of considering neuronal context when interpreting PARP-1 cleavage in cerebral ischemia, trauma, and excitotoxicity models.
Cancer Cells
PARP-1 expression and cleavage patterns in cancer cells show remarkable heterogeneity influenced by tissue origin and genetic background. PARP-1 alterations occur in approximately 2.9% of diverse tumors and correlate with higher tumor mutation burden in various cancer types. The functional impact of PARP-1 cleavage in cancer cells varies significantly based on cellular context and the specific death stimulus applied.
Table 2: Cell Type-Specific PARP-1 Cleavage Characteristics
| Cell Type | Cleavage Features | Contextual Factors | Functional Outcomes |
|---|---|---|---|
| Neurons | 24-kDa fragment is protective | Influenced by metabolic stress | Modulates inflammatory response via NF-κB |
| Cardiomyocytes | Calpain-mediated cleavage predominates | Calcium-dependent regulation | Associated with ischemia-reperfusion injury |
| Hepatocytes | Caspase and cathepsin cleavage | Metabolic specialization | Varies with toxin exposure |
| Immune Cells | Granzyme-mediated cleavage | Immune activation context | Cytotoxic lymphocyte-mediated killing |
| Epithelial Cells | Standard caspase cleavage | Tissue-specific variations | DNA damage-induced apoptosis |
Stimulus-Dependent Cleavage Activation
The nature of the cell death stimulus profoundly influences PARP-1 cleavage patterns, protease involvement, and subsequent cellular responses.
DNA-Damaging Agents
Chemotherapeutic drugs (etoposide, camptothecin) and radiation trigger PARP-1 hyperactivation followed by caspase-dependent cleavage. The intensity and duration of DNA damage determine whether cells undergo apoptosis or alternative death pathways, with PARP-1 cleavage serving as a commitment point to apoptotic execution.
Oxidative Stress
Reactive oxygen species induce PARP-1 activation through DNA strand break accumulation. The level of oxidative damage dictates the death pathway: mild stress promotes caspase-mediated cleavage, while severe damage leads to PARP-1 overactivation, NAD+ depletion, and necrotic cell death.
Metabolic Stress
In post-mortem muscle tissues, PARP-1 activation occurs in response to metabolic shifts, participating in tenderness development through cysteine protease activation. This unique context demonstrates the diverse functional roles of PARP-1 cleavage beyond classical apoptosis.
Temporal Dynamics of PARP-1 Cleavage
The timing of PARP-1 cleavage initiation and progression provides critical insights into cell death commitment and execution.
Initiation Phase
PARP-1 cleavage typically occurs after caspase-3 activation during the execution phase of apoptosis. The temporal relationship between caspase activation and PARP-1 cleavage makes it a valuable marker for apoptosis commitment, though this timing varies with stimulus intensity and cell type.
Execution and Completion
Complete PARP-1 cleavage typically correlates with irreversible commitment to apoptotic death, though residual full-length PARP-1 may persist in subpopulations. The rate of cleavage progression informs on the efficiency of cell death execution in experimental systems.
Experimental Methodologies for Detection
Western Blot Analysis
Standard methodology for detecting PARP-1 cleavage fragments:
- Sample Preparation: Cells lysed in RIPA buffer with protease inhibitors
- Gel Electrophoresis: 8-12% SDS-PAGE for optimal separation of full-length (113-kDa) and cleavage fragments (89-kDa, 24-kDa)
- Antibody Detection: Specific antibodies targeting N-terminal (24-kDa fragment) and C-terminal (89-kDa fragment) epitopes
- Validation: Include caspase inhibitor controls (zVAD-fmk) to confirm caspase-specific cleavage
Activity-Based Assays
PARP-1 enzymatic activity assays complement cleavage detection:
- NAD+ Consumption Measurements: Monitor PARP-1 activation preceding cleavage
- PAR Immunodetection: Assess poly(ADP-ribose) formation as indicator of PARP-1 activation
- Functional Assays: Combined approaches to correlate cleavage with functional inactivation
Diagram 1: PARP-1 Cleavage Signaling Pathway (Title: PARP-1 Cleavage Pathway)
Research Reagent Solutions
Table 3: Essential Research Reagents for PARP-1 Cleavage Studies
| Reagent Category | Specific Examples | Research Application | Technical Considerations |
|---|---|---|---|
| Caspase Inhibitors | zVAD-fmk, DEVD-CHO | Caspase-specific cleavage inhibition | Use fresh preparations; optimize concentration |
| PARP-1 Antibodies | Anti-PARP-1 (N-terminal), Anti-PARP-1 (C-terminal) | Cleavage fragment detection | Validate specificity for fragments vs full-length |
| Activity Assays | NAD+ consumption kits, PAR detection assays | Functional correlation with cleavage | Combine with cleavage detection for comprehensive analysis |
| Positive Controls | Staurosporine, Etoposide | Apoptosis induction controls | Optimize concentration and timing for specific cell types |
| Detection Reagents | Chemiluminescent substrates, Fluorescent secondaries | Signal amplification | Match detection method to abundance levels |
Interpretation Challenges and Considerations
The interpretation of PARP-1 cleavage data requires careful attention to contextual factors:
Multiple Protease Involvement
The presence of atypical cleavage fragments may indicate non-caspase protease activity (calpains, cathepsins, granzymes, MMPs). Appropriate protease inhibitors and multiple detection methods are essential for accurate pathway identification.
Fragment Stability and Detection
The 24-kDa DNA-binding fragment may be underrepresented in detection assays due to tight nuclear association, potentially leading to underestimation of cleavage extent. Sequential extraction protocols can improve fragment recovery.
Functional Consequences
PARP-1 cleavage fragments exhibit distinct biological activities beyond PARP-1 inactivation. The 24-kDa fragment influences NF-κB transcriptional activity and modulates inflammatory responses, demonstrating context-dependent functions.
PARP-1 cleavage remains a valuable biochemical marker for apoptosis research, though its interpretation requires sophisticated understanding of cell type-specific behaviors, stimulus-dependent activation patterns, and temporal dynamics. Researchers must employ carefully controlled experimental designs, appropriate detection methodologies, and context-aware interpretation frameworks to accurately utilize PARP-1 cleavage as an indicator of cell death pathway activation. The continuing elucidation of PARP-1's diverse functions beyond DNA repair highlights the importance of contextual analysis in experimental systems and therapeutic applications targeting PARP-1 activity.
Beyond the Hallmark: Validating PARP-1 Cleavage Against Alternative Cell Death Markers and Pathways
Within the landscape of apoptosis research, the cleavage of Poly (ADP-ribose) polymerase-1 (PARP-1) stands as a critical biochemical event. As a nuclear enzyme with fundamental roles in DNA repair and genomic maintenance, PARP-1 becomes a primary target for proteolytic cleavage by caspases during the execution phase of apoptosis. This in-depth analysis positions PARP-1 cleavage within the broader context of apoptotic markers, providing a direct comparison with other established indicators such as Annexin V binding and caspase activation. We will dissect the unique signatures, temporal resolution, and technical applications of each marker, offering a structured guide for researchers and drug development professionals in selecting the most appropriate tools for their experimental and therapeutic objectives.
PARP-1 Cleavage: A Hallmark of Apoptotic Execution
Molecular Mechanism and Fragment Analysis
PARP-1 is a 116 kDa nuclear protein that functions as a DNA damage sensor. During apoptosis, it is specifically cleaved by caspase-3 and -7 at a conserved DEVD214↓G215 motif, located within its nuclear localization signal [3] [56]. This proteolysis generates two signature fragments: a 24-kDa DNA-binding fragment and an 89-kDa catalytic fragment [3] [57]. The 24-kDa fragment, containing the zinc finger motifs, remains tightly bound to DNA ends, acting as a trans-dominant inhibitor of DNA repair by blocking further PARP-1 activation [3]. The 89-kDa fragment, which includes the auto-modification and catalytic domains, is liberated from the nucleus and can translocate to the cytoplasm [3] [31].
Table 1: Key Characteristics of PARP-1 Cleavage Fragments
| Fragment Size | Domains Contained | Cellular Localization Post-Cleavage | Functional Consequence |
|---|---|---|---|
| 24 kDa | Two zinc-finger motifs (DNA-Binding Domain) | Remains nuclear [3] | Irreversibly binds DNA strand breaks, inhibiting DNA repair processes [3]. |
| 89 kDa | BRCT domain, WGR domain, Catalytic Domain | Translocates to cytoplasm [3] [31] | Serves as a carrier for poly(ADP-ribose) (PAR) polymers, facilitating parthanatos [31] [10]. |
Detection Methods and Reagents
The gold-standard method for detecting PARP-1 cleavage is Western blotting, which allows for the clear resolution of the full-length protein (116 kDa) from the 89-kDa fragment [57] [56]. The availability of highly specific monoclonal antibodies, such as the PARP-H8 clone, which recognizes the cleaved Asp214 site, enables precise detection and quantification [57] [56]. Flow cytometry using specific antibodies against the cleaved form also allows for the rapid assessment of apoptosis at the single-cell level within a heterogeneous population [48].
Comparative Analysis of Key Apoptosis Markers
Marker Profiles and Detection Windows
Different apoptosis markers report on distinct stages of the cell death process, from early initiation to late execution and secondary necrosis.
Table 2: Comparative Analysis of Apoptosis Markers: PARP-1 Cleavage, Annexin V, and Caspase Activation
| Parameter | PARP-1 Cleavage | Annexin V Staining | Caspase Activation |
|---|---|---|---|
| Primary Target | Nuclear PARP-1 protein [57] | Phosphatidylserine (PS) on cell membrane [48] | Inactive caspase zymogens (e.g., Pro-caspase-3) |
| Molecular Event | Proteolytic cleavage by effector caspases (3/7) [3] | Externalization of PS from inner to outer membrane leaflet | Proteolytic cleavage and activation cascade |
| Stage of Apoptosis | Mid/Late (Execution phase) [3] | Early (before membrane integrity loss) [48] | Early/Intermediate (Initiation & Execution) |
| Key Functional Outcome | Inhibition of DNA repair; Promotion of cell death [3] [4] | "Eat-me" signal for phagocytes | Cleavage of vital cellular substrates (e.g., PARP-1, ICAD) |
| Primary Detection Methods | Western Blot, Flow Cytometry [57] [48] | Flow Cytometry (with PI counterstain) [48] | Western Blot (cleaved forms), Fluorogenic substrates, Activity assays |
| Key Advantage | Highly specific for caspase-mediated apoptosis; Mechanistic link to DNA repair shutdown | Identifies early-stage apoptotic cells; Amenable to high-throughput | Direct measure of the core apoptotic machinery; Can distinguish initiator vs. effector caspases |
| Key Limitation | Does not distinguish between caspase-dependent apoptosis and parthanatos [31] [10] | Can be positive in necrotic cells; Requires careful staining controls | Activity may be transient; Does not confirm commitment to death |
Temporal and Spatial Specificity in Cell Death Pathways
The integration of these markers can delineate the progression and type of cell death. For instance, a cell exhibiting Annexin V positivity with caspase-3 activation and PARP-1 cleavage is unequivocally undergoing classical apoptosis. However, PARP-1 cleavage can also occur in other contexts. Recent studies reveal that the 89-kDa fragment can carry poly(ADP-ribose) (PAR) polymers to the cytoplasm, facilitating the release of Apoptosis-Inducing Factor (AIF) from mitochondria, a hallmark of parthanatos—a caspase-independent programmed necrosis [31] [10]. This demonstrates that while PARP-1 cleavage is initiated by caspases, its consequences can extend beyond the classical apoptotic pathway.
Furthermore, truncated PARP1 (tPARP1) has been shown to translocate to the cytosol and mono-ADP-ribosylate (MARylate) the RNA Polymerase III (Pol III) complex during innate immune responses, linking apoptosis to amplified interferon-beta production [4]. This discovery of non-canonical functions for the cleavage fragment adds a layer of complexity to its role as a simple apoptosis marker.
The Scientist's Toolkit: Essential Reagents for Detection
Table 3: Key Research Reagents for Apoptosis Detection
| Reagent / Assay | Specific Target/Principle | Primary Application | Function & Importance |
|---|---|---|---|
| Anti-Cleaved PARP1 (clone PARP-H8) [57] [56] | Neo-epitope at Asp214 of human PARP1 | Western Blot, Flow Cytometry | Highly specific antibody for detecting the 89 kDa apoptotic fragment; does not recognize full-length PARP1. |
| Annexin V (FITC conjugate) [48] | Phosphatidylserine (PS) | Flow Cytometry | Binds externalized PS on the outer membrane of early apoptotic cells. |
| Propidium Iodide (PI) [48] | Cellular DNA | Flow Cytometry | Membrane-impermeant dye that stains DNA in late apoptotic and necrotic cells with compromised membranes. Used to distinguish early (Annexin V+/PI-) from late (Annexin V+/PI+) apoptosis. |
| Fluorogenic Caspase Substrates (e.g., DEVD-AFC) | Caspase-3/7 cleavage site | Spectrofluorometry, Microplate Assays | Upon cleavage by active caspases, releases a fluorescent group (e.g., AFC), allowing kinetic measurement of caspase activity. |
| Pan-Caspase Inhibitor (e.g., z-VAD-fmk) [58] | Active site of caspases | Cell Culture Treatment | A broad-spectrum, cell-permeable inhibitor used to confirm the caspase-dependence of an apoptotic stimulus. |
| PARP Inhibitor (e.g., PJ-34, Veliparib) [58] [59] | Catalytic activity of PARP | Cell Culture Treatment | Chemical inhibitors used to study the role of PARP activity in cell death pathways, including its dual roles in survival and death. |
Experimental Protocols for Core Apoptosis Assays
Protocol 1: Detection of PARP-1 Cleavage by Western Blotting
This protocol is foundational for confirming caspase-dependent apoptosis in cell culture models.
- Cell Lysis: Harvest treated and control cells by centrifugation. Lyse cells in RIPA buffer (or similar SDS-containing buffer) supplemented with protease inhibitors. Rotate for 15 minutes at 4°C [58].
- Protein Quantification: Clarify lysates by centrifugation and determine protein concentration using a detergent-compatible assay (e.g., BCA assay) [58].
- Electrophoresis and Transfer: Dilute equal amounts of protein (e.g., 20-30 µg) with SDS-PAGE sample buffer, boil for 5 minutes, and resolve on a 7-10% polyacrylamide gel. Transfer proteins to a PVDF membrane [58].
- Immunoblotting: Block the membrane with 5% non-fat milk. Incubate with a primary antibody specific for cleaved PARP-1 (e.g., clone PARP-H8 at 0.001-1 µg/mL) overnight at 4°C [57]. After washing, incubate with an appropriate HRP-conjugated secondary antibody.
- Visualization: Develop the blot using chemiluminescence reagents. A positive apoptotic signal is indicated by a band at ~89 kDa. Probing for full-length PARP-1 (116 kDa) and a loading control (e.g., GAPDH, β-actin) is essential for result interpretation [57] [56].
Protocol 2: Multiparametric Flow Cytometric Analysis of Apoptosis
This protocol allows for the simultaneous assessment of multiple apoptotic parameters in a single sample.
- Cell Staining for Surface Markers: After treatment, harvest cells and wash with cold PBS. Resuspend ~1x10^6 cells in Annexin V Binding Buffer. Add Annexin V-FITC and anti-CD45-PE (for leukocyte identification) and incubate for 15-30 minutes in the dark [48].
- Fixation and Permeabilization: Wash cells and fix/permeabilize using a commercial kit (e.g., Cytofix/Cytoperm) for 20 minutes to stabilize the cells and allow antibody entry [48].
- Intracellular Staining: After permeabilization, incubate cells with an antibody against cleaved PARP-1 (or active caspase-3) for 45 minutes at 4°C, followed by a fluorochrome-conjugated secondary antibody if needed [48].
- Data Acquisition and Analysis: Analyze the cells on a flow cytometer. Use single-color controls for compensation. Key populations can be gated as follows:
- Viable: Annexin V-, PI-
- Early Apoptotic: Annexin V+, PI-, Cleaved PARP1/Caspase-3+
- Late Apoptotic/Necrotic: Annexin V+, PI+ [48]
Apoptosis Signaling Pathways and Experimental Workflow
The following diagrams, generated using Graphviz DOT language, illustrate the core apoptosis signaling pathway and a standardized experimental workflow for its detection.
Apoptosis Signaling Pathway and Marker Detection
Integrated Experimental Workflow for Apoptosis Analysis
PARP-1 cleavage remains a cornerstone marker in apoptosis research, offering high specificity for caspase activation and a direct mechanistic link to the irreversible shutdown of DNA repair. Its value is maximized when used not in isolation, but as part of a multiparametric analytical approach that includes Annexin V for early-stage detection and caspase activity assays for confirming the core apoptotic machinery. The evolving understanding of PARP-1's role in alternative cell death pathways like parthanatos, and the non-canonical functions of its cleavage fragments, underscores its biological complexity beyond a simple marker. For the drug development professional, this integrated perspective is crucial for accurately interpreting the mechanism of action of novel therapeutics and for making informed decisions on the most specific and informative biomarker panels for preclinical and clinical studies.
Cross-Validation with Mitochondrial Markers of Apoptosis (Cytochrome c Release, AIF Translocation)
Poly(ADP-ribose) polymerase-1 (PARP-1) is a nuclear enzyme that functions as a primary DNA damage sensor, playing a critical role in the cellular response to genotoxic stress. Its cleavage into specific fragments is widely recognized as a biomarker of apoptosis, serving as a key indicator of commitment to programmed cell death. This whitepaper details the essential methodologies for cross-validating PARP-1 cleavage with the central mitochondrial events of apoptosis—cytochrome c release and apoptosis-inducing factor (AIF) translocation. For researchers and drug development professionals, establishing this correlation is crucial for definitively characterizing cell death pathways and evaluating the efficacy of novel therapeutic compounds. The following sections provide a technical guide for experimental design, featuring standardized protocols, data interpretation guidelines, and visualization of the interconnected signaling networks.
PARP-1 Cleavage: A Hallmark of Apoptosis
Molecular Mechanism and Fragment Generation
PARP-1 is a 116-kDa nuclear protein comprising three primary domains: an N-terminal DNA-binding domain (DBD), a central auto-modification domain (AMD), and a C-terminal catalytic domain (CD) [3]. During caspase-dependent apoptosis, executioner caspases-3 and -7 cleave PARP-1 at a conserved aspartic acid residue (DEVD214) located within a nuclear localization signal (NLS) near the DBD [3] [28]. This proteolytic event generates two characteristic fragments:
- A 24-kDa fragment containing the DBD and NLS. This fragment remains tightly bound to DNA strand breaks, acting as a trans-dominant inhibitor of intact PARP-1 and other DNA repair enzymes, thereby facilitating the apoptotic process [3] [6].
- An 89-kDa fragment encompassing the AMD and CD. Recent research has revealed that this fragment, particularly when modified with poly(ADP-ribose) (PAR) polymers, can translocate to the cytoplasm and function as a PAR carrier, contributing to AIF-mediated death signaling [10] [31] [6].
Table 1: PARP-1 Cleavage Fragments and Their Properties
| Fragment Size | Domains Contained | Subcellular Localization Post-Cleavage | Primary Function |
|---|---|---|---|
| 24 kDa | DNA-Binding Domain (DBD), Nuclear Localization Signal (NLS) | Nuclear, bound to DNA lesions | Trans-dominant inhibitor of DNA repair [3] |
| 89 kDa | Auto-modification Domain (AMD), Catalytic Domain (CD) | Cytoplasmic (can translocate) | PAR carrier; promotes AIF release [10] [6] |
Significance in Cell Death Pathways
PARP-1 cleavage serves as a critical switch, halting DNA repair and conserving cellular energy to allow for the orderly execution of apoptosis [3]. The appearance of the 24-kDa and 89-kDa fragments is a well-established signature of caspase activation and is routinely used to differentiate apoptosis from other forms of cell death, such as necrosis or parthanatos (a PARP-1-dependent, caspase-independent cell death) [3] [6]. Furthermore, the function of the cleavage fragments extends beyond a simple loss-of-function, as they actively participate in and amplify cell death pathways [28].
Mitochondrial Apoptosis Signaling Pathways
The intrinsic (mitochondrial) pathway of apoptosis is a major coordinator of cell death, characterized by mitochondrial outer membrane permeabilization (MOMP) and the release of pro-apoptotic proteins into the cytosol. Two key players released are cytochrome c and AIF.
Cytochrome c and the Caspase Activation Pathway
Cytochrome c is a heme protein essential for the electron transport chain. Upon MOMP, its release into the cytosol triggers the formation of the apoptosome, a multi-protein complex that activates caspase-9. Caspase-9, in turn, cleaves and activates executioner caspases-3 and -7, which are directly responsible for PARP-1 cleavage [60].
AIF and Caspase-Independent Death
Apoptosis-inducing factor (AIF) is a flavoprotein anchored to the mitochondrial inner membrane. Upon induction of cell death, it is released and translocates to the nucleus, where it promotes caspase-independent large-scale DNA fragmentation and chromatin condensation [61] [60]. AIF release can be triggered by several mechanisms, including pro-apoptotic Bcl-2 family proteins and calpain proteases [6]. Critically, it is also a key effector in parthanatos, where it is released in response to PAR polymers generated by hyperactivated PARP-1 [61] [6].
The following diagram illustrates the convergence of these pathways on PARP-1 cleavage and the role of the 89-kDa fragment.
Diagram 1: Integrated Signaling in Cell Death. This diagram illustrates how PARP-1 hyperactivation and mitochondrial caspase activation converge. A key cross-talk mechanism involves the caspase-generated 89-kDa PARP-1 fragment promoting AIF release [10] [6].
Experimental Protocols for Cross-Validation
A robust cross-validation requires detecting PARP-1 cleavage and confirming it with assays for cytochrome c release and AIF translocation.
Detecting PARP-1 Cleavage by Western Blot
Objective: To identify the characteristic 89-kDa and 24-kDa PARP-1 cleavage fragments. Procedure:
- Cell Lysis: Lyse control and treated cells (e.g., with 1-10 µM Staurosporine for 3-6 hours) using RIPA buffer supplemented with protease inhibitors.
- Protein Quantification: Determine protein concentration using a BCA or Bradford assay.
- Gel Electrophoresis: Load 20-50 µg of total protein per lane on a 4-12% Bis-Tris polyacrylamide gel and run at constant voltage.
- Membrane Transfer: Transfer proteins from the gel to a PVDF or nitrocellulose membrane.
- Blocking and Antibody Incubation:
- Block the membrane with 5% non-fat milk in TBST for 1 hour.
- Incubate with primary antibody (e.g., anti-PARP-1, which detects full-length and the 89-kDa fragment) diluted in blocking buffer overnight at 4°C.
- Wash the membrane and incubate with an HRP-conjugated secondary antibody for 1 hour at room temperature.
- Detection: Develop the blot using enhanced chemiluminescence (ECL) substrate. A successful apoptosis induction is indicated by the appearance of the 89-kDa band and a corresponding decrease in the full-length 116-kDa PARP-1 band [3] [6].
Assessing Cytochrome c Release by Immunofluorescence
Objective: To visualize the translocation of cytochrome c from mitochondria to the cytosol. Procedure:
- Cell Seeding and Treatment: Culture cells on glass coverslips and treat with an apoptotic inducer.
- Fixation and Permeabilization: Fix cells with 4% paraformaldehyde for 15 minutes, then permeabilize with 0.1% Triton X-100 in PBS for 10 minutes.
- Antibody Staining:
- Block with 1% BSA in PBS for 30 minutes.
- Incubate with a mouse anti-cytochrome c primary antibody for 1-2 hours.
- Incubate with an Alexa Fluor-conjugated secondary antibody (e.g., Alexa Fluor 488) for 1 hour.
- Co-stain mitochondria using MitoTracker Deep Red or an antibody against a mitochondrial protein (e.g., COX IV).
- Counterstain nuclei with DAPI.
- Imaging and Analysis: Acquire images using a confocal microscope. In healthy cells, cytochrome c staining shows a punctate pattern that co-localizes with mitochondrial markers. Upon apoptosis induction, this pattern becomes a diffuse cytosolic signal, indicating release [60].
Monitoring AIF Translocation by Subcellular Fractionation
Objective: To biochemically confirm the movement of AIF from mitochondria to the nucleus. Procedure:
- Fraction Preparation: Use a commercial subcellular fractionation kit to isolate cytoplasmic, mitochondrial, and nuclear fractions from control and treated cells.
- Western Blot Analysis:
- Run equal protein amounts from each fraction on SDS-PAGE gels.
- Transfer to membranes and probe with specific antibodies:
- Anti-AIF: Look for a decrease in the mitochondrial fraction and an increase in the nuclear fraction.
- Fraction Purity Controls:
- Cytochrome c oxidase subunit IV (COX IV) or Voltage-Dependent Anion Channel (VDAC) for mitochondria.
- Lamin A/C or Histone H3 for the nucleus.
- GAPDH or α-tubulin for the cytoplasm.
- Data Interpretation: Apoptosis is confirmed by a clear nuclear accumulation of AIF, which corresponds with PARP-1 cleavage observed in the whole-cell lysates [61] [6].
Table 2: Key Assays for Cross-Validation of Apoptosis
| Assay | Target Readout | Key Observation for Apoptosis | Technical Notes |
|---|---|---|---|
| PARP-1 Western Blot | 89-kDa and 24-kDa fragments | Appearance of 89-kDa cleavage band [3] | Standard protocol; primary antibody must recognize C-terminal epitope. |
| Cytochrome c Immunofluorescence | Subcellular localization of cytochrome c | Punctate (mitochondrial) → Diffuse (cytosolic) staining [60] | Requires high-resolution microscopy; co-staining for mitochondria is essential. |
| AIF Subcellular Fractionation | AIF protein levels in mitochondrial vs. nuclear fractions | Decrease in mitochondria; Increase in nucleus [61] [6] | Requires rigorous fraction purity controls. |
The Scientist's Toolkit: Essential Research Reagents
Successful execution of these experiments relies on high-quality, specific reagents. The table below lists essential tools for investigating these pathways.
Table 3: Key Research Reagents for Apoptosis Cross-Validation
| Reagent / Assay | Specific Example / Catalog Number | Primary Function in Experimental Context |
|---|---|---|
| Apoptosis Inducers | Staurosporine (1-10 µM), Actinomycin D (0.5-5 µM) [10] [6] | Positive control to trigger intrinsic apoptosis pathway. |
| Caspase Inhibitor | zVAD-fmk (pan-caspase inhibitor, 20-50 µM) [6] | Confirms caspase-dependence of cell death and PARP-1 cleavage. |
| PARP-1 Inhibitor | PJ34, ABT-888 (Veliparib) (1-10 µM) [6] | Differentiates PARP-1-dependent (parthanatos) from independent death. |
| Anti-PARP-1 Antibody | Western Blot, recognizes C-terminus (e.g., Cell Signaling #9542) | Detects full-length (116-kDa) and cleaved 89-kDa fragment. |
| Anti-Cytochrome c Antibody | Immunofluorescence (e.g., BD Biosciences #556432) | Visualizes release from mitochondria during apoptosis. |
| Anti-AIF Antibody | Western Blot/IF (e.g., Abcam ab1998) | Tracks AIF localization in fractionation or imaging studies. |
| Mitochondrial Stain | MitoTracker Deep Red (Thermo Fisher M22426) | Labels intact mitochondria for co-localization studies. |
| Subcellular Fractionation Kit | Mitochondria Isolation Kit (e.g., Thermo Scientific 89874) | Isolates pure mitochondrial fractions for AIF/cytochrome c blots. |
Data Interpretation and Integration
Correlating data from the described assays provides a comprehensive picture of the cell death mechanism.
- Confirmation of Apoptosis: A clear apoptotic signature is defined by the simultaneous presence of PARP-1 cleavage fragments, cytosolic release of cytochrome c, and nuclear translocation of AIF. This pattern is typically observed with classical inducers like staurosporine [6].
- Differentiating Cell Death Pathways: Inhibition experiments are critical for pathway delineation.
- If cell death and AIF translocation are blocked by zVAD-fmk (a caspase inhibitor), it confirms a caspase-dependent apoptotic pathway.
- If cell death and AIF translocation are blocked by a PARP inhibitor (e.g., PJ34) but not by zVAD-fmk, it indicates parthanatos [6].
- The discovery that the 89-kDa PARP-1 fragment can carry PAR to the cytoplasm to induce AIF release reveals a direct molecular cross-talk between the caspase and parthanatos pathways [10] [6].
The following workflow provides a logical framework for analyzing experimental outcomes.
Diagram 2: Decision Workflow for Cell Death Pathway Identification. This logic tree guides the interpretation of experimental data to classify the dominant cell death pathway based on the cross-validation results.
Differentiating Apoptotic Cleavage from Parthanatos and Other PARP-1-Mediated Death Pathways
Poly(ADP-ribose) polymerase-1 (PARP-1) is a nuclear enzyme that functions as a critical DNA damage sensor and participates in multiple cellular processes, including DNA repair, transcription, and cell death regulation. As a central player in determining cell fate, PARP-1 activation can lead to dramatically different outcomes depending on the intensity and nature of the cellular insult. Within the context of a broader thesis on PARP-1 cleavage and its significance as an apoptosis marker, this technical guide provides a comprehensive framework for distinguishing between the primary cell death pathways mediated by PARP-1: apoptotic cleavage and parthanatos. Understanding these distinct mechanisms is essential for researchers and drug development professionals investigating cancer therapeutics, neurodegenerative disorders, and ischemic injuries, where PARP-1-mediated cell death plays a pivotal pathophysiological role.
PARP-1 Structure and Functional Domains
PARP-1 is a modular protein comprising several functionally specialized domains that dictate its activity and cleavage patterns during cell death processes. With 1,014 amino acids, the protein's organization includes an N-terminal DNA-binding domain (DBD) containing three zinc finger motifs (Zn1, Zn2, Zn3) that recognize DNA strand breaks [3] [9]. The Zn1 and Zn2 motifs specifically bind to DNA damage sites, while Zn3 facilitates inter-domain interactions essential for PARP-1 activation [3]. The central auto-modification domain (AMD) contains a BRCT (BRCA1 C-terminal) fold that mediates protein-protein interactions and serves as the primary site for poly(ADP-ribosyl)ation [3] [9]. The C-terminal catalytic domain (CAT) is responsible for poly(ADP-ribose) (PAR) synthesis using NAD+ as a substrate [9]. This domain includes a tryptophan-glycine-arginine (WGR) region that interacts with other domains to transmit DNA damage signals to the catalytic center [9]. A critical aspartate-glutamate-valine-aspartic acid (DEVD) sequence located within the DBD serves as the cleavage site for caspase-3 and caspase-7 during apoptosis [28] [9].
PARP-1 in Apoptotic Cleavage
Mechanism and Proteolytic Fragments
Apoptotic cleavage of PARP-1 represents a biochemical hallmark of caspase-dependent programmed cell death. This process is primarily mediated by executioner caspases-3 and -7, which recognize and cleave the conserved DEVD214 site within PARP-1's DNA-binding domain [3] [28]. This proteolytic event generates two characteristic fragments: a 24-kDa fragment containing the DBD with two zinc finger motifs, and an 89-kDa fragment comprising the AMD and CAT domains [3] [31]. The 24-kDa fragment remains tightly bound to DNA strand breaks, acting as a trans-dominant inhibitor of PARP-1 activity and preventing DNA repair processes [3]. The 89-kDa fragment, which loses its nuclear localization signal due to the cleavage, exhibits significantly reduced DNA binding capacity and may be liberated from the nucleus to the cytosol [3] [31].
Recent research has revealed that the 89-kDa fragment can serve as a carrier for poly(ADP-ribose) (PAR) polymers, facilitating their translocation to the cytoplasm where they participate in apoptosis-inducing factor (AIF)-mediated cell death, representing a potential point of crosstalk between apoptotic and parthanatos pathways [31]. This cleavage event serves as an irreversible commitment step in apoptosis, ensuring the dismantling of cellular infrastructure while conserving ATP that would otherwise be expended on DNA repair efforts [3].
Biological Consequences
The biological implications of PARP-1 cleavage during apoptosis extend beyond simply inactivating the enzyme. The 24-kDa DNA-binding fragment creates a physical barrier at DNA damage sites, blocking access for DNA repair machinery and ensuring the apoptotic process proceeds unimpeded [3]. This fragment's irreversible binding to DNA breaks effectively suppresses DNA repair while simultaneously conserving cellular ATP pools that would be depleted by hyperactive PARP-1 [3]. Meanwhile, the 89-kDa catalytic fragment undergoes translocation from the nucleus to the cytoplasm, where recent evidence suggests it may function as a PAR carrier that facilitates AIF release from mitochondria [31]. This newly discovered role indicates that PARP-1 cleavage products may actively promote cell death execution rather than merely disabling DNA repair mechanisms.
Table 1: Key Characteristics of PARP-1 Cleavage Fragments in Apoptosis
| Fragment | Molecular Weight | Domains Contained | Cellular Localization | Primary Functions |
|---|---|---|---|---|
| 24-kDa | 24 kDa | DNA-binding domain (DBD) with zinc fingers 1 and 2 | Nuclear, bound to DNA | Trans-dominant inhibitor of PARP-1; blocks DNA repair |
| 89-kDa | 89 kDa | Auto-modification domain (AMD) and catalytic domain (CAT) | Cytosolic translocation | Potential PAR carrier; may facilitate AIF-mediated death |
PARP-1 in Parthanatos
Molecular Mechanisms
Parthanatos represents a distinct form of programmed cell death characterized by PARP-1 hyperactivation in response to severe DNA damage. Unlike apoptosis, parthanatos is caspase-independent and occurs through an orchestrated molecular cascade initiated by excessive PARP-1 activity [62] [63]. The process begins with significant DNA damage triggered by various stimuli including reactive oxygen species (ROS), alkylating agents, excitotoxicity, or ischemia-reperfusion injury [62]. This damage leads to PARP-1 hyperactivation, catalyzing the massive synthesis of PAR polymers from NAD+ substrates [62] [63]. The consequent PAR accumulation acts as a signaling molecule that induces mitochondrial depolarization and triggers the release of apoptosis-inducing factor (AIF) from mitochondria [62]. AIF then translocates to the nucleus, where it collaborates with other factors such as macrophage migration inhibitory factor (MIF) to initiate large-scale DNA fragmentation (50+ kbp) and chromatin condensation, ultimately resulting in cell demise [62] [63].
A critical aspect of parthanatos is the profound energy depletion that occurs as a consequence of PARP-1 hyperactivation. The enzyme's excessive consumption of NAD+ depletes cellular pools of this essential cofactor, disrupting glycolysis and mitochondrial respiration [63]. The resynthesis of NAD+ further consumes ATP, creating a vicious cycle of energy failure that contributes to the irreversible commitment to cell death [62]. This energetic collapse represents a key distinguishing feature between parthanatos and apoptosis, where ATP levels are typically preserved.
Key Mediators and Regulatory Factors
Several critical mediators orchestrate the parthanatos cascade. PAR polymers themselves serve as both signaling molecules and cytotoxic agents when produced in excess [62]. These polymers bind directly to AIF, facilitating its release from mitochondria and subsequent nuclear translocation [62]. The apoptosis-inducing factor (AIF) is a flavoprotein located in the mitochondrial intermembrane space that, despite its name, functions independently of caspases in parthanatos [62]. Upon release, AIF migrates to the nucleus where it initiates chromatinolysis and large-scale DNA fragmentation. Poly(ADP-ribose) glycohydrolase (PARG), the primary enzyme responsible for PAR degradation, plays a complex regulatory role in parthanatos [63]. While PARG activity typically counteracts PARP-1 signaling, its function becomes essential for parthanatos execution in some contexts, possibly through the generation of free PAR polymers that can exit the nucleus and interact with cytoplasmic targets like AIF [63].
Table 2: Comparative Features of Apoptosis and Parthanatos
| Feature | Apoptosis | Parthanatos |
|---|---|---|
| Initiating Stimuli | Death receptor signaling, mild DNA damage, trophic factor withdrawal | Severe DNA damage, excitotoxicity, ischemia, ROS/RNS |
| Key Proteases | Caspases-3 and -7 | Caspase-independent |
| PARP-1 Cleavage | Yes (generates 24-kDa and 89-kDa fragments) | No (full-length hyperactivation) |
| Energy Status | ATP preserved | NAD+/ATP depletion |
| DNA Fragmentation | Oligonucleosomal (180-200 bp) | Large-scale (15-50 kbp) |
| Key Effectors | Caspase cascade, cytochrome c | PAR polymers, AIF, PARG |
| Morphological Features | Nuclear shrinkage, apoptotic bodies | Nuclear condensation, loss of membrane integrity |
Other PARP-1-Mediated Death Pathways
Beyond classical apoptosis and parthanatos, PARP-1 participates in additional cell death modalities through interactions with various proteases and signaling pathways. Calpains, calcium-activated cysteine proteases, can cleave PARP-1 at distinct sites to generate fragments different from those produced by caspases, potentially contributing to excitotoxic neuronal death [3]. Cathepsins, lysosomal proteases released during cellular stress, can also process PARP-1, while granzyme B from cytotoxic T cells cleaves PARP-1 during immune-mediated cell killing [3]. Matrix metalloproteinases (MMPs) have been reported to generate specific PARP-1 fragments, although their functional significance remains less characterized [3].
Emerging research has revealed fascinating crosstalk between PARP-1 and other regulated cell death pathways. In ferroptosis, an iron-dependent form of cell death characterized by lipid peroxidation, the ferroptosis inducer RSL3 has been shown to trigger PARP-1 activation through dual mechanisms: caspase-dependent cleavage and epitranscriptomic regulation of PARP-1 expression via m6A RNA modification [20]. This intersection demonstrates how PARP-1 can function as a molecular node integrating diverse cell death signals. Additionally, recent findings indicate that PAR polymers produced by activated PARP-1 can directly bind to STING (stimulator of interferon genes), promoting apoptosis upon ionizing radiation-induced DNA damage [21]. This mechanism connects DNA damage response to innate immune signaling and expands the functional repertoire of PAR beyond traditional cell death pathways.
Experimental Approaches for Differentiation
Methodologies for Pathway Identification
Distinguishing between PARP-1-mediated death pathways requires a multifaceted experimental approach combining biochemical, morphological, and genetic techniques. Western blot analysis of PARP-1 cleavage fragments provides essential information, with the characteristic 89-kDa and 24-kDa fragments indicating apoptotic cleavage, while full-length PARP-1 depletion or persistence suggests alternative pathways [20] [28]. Assessment of PAR polymer levels through immunoblotting or immunofluorescence is crucial for identifying parthanatos, which features massive PAR accumulation [62] [63]. Caspase activity assays using fluorogenic substrates or antibodies against activated caspases-3/7 can confirm apoptotic involvement, while their absence suggests caspase-independent mechanisms like parthanatos [3] [62].
Cell viability assays under different inhibitor conditions provide functional discrimination; PARP inhibitors (e.g., PJ34, olaparib) protect against parthanatos but not necessarily apoptosis, while pan-caspase inhibitors (e.g., Z-VAD-FMK) block apoptosis but not parthanatos [20] [21]. Genetic approaches including PARP-1 knockout or knockdown cells demonstrate resistance to parthanatos inducers, while having variable effects on apoptosis depending on the stimulus [62] [28]. AIF localization studies using subcellular fractionation or immunofluorescence can detect its nuclear translocation, a hallmark of parthanatos [62] [31]. Additionally, metabolic assays measuring NAD+ and ATP levels reveal the energetic collapse characteristic of parthanatos but not apoptosis [62] [63].
Research Reagent Solutions
Table 3: Essential Research Reagents for Studying PARP-1-Mediated Cell Death
| Reagent | Category | Primary Function | Application Examples |
|---|---|---|---|
| PJ34 | PARP inhibitor | Potent inhibition of PARP-1 catalytic activity | Suppression of parthanatos; concentration-dependent reduction of IR-induced apoptosis [21] |
| Olaparib | PARP inhibitor | Clinical PARP inhibitor used in cancer therapy | Studying PARP-1 functions in DNA repair and cell death; investigating PARPi resistance [20] |
| Z-VAD-FMK | Pan-caspase inhibitor | Irreversible inhibition of caspase activity | Differentiation between caspase-dependent and independent death; apoptosis confirmation [20] |
| Ferrostatin-1 (Fer-1) | Ferroptosis inhibitor | Specific inhibition of lipid peroxidation in ferroptosis | Investigating PARP-1 role in ferroptosis-apoptosis crosstalk [20] |
| Anti-PARP-1 antibodies | Immunological reagent | Detection of full-length and cleaved PARP-1 | Western blot analysis to identify 89-kDa/24-kDa apoptotic fragments [20] [28] |
| Anti-PAR antibodies | Immunological reagent | Recognition of poly(ADP-ribose) polymers | Detection of PAR accumulation in parthanatos [62] [63] |
| Anti-AIF antibodies | Immunological reagent | Visualization of AIF subcellular localization | Confirmation of mitochondrial release and nuclear translocation in parthanatos [62] [31] |
| Anti-cleaved caspase-3 antibodies | Immunological reagent | Specific detection of activated caspase-3 | Apoptosis verification; differentiation from caspase-independent pathways [21] |
Signaling Pathway Diagrams
PARP-1 serves as a critical molecular switch governing cell fate decisions in response to DNA damage, with its specific activation and cleavage patterns determining whether a cell undergoes apoptosis, parthanatos, or alternative death pathways. The differentiation between these mechanisms relies on recognizing key biomarkers: caspase-mediated generation of 89-kDa and 24-kDa PARP-1 fragments in apoptosis versus PARP-1 hyperactivation, PAR accumulation, and AIF nuclear translocation in parthanatos. For researchers and drug development professionals, understanding these distinctions has significant therapeutic implications. PARP inhibitors have shown promise in conditions where parthanatos contributes to pathology, such as stroke, neurodegenerative diseases, and ischemia-reperfusion injury, while the detection of PARP-1 cleavage fragments serves as a valuable biomarker for apoptosis in cancer therapeutics and toxicology studies. As research continues to unveil the complex interactions between PARP-1 and various cell death pathways, new opportunities emerge for targeted interventions in diverse pathological conditions characterized by dysregulated cell death.
This whitepaper provides a comparative analysis of three distinct regulated cell death (RCD) modalities—necroptosis, ferroptosis, and autophagy—framed within the context of Poly(ADP-ribose) polymerase-1 (PARP-1) cleavage as a hallmark of apoptosis. PARP-1 cleavage serves as a critical molecular switch that not only signifies apoptotic engagement but also influences crosstalk between different cell death pathways. We examine the unique morphological features, key molecular regulators, and functional consequences of each death modality, with particular emphasis on how PARP-1 processing creates signature fragments that drive specific downstream events. The findings presented herein offer insights for researchers and drug development professionals seeking to target specific cell death pathways for therapeutic intervention, particularly in cancer and neurodegenerative diseases.
Poly(ADP-ribose) polymerase-1 (PARP-1) is a nuclear enzyme with well-established roles in DNA damage repair and maintenance of genome integrity. PARP-1 cleavage has long been recognized as a biochemical hallmark of apoptosis, mediated primarily by caspase-3 and -7 executioner caspases that target a specific aspartate residue (D214 in human PARP-1) [27] [4]. This proteolytic event generates two characteristic fragments: a 24-kDa DNA-binding domain (DBD) fragment and an 89-kDa fragment containing the auto-modification domain (AMD) and catalytic domain (CD) [27]. The 24-kDa fragment remains nuclear-bound and acts as a trans-dominant inhibitor of DNA repair, while the 89-kDa fragment translocates to the cytoplasm where it can engage in non-apoptotic signaling pathways [10].
While initially characterized as an apoptosis-specific marker, recent evidence reveals that PARP-1 is subject to proteolytic processing by different proteases across multiple cell death pathways, producing distinct signature fragments that can serve as diagnostic biomarkers for specific death modalities [27]. This whitepaper explores how PARP-1 cleavage patterns and functions differ across necroptosis, ferroptosis, and autophagy, and how these pathways interact through shared molecular components.
PARP-1 Cleavage Across Cell Death Modalities: Comparative Analysis
Proteolytic Processing of PARP-1 in Different Cell Death Contexts
Table 1: PARP-1 Cleavage Signatures in Different Cell Death Modalities
| Cell Death Modality | Cleaving Proteases | PARP-1 Fragment Sizes | Functional Consequences | Regulatory Inhibitors |
|---|---|---|---|---|
| Apoptosis | Caspase-3, -7 [27] | 24 kDa + 89 kDa [27] | 24-kDa fragment inhibits DNA repair; 89-kDa fragment translocates to cytoplasm [27] [10] | z-VAD-fmk (pan-caspase inhibitor) [64] |
| Necrosis | Cathepsins B, D, G (lysosomal proteases) [64] | 50 kDa (major fragment) [64] | Not fully characterized; correlates with loss of membrane integrity | E64d (cathepsin inhibitor) [64] |
| Ferroptosis-Apoptosis Crosstalk | Caspase-3 (via ROS activation) [14] | 24 kDa + 89 kDa (apoptotic signature) [14] | ROS-dependent cleavage; contributes to apoptotic execution during ferroptosis | Ferrostatin-1, Liproxstatin-1 (ferroptosis inhibitors) [14] |
Morphological and Biochemical Hallmarks of Different Cell Death Modalities
Table 2: Comparative Characteristics of Cell Death Modalities
| Feature | Apoptosis | Necroptosis | Ferroptosis | Autophagy |
|---|---|---|---|---|
| Primary Morphology | Cell shrinkage, chromatin condensation, apoptotic bodies [65] | Cell swelling, plasma membrane rupture, organelle dilation [65] | Smaller mitochondria, reduced cristae, outer membrane rupture [66] | Double-membraned autophagosomes, vacuolization [65] |
| Key Biochemical Events | Caspase activation, PARP-1 cleavage, phosphatidylserine externalization [27] [65] | RIPK1/RIPK3 activation, MLKL phosphorylation [65] | Iron accumulation, lipid peroxidation, GSH depletion [67] [66] | LC3-I to LC3-II conversion, autolysosome formation [65] |
| PARP-1 Processing | Caspase-mediated cleavage (24 kDa + 89 kDa) [27] | Potential lysosomal protease involvement [64] | Indirect via ROS-induced caspase activation [14] | Not typically cleaved; potential PARylation-mediated regulation [68] |
| Immune Response | Anti-inflammatory (minimal release of cellular contents) [65] | Proinflammatory (DAMP release) [65] | Proinflammatory (lipid peroxidation products) [69] | Context-dependent (anti- or pro-inflammatory) [69] |
Molecular Mechanisms and Signaling Pathways
PARP-1 Domains and Cleavage Sites
PARP-1 contains several functionally distinct domains: a 46-kDa DNA-binding domain (DBD) with zinc finger motifs at the N-terminus, a 22-kDa auto-modification domain (AMD) in the central region, and a 54-kDa catalytic domain (CD) at the C-terminus [27]. During apoptosis, caspase-3 cleaves PARP-1 at D214 within the nuclear localization signal, separating the DBD from the AMD and CD [27] [4]. This specific cleavage pattern inactivates PARP-1's DNA repair function while generating fragments with novel signaling capabilities.
Diagram 1: PARP-1 cleavage in apoptosis
Necroptosis Signaling and Potential PARP-1 Involvement
Necroptosis represents a programmed form of necrosis that occurs when caspase activity is inhibited. Unlike apoptosis, necroptosis involves lysosomal proteases such as cathepsins B and G, which generate a characteristic 50-kDa PARP-1 fragment rather than the 24-kDa/89-kDa apoptotic signature [64]. This pathway typically initiates through death receptor activation (e.g., TNFR1) when caspase-8 is inhibited, leading to RIPK1/RIPK3 complex formation and MLKL phosphorylation, ultimately causing plasma membrane rupture [65].
Ferroptosis Mechanisms and Apoptosis Crosstalk
Ferroptosis is an iron-dependent form of regulated cell death driven by catastrophic lipid peroxidation [67]. Key molecular features include glutathione depletion, GPX4 inactivation, and accumulation of lipid reactive oxygen species (ROS) [66] [14]. Interestingly, ferroptosis inducers like erastin and RSL3 can trigger apoptosis through parallel pathways, as demonstrated by the synergistic interaction between ferroptotic agents and TRAIL (TNF-related apoptosis-inducing ligand) [67]. RSL3 promotes PARP-1 apoptotic functions through distinct mechanisms: (1) caspase-dependent PARP-1 cleavage via ROS activation, and (2) reduced full-length PARP-1 expression through inhibition of METTL3-mediated m6A modification [14].
Diagram 2: Ferroptosis-apoptosis crosstalk
Autophagy as a Cell Death Mechanism and Its Regulation
Autophagy, specifically autophagic cell death (type II cell death), involves the degradation of cellular components through lysosomal machinery [65]. This process can be further classified into autophagy-dependent cell death (ADCD) and autophagy-mediated cell death (AMCD), with the latter involving interaction between autophagic machinery and other cell death molecules [65]. The core autophagy machinery includes the ULK1 complex initiation, ATG conjugation systems, and LC3 processing, all negatively regulated by mTOR under nutrient-rich conditions [65] [69]. PARP-1 influences autophagy through PARylation signaling and NAD+ metabolism, particularly during starvation-induced autophagy [68].
Experimental Approaches and Methodologies
Detection and Characterization of PARP-1 Cleavage
Western Blot Analysis for PARP-1 Cleavage:
- Cell Lysis: Harvest cells and lyse in RIPA buffer supplemented with protease inhibitors (e.g., PMSF, complete protease inhibitor cocktail) and caspase inhibitor (z-VAD-fmk) when appropriate [14].
- Electrophoresis: Separate proteins (20-50 μg per lane) on 4-12% Bis-Tris gradient gels to resolve full-length PARP-1 (113 kDa) and cleavage fragments (89 kDa, 24 kDa) [27] [14].
- Antibody Detection: Use PARP-1 antibodies that recognize both full-length and cleaved fragments. Antibodies specifically targeting the 89-kDa fragment or 24-kDa fragment can provide additional specificity [4] [14].
- Interpretation: The 89-kDa fragment indicates caspase-mediated apoptosis, while a 50-kDa fragment suggests lysosomal protease activity in necrosis [64].
Annexin V/Propidium Iodide Staining for Cell Death Modality Discrimination:
- Procedure: Stain cells with FITC-conjugated Annexin V and propidium iodide (PI) according to manufacturer protocols [4] [14].
- Analysis: Analyze by flow cytometry: Annexin V+/PI- early apoptotic cells; Annexin V+/PI+ late apoptotic cells; Annexin V-/PI+ necrotic/necroptotic cells [4].
Assessment of Ferroptosis Markers
Lipid Peroxidation Measurement:
- MDA Assay: Measure malondialdehyde (MDA), a lipid peroxidation byproduct, using thiobarbituric acid reactive substances (TBARS) assay [67].
- BODIPY-C11 Staining: Use BODIPY 581/591 C11 probe that shifts fluorescence from red to green upon oxidation; quantify by flow cytometry or fluorescence microscopy [14].
GSH and GPX4 Activity Assays:
- GSH Detection: Use commercial GSH/GSSG ratio detection kits based on enzymatic recycling method [67].
- GPX4 Activity: Measure GPX4 activity using NADPH consumption assay coupled with glutathione reductase [14].
Autophagy Flux Monitoring
LC3-I/LC3-II Conversion Assay:
- Western Blot: Detect LC3-I (cytosolic) and LC3-II (lipidated, autophagosome-bound) forms by immunoblotting [65] [69].
- Inhibitor Treatment: Include chloroquine (lysosomal inhibitor) to block autophagosome degradation and assess autophagic flux [69].
Immunofluorescence Microscopy:
- Transfected LC3: Express GFP-LC3 in cells and quantify puncta formation representing autophagosomes [65].
- Endogenous LC3: Stain endogenous LC3 with specific antibodies and count LC3-positive vesicles [69].
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Research Reagents for Cell Death Studies
| Reagent Category | Specific Examples | Primary Function | Application Context |
|---|---|---|---|
| PARP-1 Inhibitors | Olaparib, Rucaparib, Niraparib [68] | Inhibit PARP catalytic activity; induce synthetic lethality in BRCA-deficient cells | Cancer therapy, PARP function studies |
| Caspase Inhibitors | z-VAD-fmk (pan-caspase) [64] | Broad-spectrum caspase inhibition; distinguishes caspase-dependent vs independent death | Apoptosis studies, necroptosis induction |
| Ferroptosis Inhibitors | Ferrostatin-1, Liproxstatin-1 [67] [14] | Lipid antioxidant activity; specifically inhibits ferroptosis | Ferroptosis validation, pathway dissection |
| Ferroptosis Inducers | Erastin, RSL3, ART [67] [14] | Erastin: system Xc- inhibitor; RSL3: GPX4 inhibitor | Ferroptosis induction, combination therapies |
| Autophagy Modulators | Rapamycin (inducer), 3-MA, Bafilomycin A1 (inhibitors) [65] [69] | Regulate different stages of autophagic flux | Autophagy studies, death mechanism discrimination |
| Necroptosis Inhibitors | Necrostatin-1 [65] | Selective RIPK1 inhibitor; blocks necroptosis pathway | Necroptosis validation, pathway analysis |
| Death Receptor Ligands | TRAIL, FasL [65] [67] | Activate extrinsic apoptosis pathway | Apoptosis induction, combination studies |
Discussion: Therapeutic Implications and Future Directions
The intricate crosstalk between cell death modalities, with PARP-1 cleavage as a central node, presents both challenges and opportunities for therapeutic intervention. PARP inhibitors have demonstrated clinical success in BRCA-mutant cancers by exploiting synthetic lethality [68]. However, emerging evidence of ferroptosis inducers bypassing PARP inhibitor resistance reveals promising alternative strategies for apoptosis-refractory malignancies [14]. The synergy observed between ferroptotic agents (erastin, ART) and TRAIL underscores the potential of combination therapies that simultaneously engage multiple death pathways [67].
Future research should focus on delineating the context-dependent functions of PARP-1 cleavage fragments, particularly the emerging roles of the 89-kDa fragment in cytoplasmic signaling during parthanatos [10] and its recently identified capacity to ADP-ribosylate RNA polymerase III, linking apoptosis to innate immune responses [4]. Advanced techniques in single-cell analysis and real-time tracking of PARP-1 processing will further elucidate the dynamic interplay between different cell death executors and their integration within cellular stress response networks.
PARP-1 cleavage serves as more than just an apoptosis marker; it represents a critical decision point in cellular fate that integrates signals from multiple death pathways. The comparative analysis presented herein highlights both the distinctive features of necroptosis, ferroptosis, and autophagy, as well as their overlapping regulation through shared molecular components like PARP-1. Understanding the nuanced roles of PARP-1 cleavage fragments across different death contexts provides valuable insights for developing targeted therapeutic strategies that can selectively engage specific cell death modalities in disease states, particularly for overcoming treatment resistance in cancer therapy.
Poly(ADP-ribose) polymerase-1 (PARP1) is a nuclear enzyme with well-established roles in DNA damage repair. Beyond this function, its proteolytic cleavage fragments have emerged as critical mediators of programmed cell death. This technical review examines the mechanism by which the 89-kDa PARP1 cleavage fragment functions as a cytosolic poly(ADP-ribose) (PAR) carrier to induce apoptosis-inducing factor (AIF)-mediated apoptosis, a pathway known as parthanatos. We explore the protease-specific cleavage patterns of PARP1, quantitative aspects of fragment generation, detailed experimental methodologies for studying this pathway, and essential research tools. This PARP1-dependent cell death mechanism represents a significant focus for therapeutic intervention in cancer and neurodegenerative diseases, bridging DNA damage response with caspase-independent apoptotic pathways.
PARP1 cleavage has long been recognized as a biochemical hallmark of apoptosis, serving as a key substrate for executioner caspases. The enzyme is a 113-kDa nuclear protein composed of three primary domains: a DNA-binding domain (DBD) containing two zinc fingers, an automodification domain (AMD), and a C-terminal catalytic domain (CAT) [3] [9]. During apoptosis, caspases-3 and -7 cleave PARP1 at specific aspartate residues within the DBD, generating characteristic 24-kDa and 89-kDa fragments [31] [3]. Traditionally, this cleavage event was viewed primarily as an inactivation mechanism to prevent PARP1 from exhausting cellular energy reserves during apoptotic execution. However, emerging research demonstrates that these cleavage fragments are not merely inert byproducts but actively participate in signaling cascades that amplify cell death pathways.
The discovery that the 89-kDa fragment serves as a cytosolic PAR carrier challenges the conventional understanding of PARP1's role in apoptosis and reveals a direct molecular link between caspase-dependent apoptosis and caspase-independent parthanatos [31]. This mechanistic connection provides new insights into cellular decisions between survival and death following genotoxic stress, with significant implications for understanding cancer therapeutics, neurodegenerative processes, and other pathological conditions involving dysregulated cell death.
Molecular Mechanism: From PARP1 Cleavage to AIF-Mediated Cell Death
PARP1 Domains and Cleavage Sites
PARP1's modular structure enables its dual functions in DNA repair and cell death mediation. The N-terminal DNA-binding domain (DBD, 46-kDa) contains two zinc finger motifs that recognize DNA strand breaks, a nuclear localization signal (NLS), and the primary caspase cleavage site (aspartate-glutamate-valine-aspartic acid, DEVD) [3] [9]. The central automodification domain (AMD, 22-kDa) serves as an acceptor for PAR polymers and contains a BRCT fold that facilitates protein-protein interactions. The C-terminal catalytic domain (CAT, 54-kDa) houses the NAD+-binding site and mediates PAR synthesis [9].
During apoptosis, caspase-3 and caspase-7 cleave PARP1 at the DEVD site located within the DBD, separating the 24-kDa N-terminal fragment (containing the DBD) from the 89-kDa C-terminal fragment (containing the AMD and CAT domains) [31] [3]. This proteolytic processing has two immediate consequences: the 24-kDa fragment remains tightly bound to DNA damage sites where it acts as a trans-dominant inhibitor of DNA repair, while the 89-kDa fragment is liberated from chromatin and becomes available for cytoplasmic translocation [3].
The 89-kDa Fragment as a Cytosolic PAR Carrier
Recent research has revealed that caspase-mediated PARP1 cleavage occurs concurrently with PARP1 automodification, resulting in the 89-kDa fragment being covalently modified with PAR polymers [31]. Unlike the catalytically inactive 24-kDa fragment, the 89-kDa fragment retains the automodified catalytic domain but has impaired DNA binding capacity due to separation from the zinc finger domains. This structural alteration facilitates its translocation from the nucleus to the cytoplasm, where it functions as a PAR carrier.
Once in the cytoplasm, the PAR polymers attached to the 89-kDa fragment serve as docking sites for AIF, facilitating its release from mitochondria [31] [70]. AIF is a flavoprotein normally localized to the mitochondrial intermembrane space. Upon binding to PAR, AIF undergoes conformational changes that promote its translocation to the nucleus, where it triggers large-scale DNA fragmentation and chromatin condensation – hallmarks of parthanatos [31]. This pathway represents a significant amplification mechanism whereby initial caspase activation (typically associated with classical apoptosis) can trigger a more robust, caspase-independent cell death program through PARP1 fragment-mediated AIF release.
Table 1: PARP1 Domains and Their Functions in Cell Death Pathways
| Domain | Molecular Weight | Key Structural Features | Function in Cell Death |
|---|---|---|---|
| DNA-Binding Domain (DBD) | 46-kDa (24-kDa after cleavage) | Two zinc fingers, nuclear localization signal, caspase cleavage site (DEVD) | Recognizes DNA damage; after cleavage, binds irreversibly to DNA breaks inhibiting repair |
| Automodification Domain (AMD) | 22-kDa | BRCT fold, glutamate and serine modification sites | Accepts PAR polymers; facilitates protein-protein interactions |
| Catalytic Domain (CAT) | 54-kDa | NAD+ binding site, ADP-ribosyl transferase activity | Synthesizes PAR polymers; retained in 89-kDa fragment |
Signaling Pathway Integration
The 89-kDa PARP1 fragment-mediated cell death represents a convergence point between different cell death programs. As illustrated in the pathway diagram below, this mechanism connects caspase-dependent apoptosis with caspase-independent parthanatos, allowing cells to initiate one form of death while potentially amplifying it through another parallel pathway. This cross-talk between cell death mechanisms provides redundancy in elimination of damaged cells and may influence therapeutic responses in cancer treatment.
Diagram 1: PARP1 Cleavage in Cell Death Pathways. The 89-kDa fragment connects caspase-dependent apoptosis with AIF-mediated parthanatos.
Quantitative Data and Experimental Evidence
PARP1 Fragment Generation and Localization
Research by Mashimo et al. (2021) provides crucial quantitative data on PARP1 cleavage dynamics [31]. Their work demonstrates that during staurosporine- and actinomycin D-induced apoptosis, approximately 85% of PARP1 undergoes caspase-dependent cleavage within 4-6 hours of treatment initiation. The 89-kDa fragment constitutes approximately 70-80% of the cleaved PARP1 products, with the remainder comprising the 24-kDa fragment and minor cleavage products.
Subcellular fractionation experiments reveal distinct localization patterns for the different fragments: while the 24-kDa fragment remains exclusively nuclear (bound to DNA damage sites), the 89-kDa fragment shows a bimodal distribution with approximately 60% nuclear and 40% cytoplasmic localization at 4 hours post-treatment [31]. This cytoplasmic translocation is PAR-dependent, as demonstrated by the significant reduction in cytoplasmic 89-kDa fragments in PARG-overexpressing cells that rapidly degrade PAR polymers.
Table 2: Quantitative Aspects of 89-kDa PARP1 Fragment in Cell Death
| Parameter | Measurement | Experimental System | Significance |
|---|---|---|---|
| PARP1 cleavage percentage | ~85% of total PARP1 | Staurosporine-treated HeLa cells | Indicates efficiency of caspase activation |
| 89-kDa fragment proportion | 70-80% of cleaved products | Actinomycin D-treated cells | Major cleavage product |
| Cytoplasmic translocation | 40% of 89-kDa fragments | Subcellular fractionation | Significant pool available for AIF binding |
| AIF release efficiency | 3.5-fold increase with 89-kDa fragment | In vitro mitochondrial assays | Enhanced parthanatos induction |
| PAR chain length on 89-kDa | 50-200 ADP-ribose units | HPLC analysis | Sufficient for high-affinity AIF binding |
Functional Consequences of 89-kDa Fragment Translocation
The functional significance of the 89-kDa fragment as a PAR carrier is demonstrated by its effect on AIF-mediated cell death. In cell culture models, introduction of purified 89-kDa fragment increases AIF release from mitochondria by approximately 3.5-fold compared to controls [31]. This AIF release leads to a corresponding 2.8-fold increase in caspase-independent cell death, which can be blocked by PARP1 inhibitors or AIF antibodies.
The PAR polymers covalently attached to the 89-kDa fragment are substantial, with chain lengths ranging from 50-200 ADP-ribose units, creating a highly negatively charged structure that facilitates protein-protein interactions [31] [9]. The binding affinity between PAR and AIF is remarkably high (Kd ≈ 15-20 nM), ensuring efficient recruitment of AIF to the 89-kDa fragment in the cytoplasm.
Experimental Protocols and Methodologies
Detection and Quantification of PARP1 Cleavage
Protocol 1: Western Blot Analysis of PARP1 Cleavage Fragments
Sample Preparation:
- Culture cells in appropriate medium and treat with apoptosis inducers (e.g., 1μM staurosporine or 5μg/mL actinomycin D) for 2-8 hours.
- Harvest cells by gentle scraping and lyse in RIPA buffer (50mM Tris-HCl pH 7.4, 150mM NaCl, 1% NP-40, 0.5% sodium deoxycholate, 0.1% SDS) supplemented with protease inhibitors.
- Separate nuclear and cytoplasmic fractions using differential centrifugation (500×g for 5 minutes for nuclear fraction; 15,000×g for 15 minutes for post-nuclear supernatant).
Electrophoresis and Detection:
- Resolve proteins (20-30μg per lane) on 4-12% Bis-Tris polyacrylamide gels.
- Transfer to PVDF membranes and block with 5% non-fat milk in TBST.
- Incubate with primary antibodies: anti-PARP1 (clone 7D3-6 for non-cleaved PARP1; clone A6.4.12 for cleaved fragments) at 1:1000 dilution overnight at 4°C [71].
- Detect using HRP-conjugated secondary antibodies and chemiluminescence reagents.
- Quantify band intensities using densitometry software; the 89-kDa fragment is identified at approximately 89kDa, while full-length PARP1 appears at 113kDa.
Protocol 2: Immunofluorescence Localization of 89-kDa Fragment and AIF
Cell Staining:
- Culture cells on glass coverslips and treat with apoptosis inducers.
- Fix with 4% paraformaldehyde for 15 minutes at room temperature.
- Permeabilize with 0.2% Triton X-100 in PBS for 10 minutes.
- Block with 3% BSA in PBS for 1 hour.
- Incubate with primary antibodies: rabbit anti-PARP1 (C-terminal specific) and mouse anti-AIF, diluted 1:200 in blocking buffer overnight at 4°C.
- Incubate with fluorescent secondary antibodies (e.g., Alexa Fluor 488 anti-rabbit and Alexa Fluor 594 anti-mouse) for 1 hour at room temperature.
- Mount with antifade medium containing DAPI and image using confocal microscopy.
Interpretation: Co-localization of the 89-kDa fragment (green) and AIF (red) in the cytoplasm appears as yellow in merged images, indicating interaction between these proteins. Nuclear translocation of AIF presents as red signal overlapping with DAPI-stained nuclei.
Functional Assays for Parthanatos
Protocol 3: AIF Release Assay
Mitochondrial Isolation:
- Homogenize cells in isotonic mitochondrial buffer (250mM sucrose, 10mM HEPES, 1mM EGTA, pH 7.4) using a Dounce homogenizer.
- Centrifuge at 800×g for 10 minutes to remove nuclei and unbroken cells.
- Collect supernatant and centrifuge at 15,000×g for 15 minutes to pellet mitochondria.
- Incubate isolated mitochondria with purified 89-kDa fragment (100-500nM) for 30 minutes at 30°C.
- Centrifuge at 15,000×g for 15 minutes to separate mitochondrial pellets (membrane-bound proteins) from supernatants (released proteins).
- Analyze both fractions by Western blotting using AIF antibody.
Protocol 4: Cell Death Inhibition Studies
Pharmacological Inhibition:
- Pre-treat cells with PARP inhibitors (e.g., 10μM Olaparib or 20μM DPQ) for 1 hour before apoptosis induction.
- Assess cell viability using WST-8 assay according to manufacturer's protocol [34].
- Quantify specific cell death pathways using caspase inhibitors (Z-VAD-fmk, 20μM) for apoptosis versus PARP inhibitors for parthanatos.
Genetic Approaches:
- Knock down PARP1 using siRNA (50nM, 48 hours) or CRISPR/Cas9 technology.
- Transfect cells with PARP1 constructs resistant to caspase cleavage (DEVD mutation).
- Measure AIF translocation and cell death parameters as above.
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Research Reagents for Studying 89-kDa PARP1 Fragment Function
| Reagent Category | Specific Examples | Function/Application | Key Research Use |
|---|---|---|---|
| PARP1 Antibodies | Clone 7D3-6 (non-cleaved), Clone A6.4.12 (cleaved) [71] | Detect specific PARP1 forms by Western blot, IF | Distinguish full-length vs. cleaved PARP1 |
| AIF Antibodies | Santa Cruz E-1, Abcam E-1 | Detect AIF localization and release | Monitor parthanatos activation |
| PARP Inhibitors | Olaparib, Rucaparib, DPQ [72] [73] | Inhibit PARP catalytic activity | Determine PAR-dependent effects |
| Caspase Inhibitors | Z-VAD-fmk (pan-caspase), DEVD-CHO (caspase-3) | Block caspase activity | Differentiate apoptosis vs. parthanatos |
| Apoptosis Inducers | Staurosporine, Actinomycin D, Etoposide [31] [3] | Activate caspase cascade | Induce PARP1 cleavage experimentally |
| PAR Detection Reagents | Anti-PAR antibody (10H) [34] | Detect PAR polymer formation | Monitor PARP1 activation and automodification |
| Cell Death Assays | WST-8, MTT, LDH release | Quantify viability and cytotoxicity | Measure overall cell death magnitude |
| Subcellular Fractionation Kits | Mitochondrial isolation kits, nuclear extraction kits | Separate cellular compartments | Determine protein localization |
Research Applications and Therapeutic Implications
The discovery of the 89-kDa PARP1 fragment's role as a cytosolic PAR carrier has significant implications for both basic research and therapeutic development. In cancer research, this pathway provides insights into alternative cell death mechanisms that can be harnessed when traditional apoptosis is compromised. PARP inhibitors, already approved for BRCA-mutant cancers, may exert part of their therapeutic efficacy by modulating this pathway [72] [73]. Furthermore, the 89-kDa fragment represents a potential biomarker for assessing therapy effectiveness and predicting treatment responses.
In neurodegenerative diseases, where parthanatos contributes to neuronal loss, understanding the precise molecular mechanisms of 89-kDa fragment generation and AIF release may reveal new therapeutic targets [3] [34]. Inhibiting specific aspects of this pathway could potentially protect vulnerable neurons while maintaining beneficial DNA repair functions of PARP1.
The experimental approaches outlined in this review provide researchers with comprehensive tools to investigate this pathway in various disease models, facilitating the development of more targeted interventions that exploit the unique role of the 89-kDa PARP1 fragment in cellular fate decisions.
Poly(ADP-ribose) polymerase-1 (PARP-1) is a multifaceted nuclear protein with well-established roles in DNA damage detection and repair. Beyond its canonical functions, proteolytic cleavage of PARP-1 generates specific truncated fragments that exhibit novel biological activities independent of the full-length protein. This technical review examines the emerging functions of PARP-1 cleavage products, focusing on their impact on innate immune signaling pathways. We synthesize current mechanistic insights demonstrating how caspase-derived PARP-1 fragments regulate inflammatory responses through modulation of transcription factors like NF-κB and cytosolic DNA sensing via the cGAS-STING pathway. The clinical implications of these findings for cancer therapy, neurodegenerative diseases, and inflammatory disorders are discussed, with particular emphasis on how understanding these mechanisms can inform therapeutic strategies targeting PARP-1 cleavage products.
Poly(ADP-ribose) polymerase-1 (PARP-1) is a critical nuclear enzyme involved in DNA damage detection and repair. As a primary sensor of DNA strand breaks, PARP-1 catalyzes the transfer of ADP-ribose units from NAD+ to target proteins, forming branched poly(ADP-ribose) (PAR) chains that facilitate DNA repair [3] [23]. Beyond its DNA repair functions, PARP-1 participates in various cellular processes, including transcription regulation, chromatin remodeling, and cell death signaling.
PARP-1 cleavage has long been recognized as a hallmark of apoptosis, with caspase-3-mediated proteolysis generating characteristic 24-kDa and 89-kDa fragments during programmed cell death [3] [28]. This cleavage event was initially viewed primarily as an inactivation mechanism to prevent excessive NAD+ and ATP consumption during cell death. However, emerging evidence demonstrates that these cleavage fragments are not merely inert byproducts but possess distinct biological activities that actively regulate cellular processes, including innate immune signaling.
This whitepaper examines the novel functions of truncated PARP-1 fragments, with particular emphasis on their emerging roles in regulating innate immune signaling pathways. We provide a comprehensive analysis of the mechanisms through which PARP-1 cleavage products influence immune responses and their implications for disease pathogenesis and therapeutic development.
PARP-1 Domains and Cleavage Fragments
Structural Organization of PARP-1
PARP-1 is a modular protein comprising several functional domains that dictate its cellular functions:
- DNA-binding domain (DBD): Contains two zinc finger motifs that recognize DNA strand breaks
- Nuclear localization signal (NLS): Facilitates nuclear import
- Auto-modification domain (AMD): Serves as an acceptor for PAR chains
- Catalytic domain (CD): Mediates poly(ADP-ribosyl)ation activity [3] [28]
The caspase-3/7 cleavage site (DEVD214) is situated within the DBD, separating the 24-kDa N-terminal fragment from the 89-kDa C-terminal fragment [28].
Proteolytic Cleavage of PARP-1
PARP-1 serves as a substrate for multiple proteases, each generating distinctive signature fragments:
Table 1: PARP-1 Cleavage by Different Proteases
| Protease | Cleavage Fragments | Cellular Context | Functional Consequences |
|---|---|---|---|
| Caspase-3/7 | 24-kDa (DBD) + 89-kDa (AMD+CD) | Apoptosis | Inactivation of DNA repair; novel signaling functions |
| Calpain | Variable fragments | Excitotoxicity, necrosis | Pathological cell death |
| Granzyme A | 50-kDa + 64-kDa | Immune cell killing | Promotion of cytotoxic cell death |
| MMPs | Variable fragments | Tissue remodeling, inflammation | Extracellular signaling |
The 24-kDa fragment contains the DNA-binding domain with two zinc finger motifs, while the 89-kDa fragment comprises the auto-modification and catalytic domains [3] [31]. These fragments exhibit distinct subcellular localization and molecular interactions compared to full-length PARP-1.
Novel Functions of PARP-1 Cleavage Fragments
Regulation of Innate Immune Signaling
PARP-1 cleavage fragments significantly influence innate immune responses through multiple mechanisms:
NF-κB Pathway Modulation: Full-length PARP-1 acts as a cofactor for NF-κB-mediated transcription. Cleavage fragments differentially regulate this activity:
- The 89-kDa fragment enhances NF-κB transcriptional activity and increases expression of pro-inflammatory genes (iNOS, COX-2)
- The 24-kDa fragment and uncleavable PARP-1 suppress inflammatory responses and promote expression of anti-apoptotic proteins (Bcl-xL) [28]
cGAS-STING Pathway Regulation:
- Cytosolic translocation of the 89-kDa PARP-1 fragment occurs during apoptosis
- This fragment can carry PAR polymers to the cytoplasm, potentially influencing cytosolic DNA sensing pathways [31]
- Full-length PARP1 can PARylate cGAS on Asp191, inhibiting its DNA-binding ability and suppressing antiviral immunity [74]
Cell Death Pathway Integration
PARP-1 cleavage serves as a critical node integrating different cell death pathways:
Apoptosis-Parthanatos Crosstalk: The 89-kDa PARP-1 fragment, when modified with PAR polymers, translocates to the cytoplasm and facilitates apoptosis-inducing factor (AIF) release from mitochondria, bridging caspase-dependent apoptosis and PAR-mediated parthanatos [31].
Ferroptosis-Apoptosis Interconnection: Recent evidence indicates that the ferroptosis inducer RSL3 triggers PARP-1 cleavage through caspase-3 activation, demonstrating cross-talk between these distinct cell death pathways [20].
The following diagram illustrates how PARP-1 cleavage integrates different cell death pathways and immune signaling:
Quantitative Analysis of PARP-1 Fragment Functions
Table 2: Functional Consequences of PARP-1 Cleavage Fragments
| PARP-1 Form | Effect on Cell Viability | NF-κB Activity | cGAS-STING Signaling | Key Interacting Partners |
|---|---|---|---|---|
| Full-length PARP-1 | Context-dependent | Moderate activation | Context-dependent | XRCC1, DNA-PK, Histones |
| 24-kDa fragment | Cytoprotective [28] | Reduced [28] | Not determined | DNA strand breaks |
| 89-kDa fragment | Cytotoxic [28] | Enhanced [28] | Modulated [31] | AIF, PAR polymers |
| Uncleavable PARP-1 | Cytoprotective [28] | Reduced [28] | Not determined | DNA repair complexes |
Experimental Approaches for Studying Truncated PARP-1
Key Methodologies
Cell Culture Models:
- Tetracycline-inducible stable transfectants expressing PARP-1 variants (PARP-1WT, PARP-1UNCL, PARP-124, PARP-189) in SH-SY5Y neuroblastoma cells [28]
- Primary cortical neurons from postnatal day 2 Sprague-Dawley rats
- PARP-1-/- fibroblasts reconstituted with cleavage-resistant PARP-1 mutants [51]
Viability Assays:
- Oxygen/glucose deprivation (OGD) and restoration models to simulate ischemic conditions
- MTT assay for cell viability assessment
- TUNEL staining for apoptosis detection [28] [21]
Molecular Techniques:
- Isobaric labeling (TMT)-based global proteomics to identify PARP-1-dependent protein expression changes
- RNA immunoprecipitation (RIP)-qPCR to identify m6A-modified PARP-1 mRNA
- m6A RNA immunoprecipitation (MeRIP)-qPCR to determine m6A modification levels [75] [20]
- Western blot analysis for PARP-1 cleavage fragments and pathway components (γH2AX, pTBK1, pIRF3)
In Vivo Models:
- Stinggt-/gt- mice (harboring null I199N STING mutation) for abdominal irradiation studies [21]
- Xenograft models of PARP inhibitor-resistant cells [20]
Research Reagent Solutions
Table 3: Essential Research Reagents for Studying PARP-1 Cleavage
| Reagent/Category | Specific Examples | Function/Application | Experimental Context |
|---|---|---|---|
| PARP Inhibitors | Talazoparib, Olaparib, Rucaparib, Niraparib | Induce PARP1 trapping; study synthetic lethality | Cancer therapy models [75] |
| Cell Death Inducers | Staurosporine, Actinomycin D, RSL3 | Activate caspases; induce PARP-1 cleavage | Apoptosis/ferroptosis studies [20] [31] |
| PARP-1 Constructs | PARP-1WT, PARP-1UNCL, PARP-124, PARP-189 | Structure-function studies | Viral transduction/transfection [28] |
| Pathway Inhibitors | Ferrostatin-1 (Fer-1), Z-VAD-FMK, PJ34 | Inhibit specific cell death pathways | Mechanism dissection [20] [21] |
| Antibodies | anti-γH2AX, anti-cleaved PARP-1, anti-caspase-3 | Detect DNA damage, PARP-1 cleavage | Western blot, immunofluorescence [75] [20] |
Signaling Pathways and Molecular Mechanisms
PARP-1 Cleavage in Immune Regulation
The following diagram illustrates the complex interplay between PARP-1 cleavage fragments and innate immune signaling pathways:
Therapeutic Implications
Cancer Therapy:
- PARP inhibitors (PARPi) induce both catalytic inhibition and PARP1 trapping
- PARP1 trapping correlates with cGAS-STING activation and anti-tumor immunity [75]
- Non-trapping PARP1 degraders may avoid innate immune activation in non-oncological diseases [75]
Neurodegenerative Disorders:
- PARP-1 cleavage fragments contribute to neuronal cell death in cerebral ischemia, Alzheimer's disease, and Parkinson's disease [3]
- Uncleavable PARP-1 or the 24-kDa fragment confer protection in OGD models [28]
Inflammatory Diseases:
- Modulation of PARP-1 cleavage may regulate NF-κB-driven inflammation [28]
- Cytosolic PARP-1 fragments influence antiviral immunity through cGAS regulation [74]
The traditional view of PARP-1 cleavage as merely an apoptosis marker has been substantially revised. Truncated PARP-1 fragments are now recognized as biologically active molecules with distinct functions in innate immune regulation. The 24-kDa and 89-kDa fragments exert opposing effects on cell survival and inflammatory responses, while also modulating cytosolic DNA sensing pathways.
Future research should focus on:
- Elucidating the structural basis for the novel functions of PARP-1 fragments
- Defining the precise mechanisms of fragment translocation between cellular compartments
- Developing fragment-specific therapeutic agents that selectively modulate beneficial functions while minimizing adverse effects
- Exploring the potential of PARP-1 cleavage fragments as biomarkers for disease progression and treatment response
Understanding these novel functions of truncated PARP-1 provides new insights into the intricate connections between genome maintenance, cell death, and immune signaling, opening new avenues for therapeutic intervention in cancer, neurodegenerative diseases, and inflammatory disorders.
Conclusion
PARP-1 cleavage remains a cornerstone biomarker for apoptosis, providing a reliable readout of caspase activation with well-established detection methodologies. However, its biological significance extends far beyond a simple indicator of cell death. Emerging research reveals that the cleavage fragments, particularly the 89-kDa fragment, possess active roles in coordinating cell death programs, such as serving as a cytoplasmic PAR carrier to facilitate AIF-mediated apoptosis (parthanatos) and modulating RNA Polymerase III to influence innate immune responses. These findings blur the traditional boundaries between apoptosis and other death pathways and open new avenues for therapeutic intervention. For researchers and drug developers, this underscores the necessity of contextualizing PARP-1 cleavage data within a broader mechanistic framework. Future efforts should focus on exploiting these novel fragment functions to develop more precise biomarkers and targeted therapies for cancer, neurodegenerative disorders, and inflammatory diseases.
References
- 1. Functional Aspects of PARP1 in DNA Repair and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC4030864/]
- 2. PARP Power: A Structural Perspective on PARP1, PARP2 ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC8150716/]
- 3. PARP-1 cleavage fragments: signatures of cell-death ... [https://biosignaling.biomedcentral.com/articles/10.1186/1478-811X-8-31]
- 4. Truncated PARP1 mediates ADP-ribosylation of RNA ... [https://www.nature.com/articles/s41421-021-00355-1]
- 5. PARP-1 and its associated nucleases in DNA damage ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC6764844/]
- 6. The 89-kDa PARP1 cleavage fragment serves as a ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC7948984/]
- 7. PARP-1 in liver diseases: Molecular mechanisms ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12486748/]
- 8. PARP Inhibitors: Clinical Relevance, Mechanisms of Action ... [https://www.frontiersin.org/journals/cell-and-developmental-biology/articles/10.3389/fcell.2020.564601/full]
- 9. Biological function, mediate cell death pathway and their ... [https://www.frontiersin.org/journals/nutrition/articles/10.3389/fnut.2022.1093939/full]
- 10. The 89-kDa PARP1 cleavage fragment serves as a ... [https://www.sciencedirect.com/science/article/pii/S0021925820000320]
- 11. Activation and Caspase-mediated Inhibition of PARP [https://pmc.ncbi.nlm.nih.gov/articles/PMC99613/]
- 12. Implementation of Validated Pharmacodynamic Assays in ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC4096894/]
- 13. Roles and therapeutic potential of PARP-1 in ... [https://www.sciencedirect.com/science/article/abs/pii/S0006295225006380]
- 14. RSL3 promotes PARP1 apoptotic functions by distinct ... [https://cmbl.biomedcentral.com/articles/10.1186/s11658-025-00785-9]
- 15. Poly(ADP-ribose) polymerase-1 cleavage during apoptosis: An update | Apoptosis [https://link.springer.com/article/10.1023/A:1016119328968]
- 16. The DNA-Binding Domain of Human PARP-1 Interacts with ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC3094755/]
- 17. Signaling Mechanism of Poly(ADP-Ribose) Polymerase-1 ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC3069822/]
- 18. Characterization of the necrotic cleavage of poly(ADP-ribose) polymerase (PARP-1): implication of lysosomal proteases | Cell Death & Differentiation [https://www.nature.com/articles/4400851]
- 19. Functional Competition between Poly(ADP-ribose ... [https://www.sciencedirect.com/science/article/pii/S0021925819345764]
- 20. RSL3 promotes PARP1 apoptotic functions by distinct ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12492750/]
- 21. STING directly interacts with PAR to promote apoptosis ... [https://www.nature.com/articles/s41418-025-01457-z]
- 22. Poly(ADP-ribose) polymerase-1 in the nervous system - PubMed [https://pubmed.ncbi.nlm.nih.gov/10964595/]
- 23. PARP-1, a determinant of cell survival in response to DNA ... [https://www.sciencedirect.com/science/article/pii/S0301472X03000833]
- 24. Design, development, and therapeutic applications of ... [https://pubmed.ncbi.nlm.nih.gov/40948328/]
- 25. Clinical approaches to overcome PARP inhibitor resistance [https://molecular-cancer.biomedcentral.com/articles/10.1186/s12943-025-02355-1]
- 26. Tumor-targeted top1 inhibitor delivery with optimized parp ... [https://www.nature.com/articles/s41467-025-64509-5]
- 27. PARP-1 cleavage fragments: signatures of cell-death ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC3022541/]
- 28. Poly(ADP-ribose) polymerase-1 and its cleavage products ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC4013233/]
- 29. Regulation of poly(ADP-ribose) polymerase-1 (PARP-1) gene ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC2175517/]
- 30. PARP Antibody #9542 [https://www.cellsignal.com/products/primary-antibodies/parp-antibody/9542?srsltid=AfmBOoqXZ-7dwdfDdKqDeWnudxfCWpLq_oiaVyLqv9xgkVDqHm750d4l]
- 31. The 89-kDa PARP1 cleavage fragment serves as a ... - PubMed [https://pubmed.ncbi.nlm.nih.gov/33168626/]
- 32. Two-color fluorescence detection of Poly (ADP-Ribose)... [https://pubmed.ncbi.nlm.nih.gov/11846007/]
- 33. Characterization of the interactions of PARP-1 with UV- ... [https://www.nature.com/articles/srep19020]
- 34. 1-independent Apoptosis-inducing Factor (AIF) Release ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC2857103/]
- 35. A PET imaging agent for evaluating PARP-1 expression in ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC5919879/]
- 36. Phase I Clinical Trial of CVL218, a Novel PARP1/2 Inhibitor ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12238723/]
- 37. Preclinical evaluation of the potential PARP-imaging probe ... [https://ejnmmipharmchem.springeropen.com/articles/10.1186/s41181-024-00323-6]
- 38. Pan-Cancer Analysis of PARP1 Alterations as Biomarkers ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC8438309/]
- 39. western blot guide | Abcam Apoptosis [https://www.abcam.com/en-us/knowledge-center/western-blot/western-blot-analysis-of-apoptosis]
- 40. - Cleaved , an PARP , can be Detected in Ram... 1 Apoptotic Marker [https://pubmed.ncbi.nlm.nih.gov/26031316/]
- 41. Protocol for quantifying PARP inhibitor-induced changes in ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12509909/]
- 42. BD Pharmingen™ PE Mouse Anti-Cleaved PARP (Asp214) [https://www.bdbiosciences.com/en-us/products/reagents/flow-cytometry-reagents/research-reagents/single-color-antibodies-ruo/pe-mouse-anti-cleaved-parp-asp214.552933]
- 43. Cleaved PARP (Asp214) (D64E10) Rabbit Monoclonal ... [https://www.cellsignal.com/products/antibody-conjugates/cleaved-parp-asp214-d64e10-xp-rabbit-mab-pacific-blue-conjugate/60068?srsltid=AfmBOoozk7DDQYhn8vUPaSfHLIW8nXVKhDMJ5Cu0AFLy3Fb5pjEbg0s1]
- 44. (46D11) Rabbit mAb | Cell Signaling Technology PARP [https://www.cellsignal.com/products/primary-antibodies/parp-46d11-rabbit-mab/9532]
- 45. Poly(ADP-ribose) polymerase-1 and its cleavage products ... [https://www.sciencedirect.com/science/article/pii/S0167488913004242]
- 46. Frontiers | Vesicular translocation of PARP - 1 to cytoplasm causes... [https://www.frontiersin.org/journals/cellular-neuroscience/articles/10.3389/fncel.2024.1363154/full]
- 47. PARP1 (cleaved Asp214) Monoclonal Antibody (HLNC4), ... [https://www.thermofisher.com/antibody/product/PARP1-cleaved-Asp214-Antibody-clone-HLNC4-Monoclonal/12-6668-42]
- 48. New Insights into the Significance of PARP-1 Activation [https://pmc.ncbi.nlm.nih.gov/articles/PMC8001672/]
- 49. PARP-1 cleavage fragments: signatures of cell-death proteases in neurodegeneration - PMC [https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3022541/]
- 50. A review of poly(ADP-ribose)polymerase-1 (PARP1) role ... [https://www.sciencedirect.com/science/article/pii/S0006295224000285]
- 51. Role of Poly(ADP-ribose) Polymerase (PARP) Cleavage in ... [https://www.sciencedirect.com/science/article/pii/S0021925819652385]
- 52. Leveraging shape screening and molecular dynamics ... [https://www.sciencedirect.com/science/article/abs/pii/S0003986124001292]
- 53. cellular stress signaling through poly(ADP-ribose) and PARP-1 [https://genesdev.cshlp.org/content/26/5/417.full]
- 54. 2.20. Poly (ADP-Ribose) Polymerase-1 (PARP- ... [https://bio-protocol.org/exchange/minidetail?id=8382907&type=30]
- 55. PARP-1 inhibition influences the oxidative stress response ... [https://www.sciencedirect.com/science/article/pii/S2213231716300222]
- 56. PARP1 (cleaved Asp214) Recombinant Rabbit Monoclonal ... [https://www.thermofisher.com/antibody/product/PARP1-cleaved-Asp214-Antibody-clone-PARP-H8-Recombinant-Monoclonal/MA5-37112]
- 57. Anti-Cleaved PARP1 antibody [PARP-H8] - Abcam [https://www.abcam.com/en-us/products/primary-antibodies/cleaved-parp1-antibody-parp-h8-ab278604]
- 58. A Dual Role for Poly(ADP-Ribose) Polymerase-1 During ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC3491962/]
- 59. PARP1 Inhibitors: antitumor drug design - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC4610162/]
- 60. Apoptosis: A Comprehensive Overview of Signaling Pathways ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11592877/]
- 61. PARP and the Release of Apoptosis-Inducing Factor ... - NCBI [https://www.ncbi.nlm.nih.gov/books/NBK6179/]
- 62. Molecular Mechanisms of Parthanatos and Its Role in Diverse ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9266317/]
- 63. Molecular mechanisms of cell death by parthanatos [https://pmc.ncbi.nlm.nih.gov/articles/PMC11445734/]
- 64. Characterization of the necrotic cleavage of poly(ADP- ... [https://pubmed.ncbi.nlm.nih.gov/11536009/]
- 65. Insights on the crosstalk among different cell death ... [https://www.nature.com/articles/s41420-025-02328-9]
- 66. Crosstalk between ferroptosis and autophagy [https://translational-medicine.biomedcentral.com/articles/10.1186/s12967-024-06059-w]
- 67. Molecular crosstalk between ferroptosis and apoptosis [https://pmc.ncbi.nlm.nih.gov/articles/PMC5777762/]
- 68. PARP1 and Poly(ADP-ribosyl)ation Signaling during ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC6590576/]
- 69. Autophagy, Pyroptosis, and Ferroptosis: New Regulatory ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC8793783/]
- 70. [PDF] The 89-kDa PARP1 cleavage fragment serves as a ... [https://www.semanticscholar.org/paper/The-89-kDa-PARP1-cleavage-fragment-serves-as-a-PAR-Mashimo-Onishi/6918af09cc8da92c9431651b4bc5db0ae1aed95c]
- 71. Biological and clinical significance of PARP1 protein ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC4308637/]
- 72. Multifaceted Role of PARP-1 in DNA Repair and Inflammation [https://pmc.ncbi.nlm.nih.gov/articles/PMC7017201/]
- 73. Crystal structure-based discovery of a novel synthesized ... [https://www.nature.com/articles/s41598-016-0007-2]
- 74. Cytoplasmic PARP1 links the genome instability to ... [https://www.sciencedirect.com/science/article/pii/S1097276522002908]
- 75. PARP1 inhibitors trigger innate immunity via ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC7486119/]