Surface Engineering of Mitochondria with Cell-Penetrating Peptides for Enhanced Targeted Delivery
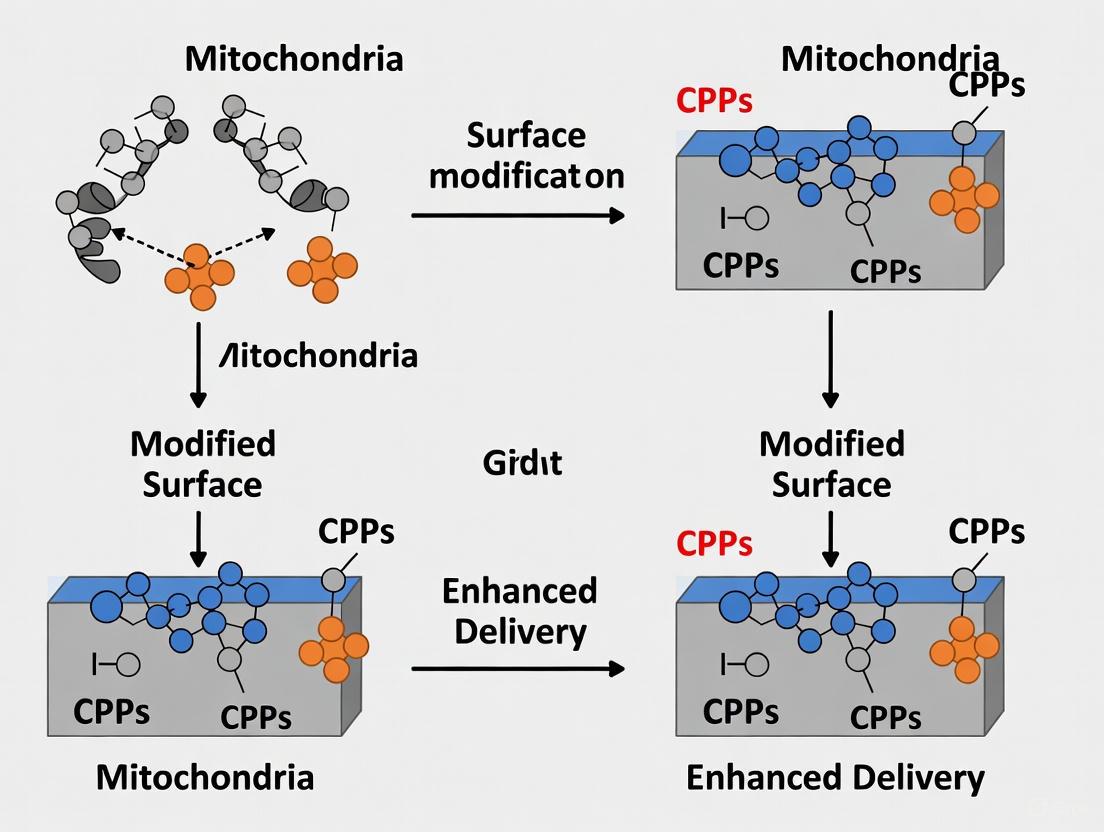
This article explores the cutting-edge biotechnological strategy of surface-modifying mitochondria with Cell-Penetrating Peptides (CPPs) to overcome critical barriers in therapeutic mitochondrial transplantation.
Surface Engineering of Mitochondria with Cell-Penetrating Peptides for Enhanced Targeted Delivery
Abstract
This article explores the cutting-edge biotechnological strategy of surface-modifying mitochondria with Cell-Penetrating Peptides (CPPs) to overcome critical barriers in therapeutic mitochondrial transplantation. We examine the foundational principles of mitochondrial dysfunction in diseases and the limitations of current delivery methods. The content delves into innovative surface engineering techniques, including lipid-polymer coatings and novel amphipathic CPP designs, that enhance cellular uptake, specificity, and bioenergetic integration. We further analyze challenges in optimization, scalability, and immune response, while evaluating functional validation through advanced metabolic assays and comparative studies with other targeting moieties. This synthesis provides researchers and drug development professionals with a comprehensive roadmap for developing clinically viable, mitochondria-based nanotherapies for conditions ranging from neurodegenerative to cardiovascular diseases.
The Rationale for Mitochondrial Surface Engineering: Addressing Dysfunction and Delivery Hurdles
Understanding Mitochondrial Dysfunction in Disease Pathogenesis
Mitochondria are essential organelles that function as cellular power plants, generating adenosine triphosphate (ATP) through oxidative phosphorylation (OXPHOS) [1] [2]. Beyond energy production, they play crucial roles in regulating oxidative stress, calcium homeostasis, immune responses, and apoptosis [1]. Mitochondrial dysfunction arises from defects in the electron transport chain, mutations in mitochondrial DNA (mtDNA), impaired mitochondrial dynamics, and disrupted quality control mechanisms, leading to reduced ATP synthesis, increased reactive oxygen species (ROS) production, and activation of inflammatory pathways [1]. These dysfunctional processes contribute to the pathogenesis of a broad spectrum of diseases, including neurodegenerative disorders, cardiovascular diseases, and oral inflammatory diseases such as periodontitis [3] [2].
The emerging therapeutic strategy of mitochondrial transfer and transplantation (MTT) aims to rescue dysfunctional cells by introducing healthy mitochondria to restore bioenergetics and cellular homeostasis [3]. However, clinical translation faces significant challenges, particularly in delivering functional mitochondria to target cells efficiently. Only a small proportion (approximately 10%) of injected mitochondria typically reach target cells, and the transfer lacks specificity [3]. To overcome these limitations, biotechnological approaches involving surface modification of mitochondria with cell-penetrating peptides (CPPs) have shown promise for enhancing delivery efficiency and specificity [4] [3]. This Application Note details protocols and methodologies for investigating mitochondrial dysfunction and developing CPP-enhanced mitochondrial delivery systems for research and therapeutic development.
Quantitative Analysis of Mitochondrial Dysfunction and CPP Performance
Key Biomarkers of Mitochondrial Dysfunction
Table 1: Quantitative Biomarkers of Mitochondrial Dysfunction in Disease Models
| Biomarker Category | Specific Parameter | Healthy Control Values | Dysfunctional State Values | Measurement Techniques |
|---|---|---|---|---|
| Bioenergetics | ATP Production | Cell/tissue dependent | Up to 5-fold reduction in chronic periodontitis [2] | Luminescence assay, HPLC |
| Oxygen Consumption Rate (OCR) | Cell/tissue dependent | 5-fold reduction in gingival cells of CP patients [2] | Seahorse XF Analyzer | |
| Oxidative Stress | Mitochondrial ROS (mtROS) | Cell/tissue dependent | 18% increase in chronic periodontitis patients [2] | MitoSOX Red, DCFDA assay |
| Lipid Peroxidation | Baseline levels | Significantly increased in periodontitis with kidney injury [2] | TBARS assay, MDA detection | |
| Mitochondrial DNA | mtDNA Copy Number | Varies by cell type | Decreased in periodontitis patients [2] | qPCR, digital PCR |
| mtDNA Mutations | Low mutation load | 14 unique mutations in CP gingival tissue [2] | Sequencing, RFLP | |
| Membrane Integrity | Mitochondrial Membrane Potential (ΔΨm) | High polarization (e.g., JC-1 red/green >5) | 4-fold reduction in CP patients [2] | JC-1, TMRM, TMRE staining |
| Mitochondrial Dynamics | Fusion/Fission Balance | Balanced ratio | Excessive fission in metabolic dysfunction [1] | Western blot (DRP1, MFN2, OPA1) |
CPP Efficiency and Mitochondrial Transfer Quantification
Table 2: Performance Metrics of Mitochondrial Delivery Systems
| Delivery System | Target Cell/Model | Transfer Efficiency | Key Functional Outcomes | References |
|---|---|---|---|---|
| Droplet Microfluidics | C2C12 myoblasts | Controlled transfer (8, 14, or 31 mitochondria/cell) | Cells with 31 mitochondria showed significant functional improvement [5] | [5] |
| Pep-1 Conjugation | Parkinson's disease models | 60.5% of cells received mitochondria vs. 14.5% with naked mitochondria [3] | Anti-apoptotic effects, reduced oxidative stress, improved locomotion [3] | [3] |
| TAT-dextran Coating | Cardiomyocyte reperfusion injury | 182.8% increase vs. free mitochondria [3] | Prevention of oxidative phosphorylation impairment [3] | [3] |
| Free Mitochondria Uptake | Various cell lines (HeLa, A431, SKOV3) | 1-2% of applied mitochondria [6] | Subset escapes endosomes, potential network integration [6] | [6] |
| CPP-Mediated mtDNA Delivery | ARPE-19 epithelial cells | Retention for ≥4 weeks [7] | Mitochondrial RNA and protein production [7] | [7] |
Experimental Protocols for Mitochondrial Dysfunction Analysis and CPP-Mediated Delivery
Protocol 1: Assessment of Mitochondrial Bioenergetics and ROS Production in Cellular Models
Purpose: To quantitatively evaluate mitochondrial dysfunction through bioenergetic profiling and oxidative stress parameters in vitro.
Materials:
- Cell culture system of interest (e.g., HGFs, PDLSCs for oral disease models) [2]
- Seahorse XF Analyzer and XF Cell Culture Microplates
- Mitochondrial Stress Test Kit (Agilent)
- MitoSOX Red mitochondrial superoxide indicator (Invitrogen)
- ATP determination kit (luminescence-based)
- JC-1 Mitochondrial Membrane Potential Assay Kit
- Lysis buffer (e.g., RIPA buffer with protease and phosphatase inhibitors)
Procedure:
- Cell Culture and Preparation: Plate cells at optimal density (e.g., 2×10⁴ cells/well for Seahorse analysis) in appropriate growth medium and culture for 24 hours.
- Mitochondrial Stress Test:
- Hydrate Seahorse sensor cartridge in XF Calibrant at 37°C in non-CO₂ incubator overnight.
- Replace growth medium with Seahorse XF Base Medium supplemented with 1mM pyruvate, 2mM glutamine, and 10mM glucose.
- Incubate cells for 1 hour at 37°C in non-CO₂ incubator.
- Load cartridge with mitochondrial inhibitors: oligomycin (ATP synthase inhibitor, 1μM), FCCP (uncoupler, 1μM), and rotenone/antimycin A (complex I/III inhibitors, 0.5μM).
- Run assay with 3 measurement cycles for each injection.
- ATP Production Measurement:
- Lyse cells in ATP assay-compatible lysis buffer.
- Combine lysate with ATP reaction mix and measure luminescence immediately.
- Normalize ATP values to protein concentration.
- Mitochondrial ROS Detection:
- Load cells with 5μM MitoSOX Red in buffer and incubate for 10 minutes at 37°C.
- Wash cells with warm buffer and measure fluorescence (excitation/emission: 510/580nm).
- Mitochondrial Membrane Potential:
- Incubate cells with JC-1 dye (2μM) for 15-20 minutes at 37°C.
- Analyze red (590nm) and green (529nm) fluorescence; calculate red/green ratio.
Data Analysis: Calculate basal respiration, ATP-linked respiration, proton leak, maximal respiration, and spare respiratory capacity from Seahorse data. Normalize all parameters to cell number or protein content. Compare test conditions to appropriate controls using statistical tests (t-test, ANOVA).
Protocol 2: Surface Modification of Mitochondria with CPPs for Enhanced Cellular Delivery
Purpose: To isolate functional mitochondria and modify their surface with cell-penetrating peptides to improve cellular uptake and targeting efficiency.
Materials:
- Mitochondrial Isolation Kit (e.g., Thermo Scientific)
- Cell-penetrating peptides (e.g., TAT, R8, Pep-1, Penetratin, Transportan)
- Fluorescent mitochondrial dyes (MitoTracker Green FM, MitoTracker Deep Red)
- CellMask plasma membrane stains (optional)
- Ultracentrifuge and rotors
- Droplet microfluidics system (optional, for high-throughput transfer) [5]
- Confocal microscopy imaging system
- Lysis buffer for mitochondrial extraction
Procedure:
- Mitochondrial Isolation:
- Homogenize donor cells (e.g., HEK293, mesenchymal stem cells) in ice-cold mitochondrial isolation buffer using Dounce homogenizer (30-40 strokes).
- Centrifuge homogenate at 800×g for 10 minutes at 4°C to remove nuclei and unbroken cells.
- Transfer supernatant to new tube and centrifuge at 12,000×g for 15 minutes at 4°C.
- Resuspend mitochondrial pellet in isolation buffer.
- Determine mitochondrial protein concentration using BCA assay.
Mitochondrial Labeling (Optional):
- Incubate isolated mitochondria with 100nM MitoTracker Green FM for 15-20 minutes at 37°C.
- Wash excess dye with cold isolation buffer by centrifugation at 12,000×g for 10 minutes.
CPP Modification:
- Prepare CPP solution in conjugation buffer (e.g., 10mM HEPES, pH 7.4).
- Combine mitochondria (0.5-1mg protein) with CPP at varying molar ratios (optimize between 10:1 to 100:1 CPP:mitochondrial protein).
- Incubate for 30-60 minutes at 4°C with gentle agitation.
- Remove unbound CPP by centrifugation at 12,000×g for 10 minutes.
Quality Assessment:
- Verify mitochondrial integrity by measuring membrane potential with JC-1 or TMRM.
- Assess respiratory function using oxygen consumption measurements.
Cellular Delivery:
- Incubate CPP-modified mitochondria with recipient cells at varying concentrations (10-100μg mitochondrial protein/mL media) for 2-24 hours.
- For droplet microfluidics approach: co-encapsulate cells and mitochondria in droplets using flow-focusing device [5].
- Remove uninternalized mitochondria by washing with heparin sulfate solution (to remove surface-bound mitochondria) [8].
Uptake Quantification:
- Analyze mitochondrial uptake by flow cytometry for fluorescently labeled mitochondria.
- Perform confocal microscopy with z-stacking to verify intracellular localization.
- Use luminescence-based assays for quantitative uptake measurements with engineered mitochondrial markers [6].
Data Analysis: Calculate uptake efficiency as percentage of cells with internalized mitochondria. Determine average number of mitochondria per cell through quantitative image analysis. Assess functional integration by measuring rescue of mitochondrial function in recipient cells.
Diagram 1: Pathogenic pathways of mitochondrial dysfunction and CPP-based therapeutic strategy. Mitochondrial damage triggers bioenergetic failure, oxidative stress, and DAMP release, driving inflammation. CPP-modified mitochondria overcome delivery barriers to rescue cellular function.
The Scientist's Toolkit: Essential Research Reagents and Solutions
Table 3: Key Reagents for Mitochondrial Dysfunction and CPP Delivery Research
| Reagent Category | Specific Examples | Research Application | Key Considerations |
|---|---|---|---|
| CPP Sequences | TAT (GRKKRRQRRRPPQ), R8 (octaarginine), Penetratin, Transportan, Pep-1 [8] [3] | Mitochondrial surface modification for enhanced cellular uptake | Uptake depends on both cell type and CPP sequence; Transportan and R8 show higher uptake [8] |
| Mitochondrial Dyes | MitoTracker Green FM, MitoTracker Deep Red, TMRM, JC-1, MitoSOX Red [5] [9] | Tracking mitochondrial localization, membrane potential, and ROS | Avoid dye transfer artifacts; use stable transfection with mitochondrial-targeted fluorescent proteins when possible [6] |
| Fluorescent Tags | TAMRA, FITC, Cy5.5, Cy7, NanoLuciferase [8] [6] [10] | Quantifying uptake and intracellular trafficking | TAMRA fluorescence affected by sequence, buffer conditions, and tryptophan quenching [9] |
| Mitochondrial Isolation Kits | Commercial kits (eilcrochondria Isolation Kit), differential centrifugation protocols | Obtaining functional mitochondria for transfer studies | Isolate quickly and maintain cold conditions; functional lifespan ~2 hours post-isolation [3] |
| Uptake Inhibitors | Heparin sulfate, chlorpromazine, amiloride, wortmannin, temperature block (4°C) [8] [6] | Mechanistic studies of uptake pathways | Temperature block (4°C) virtually eliminates uptake, suggesting energy-dependent process [6] |
| Functional Assays | Seahorse XF Analyzer, ATP luminescence assays, oxygen consumption measurements | Assessing mitochondrial functional recovery | Normalize to protein content or cell number; include appropriate controls for accurate interpretation |
Advanced Technical Considerations
Critical Factors in CPP Selection and Application
The efficiency of CPP-mediated mitochondrial delivery depends on several critical factors. Research indicates that cellular uptake depends on both cell type and CPP sequence, with the ranking of efficiency typically being Transportan > R8 > Penetratin ≈ TAT > Xentry across various cell lines [8]. The mechanism of cellular uptake appears to be primarily through endocytosis, with most CPP-fusion proteins localizing to endosomes after internalization [8]. A significant challenge is that internalized materials must escape the endosomal pathway to reach the cytosol and other intracellular compartments, with evidence suggesting that fewer than 10% of internalized mitochondria successfully escape endosomal entrapment [6].
CPP sequence modification can enhance uptake efficiency. Cyclization of CPP sequences or addition of an HA-sequence has been shown to increase cellular uptake, though this does not necessarily improve endosomal escape [8]. For mitochondrial targeting specifically, incorporating mitochondrial presequences that form helical structures recognized by translocase of the outer mitochondrial membrane (Tom) complexes can significantly improve mitochondrial localization [4].
Methodological Considerations for Accurate Quantification
Accurate quantification of mitochondrial uptake and function requires careful methodological considerations. Fluorescence-based quantification approaches are susceptible to various artifacts, including tryptophan-mediated quenching of fluorophores like TAMRA, particularly for CPPs with high tryptophan content [9]. The physicochemical properties of fluorescent dyes conjugated to CPPs can significantly affect their cellular distribution and membrane interaction, potentially leading to over- or under-estimation of internalization [9].
For optimal quantification of mitochondrial transfer, researchers should employ multiple complementary approaches. Luminescence-based assays using engineered mitochondrial markers like NLuc-HA-OMP25 provide highly sensitive quantification of mitochondrial uptake [6]. Flow cytometry offers rapid screening capabilities but may not be suitable for all cell types, particularly large or polarized cells [9]. Protease protection assays are essential for distinguishing between internalized mitochondria and surface-adherent material [6].
Diagram 2: Experimental workflow for CPP-modified mitochondrial delivery, from isolation to functional analysis in recipient cells.
The strategic application of CPP-modified mitochondria represents a promising frontier in addressing mitochondrial dysfunction across various disease contexts. The protocols and analytical frameworks presented herein provide researchers with comprehensive tools to investigate mitochondrial pathogenesis and develop enhanced delivery systems. Critical to success is the careful selection of CPP sequences based on target cell types, implementation of appropriate quantification methods that account for technical artifacts, and thorough functional validation of mitochondrial recovery. As this field advances, standardization of these methodologies across research groups will accelerate the translation of mitochondrial therapies toward clinical application.
The Therapeutic Promise and Pitfalls of Mitochondrial Transplantation
Mitochondrial transplantation (MT) is an emerging therapeutic strategy aimed at restoring cellular function by introducing healthy exogenous mitochondria into cells with compromised organelle function. This approach has garnered significant attention for treating a wide range of conditions, including cardiovascular diseases, neurodegenerative disorders, and spinal cord injuries [11] [12]. The core premise of MT is to rescue dysfunctional cells by augmenting mitochondrial quantity and enhancing overall cellular performance, particularly in tissues with high energy demands [11].
The therapeutic potential of MT is primarily attributed to the central role mitochondria play in cellular homeostasis. As essential organelles, mitochondria are responsible for adenosine triphosphate (ATP) generation through oxidative phosphorylation, regulation of apoptosis, calcium ion storage, and maintenance of redox balance [13]. Mitochondrial dysfunction disrupts these critical processes, leading to impaired protein synthesis, disrupted biogenesis, and faulty apoptosis regulation [11]. By introducing functional mitochondria, MT seeks to restore bioenergetic capacity and mitigate the adverse effects of mitochondrial impairment.
Despite promising preclinical results, the clinical translation of MT faces several challenges, including low transfer efficiency, limited stability of isolated mitochondria, and imperfect cellular integration [11]. Recent advances in biotechnology, particularly surface modification of mitochondria with cell-penetrating peptides (CPPs), have shown potential to overcome these limitations by enhancing targeting precision, improving cellular uptake, and increasing therapeutic efficacy [11] [14]. This application note explores both the promise and pitfalls of mitochondrial transplantation, with specific focus on CPP-enhanced delivery systems.
Therapeutic Applications and Supporting Evidence
Mitochondrial transplantation has demonstrated therapeutic potential across diverse disease models. The table below summarizes key applications, observed outcomes, and supporting evidence from recent studies.
Table 1: Therapeutic Applications of Mitochondrial Transplantation
| Disease Area | Experimental Model | Therapeutic Outcomes | Key Evidence |
|---|---|---|---|
| Spinal Cord Injury (SCI) | In vivo animal models | Enhanced neuronal energy metabolism; Reduced oxidative stress; Promoted axonal regeneration and functional recovery [13]. | Transplanted healthy mitochondria into neurons improved energy metabolism and reduced oxidative stress, promoting survival and functional recovery [13]. |
| Ischemic Heart Disease (IHD) | In vivo models; Few clinical trials in humans [12]. | Restoration of ATP production; Improved myocardial function; Reduction of infarct size; Induction of angiogenesis [12]. | Injected mitochondria fostered healing via ATP restoration, Ca2+ regulation, ROS homeostasis, and activation of survival mechanisms in cardiomyocytes [12]. |
| Ocular Disorders | In vitro and in vivo models for corneal, optic nerve, and retinal disorders [15]. | Bioenergetic rescue of metabolically dysfunctional cells; Alleviation of oxidative stress [15]. | Application of MT to ocular metabolic disorders showed promising therapeutic outcomes, though it remains an emerging therapy in ophthalmology [15]. |
| Aging-Related Diseases | Preclinical models of cardiovascular and neurodegenerative disorders [14]. | Restoration of bioenergetics; Alleviation of oxidative stress; Improved organ function [14]. | MT was shown to revive ATP production and alleviate oxidative stress in models of ischemia and neurodegeneration [14]. |
The mechanisms through which transplanted mitochondria exert their beneficial effects are multifaceted. Beyond simply replacing defective organelles, they can enhance metabolic regulation, support cellular survival under stress conditions, and promote tissue regeneration [11]. In the context of ischemic heart disease, transplanted mitochondria have been shown to induce angiogenesis, facilitating the formation of new blood vessels and improving blood supply to damaged tissues [12]. Furthermore, emerging evidence suggests that mitochondrial transfer can stimulate the proliferation and differentiation of local tissue stem cells, contributing to regenerative processes [12].
Pitfalls and Technical Challenges
Despite its considerable promise, the clinical translation of mitochondrial transplantation faces significant technical and biological challenges that must be addressed for therapeutic application.
Table 2: Key Challenges in Mitochondrial Transplantation
| Challenge Category | Specific Limitations | Impact on Therapy |
|---|---|---|
| Efficiency & Stability | Short lifespan of isolated mitochondria (significant loss of respiratory function after ~2 hours) [11]. | Limits therapeutic window and requires rapid isolation-to-delivery protocols. |
| Low transfer efficiency (only ~10% of injected mitochondria reach target cells) [11]. | Reduces therapeutic efficacy and necessitates high initial mitochondrial doses. | |
| Cellular Integration | Mitochondria must avoid lysosomal degradation and integrate into existing network [11]. | Inefficient integration limits functional benefits; mechanisms of fusion remain unclear. |
| Uncertain survival period and functional duration of transplanted mitochondria [11]. | Makes dosing regimens and treatment frequency difficult to establish. | |
| Immunological Concerns | Mitochondria recognized as foreign entities, potentially triggering immune responses [11]. | May compromise therapeutic efficacy and cause adverse inflammatory reactions. |
| Non-viable or damaged mitochondria can release damage-associated molecular patterns (DAMPs) [11]. | Potentially exacerbates inflammation and tissue damage. | |
| Delivery & Scalability | Lack of specificity for target cells; nonspecific distribution with systemic injection [11]. | Limits precision and raises potential off-target effects. |
| Transition from small-scale experiments to widespread clinical use requires standardized, cost-effective protocols [11]. | Currently hinders clinical translation and commercial viability. |
The extracellular environment presents additional hurdles. Isolated mitochondria face an inhospitable milieu characterized by high calcium concentrations and, in pathological conditions, elevated reactive oxygen species (ROS) [11]. Furthermore, injected mitochondria must maintain stability in the extracellular environment, avoiding aggregation, swelling, and structural changes that could compromise function or trigger immune responses [11].
Surface Modification with Cell-Penetrating Peptides (CPPs)
Surface modification of mitochondria with cell-penetrating peptides represents a promising biotechnological approach to overcome the limitations of conventional mitochondrial transplantation. CPPs are typically short, positively charged peptides that facilitate interactions with negatively charged cell membranes, enabling efficient cellular entry and subsequent cargo delivery [11].
CPP Mechanisms and Applications
The application of CPPs to mitochondrial medicine has shown substantial promise in enhancing delivery efficiency. Two well-characterized CPPs, HIV-1 TAT protein and Pep-1, have demonstrated efficacy in facilitating mitochondrial uptake and functional delivery:
- TAT-mediated delivery: The HIV-1 TAT protein has been shown to mediate the import of functionally active mitochondrial enzymes via covalent coupling, promoting the restoration of essential cellular functions [11].
- Pep-1-mediated delivery: Pep-1 can translocate cargo through non-covalent self-assembly. Once inside the cell, cargo is released via an endocytosis-independent mechanism [11].
Research by Chang et al. investigated mitochondrial conjugation with the peptide transporter Pep-1 (Pep-1 mediated mitochondria delivery, PMD) and compared its efficiency to cell-free mitochondria delivery. The Pep-1/mitochondria complex was prepared at a weight ratio of 1750:1 by incubation at 37°C for 30 minutes [11]. This modified system was tested in various disease models, including neurotoxin-induced PC12 cells, Parkinson's disease rat models, and a cybrid cell model of mitochondrial myopathy, encephalopathy, lactic acidosis, and stroke-like episodes (MELAS) [11].
Experimental Protocol: Pep-1-Mediated Mitochondrial Delivery
Objective: To efficiently deliver functional mitochondria to target cells using Pep-1 conjugation.
Materials:
- Isolated functional mitochondria (from autologous or donor tissues)
- Pep-1 peptide (commercially available)
- Isolation buffer (typically containing sucrose, EGTA, and HEPES)
- Cell culture medium
- Target cells (e.g., neurons, cardiomyocytes)
Procedure:
- Mitochondrial Isolation: Isolate mitochondria from donor tissue using differential centrifugation. Maintain mitochondria on ice throughout the process to preserve function [11].
- Complex Formation:
- Prepare Pep-1 solution in appropriate buffer.
- Combine Pep-1 with isolated mitochondria at a weight ratio of 1750:1 (Pep-1:mitochondria) [11].
- Incubate the mixture at 37°C for 30 minutes to form stable complexes.
- Quality Assessment:
- Verify mitochondrial membrane potential using fluorescent dyes (e.g., JC-1 or TMRE).
- Assess structural integrity via electron microscopy if available.
- Cellular Delivery:
- Add Pep-1/mitochondria complexes to target cells in culture.
- Incubate for desired duration (typically 2-24 hours).
- Uptake Validation:
- Label mitochondria with fluorescent markers (e.g., MitoTracker) before conjugation to track uptake.
- Confirm intracellular localization using confocal microscopy.
- Assess functional integration through measurements of ATP production, oxygen consumption rate, or restoration of metabolic activity.
Technical Notes:
- The isolation method must minimize loss of function/vitality and structural integrity [11].
- Use viable and functional mitochondria to prevent DAMP release and immune activation [11].
- Optimal incubation times may vary by cell type and should be determined empirically.
Diagram 1: This workflow illustrates the key steps for modifying mitochondria with cell-penetrating peptides (CPPs) like Pep-1 to enhance cellular delivery, covering isolation, complex formation, quality control, and functional validation.
Nanoengineered Mitochondria and Advanced Delivery Systems
Beyond CPP conjugation, more sophisticated nanoengineering approaches are emerging to enhance mitochondrial delivery. These biohybrid systems integrate isolated mitochondria with synthetic nanomaterials or biomolecules to confer new functionalities and overcome the limitations of conventional MT [14].
Table 3: Nanoengineering Strategies for Enhanced Mitochondrial Delivery
| Strategy | Design Approach | Mechanism of Action | Applications |
|---|---|---|---|
| Liposome Encapsulation | Coating mitochondria with DOTAP/DOPE lipid bilayers [14]. | Cationic lipid coating enhances surface charge and stability, improving uptake and lowering immunogenicity [14]. | Cerebral ischemia models; acute respiratory distress syndrome (ARDS) [14]. |
| Chemical Conjugation | Conjugation of targeting peptides (e.g., TPP, CAQK) to mitochondrial surface [13] [14]. | Ligand-receptor recognition enables targeted delivery to specific tissues; TPP+ leverages mitochondrial membrane potential [14]. | Ischemia-reperfusion injury; spinal cord injury [13] [14]. |
| Polymer Coating | Functionalization with hydrophilic, biocompatible polymers [11]. | Provides protective microenvironment; shields from enzymatic degradation and immune detection [11]. | Under investigation for various applications. |
| Extracellular Vesicle Packaging | Encapsulating mitochondria in natural or engineered vesicles [11]. | Utilizes natural trafficking mechanisms; enhances biocompatibility and targeting [11]. | Cardiovascular and neurodegenerative diseases [11]. |
These nanoengineered mitochondria represent a significant advancement toward harnessing the full therapeutic potential of MT. They provide a protective microenvironment for mitochondria during transit, shielding them from enzymatic degradation, immune detection, and oxidative damage [11]. Furthermore, these delivery vectors can be engineered with targeting ligands, enabling enhanced interaction with specific cell types and allowing for controlled release of mitochondrial cargo [11].
The functionalization of mitochondrial surfaces with hydrophilic, biocompatible polymers has shown particular promise in improving stability and circulation time. Additionally, the use of stimulus-responsive systems (e.g., pH/ROS-sensitive polymers) can guide mitochondria to inflammatory sites, while externally driven platforms (e.g., magnetically steered nanocapsules) offer precise spatial control over delivery [14].
The Scientist's Toolkit: Essential Research Reagents
Successful implementation of mitochondrial transplantation and surface modification protocols requires specific reagents and materials. The following table details key components for research in this field.
Table 4: Essential Research Reagents for Mitochondrial Transplantation Studies
| Reagent/Material | Function/Application | Examples/Specifications |
|---|---|---|
| Cell-Penetrating Peptides (CPPs) | Enhance mitochondrial uptake and precision delivery [11]. | Pep-1, HIV-1 TAT protein [11]. |
| Mitochondrial Isolation Kits | Obtain functional mitochondrial fractions from tissues or cells. | Sucrose-based buffers, differential centrifugation protocols [11]. |
| Membrane Potential Dyes | Assess mitochondrial viability and function pre- and post-transplantation. | JC-1, TMRE, MitoTracker probes [11]. |
| Targeting Ligands | Direct modified mitochondria to specific tissues or cell types. | TPP+ (triphenylphosphonium cation), CAQK peptide [13] [14]. |
| Lipid Formulations | Create protective vesicles or coatings for mitochondria. | DOTAP/DOPE liposomes [14]. |
| Fluorescent Tags | Track mitochondrial uptake, distribution, and persistence. | MitoTracker Green/Red, GFP-labeled mitochondria [11]. |
Diagram 2: This diagram outlines the primary therapeutic mechanisms by which transplanted mitochondria exert their beneficial effects, including bioenergetic rescue, signaling modulation, and mechanical support.
Mitochondrial transplantation represents a promising therapeutic modality for a spectrum of diseases characterized by mitochondrial dysfunction. While challenges in efficiency, stability, and delivery remain significant, emerging technologies—particularly surface modification with CPPs and nanoengineering approaches—offer viable paths toward clinical translation. The continued refinement of mitochondrial delivery protocols, coupled with rigorous assessment of long-term safety and efficacy, will be essential for realizing the full potential of this innovative therapy. As the field advances, mitochondrial transplantation may ultimately establish itself as a mainstream therapeutic strategy for restoring cellular homeostasis in diverse pathological contexts.
Cell-penetrating peptides (CPPs) are short peptides, typically consisting of 5–40 amino acids, characterized by their unique ability to cross cellular membranes. Since the initial discovery of the trans-activating transcriptional activator (TAT) protein from HIV-1 in 1988, over 1,850 distinct CPPs have been cataloged, offering a versatile toolkit for intracellular delivery [16] [17]. These peptides facilitate the transport of diverse therapeutic and diagnostic cargoes—including small molecules, nucleic acids, proteins, and nanoparticles—into cells, overcoming the fundamental biological barrier of the plasma membrane [18] [19]. Their efficiency in delivering cargo across formidable barriers, including the blood-brain barrier, positions CPPs as powerful vectors for addressing previously inaccessible intracellular targets [17].
In the specific context of mitochondrial research, CPPs offer a promising strategy to overcome the central challenge of mitochondrial membrane permeabilization. Mitochondria possess their own genome (mtDNA), and mutations in this DNA are associated with a wide spectrum of metabolic, neurodegenerative, and endocrine diseases, as well as cancer [20]. Traditional therapeutic approaches often fail to effectively deliver macromolecules to the mitochondrial matrix. The functionalization of delivery systems with CPPs, especially those conjugated with mitochondrial targeting signals (MTS), presents a sophisticated approach to transfect mammalian mitochondria directly. This strategy aims to restore normal mitochondrial function by addressing genetic defects at their source, making CPPs a cornerstone of emerging mitochondrial gene therapy and surface modification strategies [20] [4].
Classification and Selection of CPPs
Types of Cell-Penetrating Peptides
CPPs are categorized based on their physicochemical properties, such as charge and origin. The selection of an appropriate CPP is critical and depends on the cargo's properties, the target cell type, and the desired intracellular destination [21].
Table 1: Major Classes of Cell-Penetrating Peptides
| Classification | Characteristics | Key Examples (Sequence) | Primary Applications & Notes |
|---|---|---|---|
| Cationic CPPs | Net positive charge at physiological pH; rich in arginine (R) and lysine (K) residues [16] [17]. | TAT (GRKKRRQRRRPPQ) [16], Penetratin (RQIKIWFQNRRMKWKK) [16], Oligoarginines (e.g., R8) [22]. | General intracellular delivery; high transduction efficiency; the guanidinium group in arginine is crucial for membrane interaction [17]. |
| Amphipathic CPPs | Contain distinct hydrophobic and hydrophilic regions [16]. | Transportan (GWTLNSAGYLLGKINLKALAALAKKIL) [16], MAP (KLALKLALKALKAALKLA) [16], MPG (GALFLGFLGAAGSTMGAWSQPKKKRKV) [22]. | Effective for delivering hydrophobic cargos and large biomolecules; most common class of CPPs [21] [16]. |
| Anionic CPPs | Carry a net negative charge; less common [16]. | p28 (LSTAADMQGWTDGMASGLDKDYLKPDD) [16]. | Exhibit cancer-preferential entry; mechanism of internalization differs from cationic CPPs [16]. |
Guidelines for CPP Selection
Choosing the optimal CPP requires a balanced consideration of multiple factors [21]:
- Cargo Properties: Larger cargos may require CPPs with high transport capacity like Transportan or Pep-1. Negatively charged cargoes (e.g., nucleic acids) are most effectively delivered by cationic CPPs like TAT or oligoarginines. Hydrophobic molecules benefit from CPPs with increased hydrophobic character [21].
- Target Cell Type: Membrane composition and dominant endocytic pathways vary by cell type, affecting CPP uptake efficiency. For instance, HEK293 cells can be more challenging for CPP delivery than HeLa cells, potentially requiring modified CPPs or endosomal escape enhancers [21].
- Intracellular Localization: If mitochondrial targeting is desired, CPPs must be modified. This is typically achieved by conjugating the CPP with a Mitochondrial Targeting Signal (MTS), such as the presequence of mitochondrion-localized proteins, or by using peptides with inherent mitochondrial affinity like (KLAKLAK)₂ [20] [21] [4].
Application Note: CPPs for Mitochondrial Delivery
Workflow and Signaling Pathways
The following diagram illustrates the conceptual workflow for utilizing CPPs to deliver cargo to mitochondria, highlighting key biological pathways involved in mitochondrial targeting and internalization.
Protocol: Functionalizing a Cargo with a CPP for Mitochondrial Delivery
This protocol details the process of creating a CPP-MTS conjugate for delivering a protein cargo (e.g., a nuclease for genome editing) to mitochondria in mammalian cells.
Principle: A cationic or amphipathic CPP is covalently linked to both a Mitochondrial Targeting Signal (MTS) and the therapeutic cargo. The CPP facilitates cellular uptake, while the MTS directs the complex to the mitochondria via the TOM complex machinery [20] [4].
Materials:
- CPP: Synthetic TAT peptide (GRKKRRQRRRPPQ) with an N-terminal cysteine residue.
- MTS: Synthetic peptide representing the presequence of a mitochondrial protein (e.g., Cox IV).
- Cargo: Recombinant protein with a C-terminal AviTag for site-specific biotinylation.
- Crosslinker: Maleimide-PEG₂-Biotin.
- Coupling Agent: Streptavidin.
- Cell Line: Human HeLa cells.
- Buffers: Phosphate-Buffered Saline (PBS), HEPES-Buffered Saline (HBS).
Procedure:
- Activation of the MTS Peptide:
- Dissolve the MTS peptide in degassed PBS to a final concentration of 1 mM.
- Add a 5-fold molar excess of Maleimide-PEG₂-Biotin crosslinker to the MTS solution.
- Incubate for 2 hours at room temperature with gentle agitation.
- Purify the biotinylated MTS peptide using size-exclusion chromatography or dialysis.
Conjugation of CPP-MTS-Cargo:
- Biotinylate the cargo protein at its AviTag using BirA enzyme per standard protocols.
- Combine the biotinylated MTS and biotinylated cargo at a 1:1 molar ratio.
- Add a 2-fold molar excess of Streptavidin to the mixture. Incubate for 1 hour on ice to form the MTS-Cargo complex via streptavidin-biotin bridging.
- Add a 5-fold molar excess of the cysteine-containing CPP to the complex. The maleimide group on the MTS-biotin-crosslinker will react with the sulfhydryl group on the CPP's cysteine. Incubate for 4 hours at 4°C.
Purification and Validation:
- Purify the final CPP-MTS-Cargo conjugate using fast protein liquid chromatography (FPLC).
- Validate the conjugation and purity via SDS-PAGE and Western blotting using antibodies against the CPP, the cargo, and streptavidin.
Cellular Transfection and Analysis:
- Seed HeLa cells in a 24-well plate at a density of 5 x 10⁴ cells/well and culture overnight.
- Replace the medium with serum-free medium containing the CPP-MTS-Cargo conjugate (1-10 µM final concentration).
- Incubate for 4 hours at 37°C.
- Replace the transfection medium with complete growth medium and incubate for an additional 20 hours.
- Analyze mitochondrial localization using confocal microscopy by co-staining with MitoTracker Deep Red. Assess mitochondrial function via ATP assays and superoxide detection kits [23].
The Scientist's Toolkit: Key Reagents and Materials
Table 2: Essential Research Reagents for CPP-Based Mitochondrial Delivery
| Reagent/Material | Function/Description | Example Use Case |
|---|---|---|
| Cationic CPPs (e.g., TAT, R8) | Facilitate initial cellular uptake via electrostatic interactions with the negatively charged cell membrane [16] [17]. | General-purpose delivery of various cargo types into a wide range of cells. |
| Amphipathic CPPs (e.g., Transportan, MAP) | Balance hydrophobic and hydrophilic domains for efficient membrane interaction and cargo delivery [16]. | Particularly effective for delivering large or hydrophobic cargo molecules. |
| Mitochondrial Targeting Signal (MTS) | Peptide sequence that directs the conjugated complex to the mitochondria, recognized by the TOM complex [4]. | Essential component for rerouting CPP-cargo complexes from the cytosol to the mitochondrial matrix. |
| Endosomal Escape Enhancers | Agents (e.g., chloroquine, fusogenic peptides) that disrupt endosomes, preventing lysosomal degradation of the cargo [21] [19]. | Added to transfection media to significantly improve the cytosolic release and efficacy of endocytosed CPP-cargo complexes. |
| Click Chemistry Kits | Enable efficient, specific, and bioorthogonal covalent conjugation between CPPs and cargo molecules [18] [21]. | Simplifying the reproducible creation of stable CPP-cargo constructs, especially for sensitive biomolecules. |
| Cationized Gelatin Nanospheres (cGNS) | Nanocarriers that can be electrostatically associated with cargo to enhance cellular internalization efficiency [23]. | Demonstrated to improve mitochondrial internalization and subsequent functional enhancement (e.g., increased ATP production) [23]. |
Quantitative Data and Market Outlook
The efficacy and growing adoption of CPP-based strategies are reflected in both experimental data and market analysis.
Table 3: Quantitative Data on CPP Performance and Market
| Parameter | Quantitative Finding | Context & Source |
|---|---|---|
| Cellular Uptake Enhancement | ~10-fold increase in small molecule transport with specific CPPs (e.g., CL peptide) [18]. | Demonstrates the significant improvement in delivery efficiency achievable with CPPs. |
| Optimal Arginine Length | 8-10 residues (R8 to R10) [17]. | Defines the chain length for optimal membrane penetration efficiency of oligoarginine CPPs. |
| Global CPP Market Size (2024) | USD 1.94 Billion [24]. | Indicates the substantial and established commercial interest in CPP technology. |
| Projected Market CAGR (2025-2035) | 10.9% [24]. | Signals strong expected growth and continued investment in CPP research and applications. |
| Projected Market Size (2035) | USD 6.05 Billion [24]. | Highlights the anticipated long-term expansion and commercial significance of the CPP field. |
Mitochondrial transplantation has emerged as a promising therapeutic strategy for a spectrum of diseases involving mitochondrial dysfunction, including neurodegenerative, cardiovascular, and metabolic disorders [25] [3]. The fundamental premise is to restore cellular bioenergetics and homeostasis by introducing healthy, functional mitochondria into compromised cells. However, the transition of this approach from preclinical models to clinical application faces three significant biological barriers: rapid immune clearance, low cellular uptake efficiency, and intracellular lysosomal degradation of delivered mitochondria [3]. These hurdles drastically reduce the therapeutic potential and efficacy of mitochondrial therapies. Surface modification of mitochondria with Cell-Penetrating Peptides (CPPs) presents a sophisticated strategy to overcome these challenges. CPPs, short peptides capable of traversing cellular membranes, can be engineered onto the mitochondrial surface to enhance delivery [18] [26] [3]. This application note details the key barriers and provides structured protocols for leveraging CPP-modified mitochondria, supported by quantitative data and actionable methodologies for researchers and drug development professionals.
Analyzing the Key Barriers to Mitochondrial Delivery
Immune Clearance
Upon systemic administration, isolated mitochondria are recognized by the host immune system as foreign entities, triggering rapid clearance. This occurs because mitochondrial components, including mitochondrial DNA (mtDNA), share structural similarities with bacterial pathogens due to their prokaryotic origin [25] [27]. Specifically, mtDNA contains hypomethylated CpG motifs that are recognized by Toll-like receptor 9 (TLR9) and other pattern recognition receptors,potentially initiating an inflammatory response [28] [27]. This immunogenic recognition leads to opsonization and phagocytosis, significantly reducing the circulation time and bioavailability of the transplanted mitochondria [3].
Low Uptake Efficiency
A critical bottleneck is the inherently low efficiency of cellular internalization of naked mitochondria. Studies indicate that only a small fraction (approximately 10-15%) of injected mitochondria successfully enter target cells under standard conditions [3]. This inefficiency stems from the negative surface charge of both the mitochondrial outer membrane and the target cell plasma membrane, creating electrostatic repulsion that hinders close contact and subsequent uptake [3]. Furthermore, the process lacks specificity, as mitochondria do not inherently possess targeting motifs to direct them to particular cell types [3].
Lysosomal Degradation
Even upon successful cellular entry, a significant portion of internalized mitochondria are trafficked to the endolysosomal compartment. The acidic and enzymatic environment of lysosomes leads to the degradation of mitochondria, preventing their functional integration into the endogenous mitochondrial network [3]. The inability to escape the endolysosomal pathway nullifies the potential bioenergetic benefits of the transplantation.
Table 1: Key Barriers and Their Impact on Mitochondrial Delivery
| Barrier | Underlying Cause | Consequence |
|---|---|---|
| Immune Clearance | Recognition of mtDNA/proteins as foreign via TLR9 and other receptors [28] [27] | Rapid clearance from circulation, potential inflammatory response |
| Low Uptake Efficiency | Electrostatic repulsion; lack of targeting motifs [3] | <15% of administered mitochondria enter cells [3] |
| Lysosomal Degradation | Entrapment in the endocytic pathway [3] [29] | Functional degradation before cytosolic integration |
CPP-Based Strategies to Overcome Delivery Barriers
Cell-Penetrating Peptides (CPPs) are short (5-30 amino acids), cationic, or amphipathic peptides that facilitate the cellular uptake of various cargoes [18] [26]. Their application in mitochondrial transplantation primarily involves covalent or non-covalent conjugation to the mitochondrial surface to enhance delivery.
Enhancing Uptake and Avoiding Lysosomes: CPPs like TAT (YGRKKRRQRRR) and Pep-1 exploit various mechanisms, including direct translocation and endocytosis, to cross cell membranes [26] [3]. When coated on mitochondria, they shield the negative surface charge, reducing electrostatic repulsion and promoting membrane interaction. Some CPPs and associated strategies also facilitate endosomal escape, a crucial step for avoiding lysosomal degradation [29]. The cationic nature of many CPPs can disrupt the endosomal membrane, releasing the cargo into the cytosol.
Combining with Targeting and Shielding: To address immune clearance, CPPs can be used in conjunction with stealth coatings. For instance, conjugating CPPs with biocompatible polymers like dextran can mask immunogenic surface markers on mitochondria, reducing immune recognition while retaining cell-penetrating capability [3].
Table 2: CPPs and Functional Moieties for Enhanced Mitochondrial Delivery
| CPP/Moiety | Sequence/Description | Primary Function in Delivery | Reported Outcome |
|---|---|---|---|
| TAT [26] | YGRKKRRQRRR | Enhances cellular uptake | 182.8% increase in transfer over free mitochondria [3] |
| Pep-1 [26] | KETWWETWWTEWSQPKKKRKV | Enhances cellular uptake | 60.5% of cells received mitochondria vs. 14.5% with naked mitochondria [3] |
| Dextran Polymer [3] | Polysaccharide coating | Shields from immune clearance, reduces aggregation | Improves stability in circulation |
| Triphenylphosphonium (TPP) [4] | Mitochondria-targeting cation | Drives accumulation inside mitochondria | Used in combination with peptides for targeted delivery [3] |
Experimental Protocols for Mitochondrial Surface Modification and Analysis
Protocol 4.1: Mitochondrial Isolation and Surface Modification with CPP-Dextran Conjugates
This protocol outlines a method for isolating functional mitochondria and coating them with a TAT-dextran conjugate to enhance uptake and reduce immune clearance [3].
Research Reagent Solutions:
- Isolation Buffer: 250 mM sucrose, 10 mM HEPES, 1 mM EGTA, pH 7.4.
- Conjugation Buffer: 0.22 M sucrose, 10 mM succinate, 5 mM glutamate, 2 mM K2HPO4, 10 mM HEPES, 1 mM EGTA, pH 7.4.
- TAT-Dextran Conjugate: Synthesized by conjugating the TAT peptide (YGRKKRRQRRR) to an amine-reactive dextran polymer (e.g., 40-70 kDa) via a stable amine bond.
Procedure:
- Mitochondrial Isolation: Fresh tissue (e.g., liver) or cultured cells are homogenized on ice in chilled Isolation Buffer using a Potter-Elvehjem homogenizer. The homogenate is centrifuged at 800 × g for 10 min at 4°C to remove nuclei and debris. The supernatant is then centrifuged at 10,000 × g for 15 min at 4°C to pellet the mitochondrial fraction. Wash the pellet twice and resuspend in a small volume of Conjugation Buffer.
- Protein Quantification: Determine the mitochondrial protein concentration using a Bradford or BCA assay. Adjust the concentration to 5-10 mg/mL with Conjugation Buffer.
- Surface Modification: Incubate the mitochondrial suspension with the TAT-Dextran conjugate at a final concentration of 100-200 µg conjugate per mg of mitochondrial protein for 30-60 minutes at 4°C with gentle agitation.
- Purification: Remove unbound conjugate by centrifuging the mixture at 12,000 × g for 10 min at 4°C. Gently resuspend the pellet (modified mitochondria) in fresh Isolation Buffer.
- Quality Control: Assess mitochondrial membrane potential (ΔΨm) using a fluorescent dye like JC-1 or TMRE to ensure functional integrity post-modification. Purity can be checked by Western blot for mitochondrial markers (e.g., COX IV) and absence of markers for other organelles.
Protocol 4.2: Quantifying Cellular Uptake Efficiency
This protocol describes a flow cytometry-based method to quantify the uptake of fluorescently labeled mitochondria into recipient cells [3].
Research Reagent Solutions:
- Labeling Dye: MitoTracker Deep Red (or other lipophilic cationic dyes that accumulate in active mitochondria).
- Cell Culture Medium: Appropriate medium for the recipient cells (e.g., DMEM for HEK293).
- Fixative: 4% paraformaldehyde (PFA) in PBS.
- Flow Cytometry Buffer: PBS containing 1% BSA.
Procedure:
- Mitochondrial Labeling: Prior to surface modification, label isolated mitochondria with MitoTracker Deep Red (e.g., 100 nM) for 20-30 min at 37°C. Remove excess dye by centrifugation and washing as in Protocol 4.1.
- Co-culture: Seed recipient cells in a 12-well plate and culture until ~70% confluency. Replace the medium with fresh medium containing either labeled naked mitochondria or CPP-modified mitochondria (e.g., 50 µg mitochondrial protein per well).
- Incubation: Co-culture for 4-6 hours at 37°C in a 5% CO2 incubator.
- Cell Harvest and Analysis:
- Gently wash the cells twice with PBS to remove non-internalized mitochondria.
- Trypsinize the cells, quench with complete medium, and collect by centrifugation.
- Wash the cell pellet with Flow Cytometry Buffer and resuspend in a fixed volume of buffer containing 1% PFA.
- Analyze samples using a flow cytometer. Excite MitoTracker Deep Red at 640 nm and detect emission at ~665 nm. The percentage of fluorescent-positive cells and the mean fluorescence intensity (MFI) are indicators of uptake efficiency and quantity, respectively.
Diagram 1: Workflow for CPP-modified mitochondrial delivery, showing the critical steps from isolation to functional integration, with the key branching point at endosomal escape determining the outcome.
The following table consolidates key quantitative findings from recent studies utilizing CPPs and other surface modifications to enhance mitochondrial delivery.
Table 3: Efficacy of Different Mitochondrial Delivery Strategies
| Delivery Strategy | Pathology / Cell Model | Transfer Efficiency / Uptake Metric | Key Biological Effects |
|---|---|---|---|
| Pep-1-conjugated Mito [3] | Parkinson's disease models (in vitro & in vivo) | 60.5% of cells received mitochondria vs. 14.5% with naked mitochondria | Anti-apoptotic effect, reduced oxidative stress, improved locomotion |
| TAT-dextran-coated Mito [3] | Reperfusion injury in cardiomyocytes | 182.8% increase in transfer over free mitochondria | Prevented oxidative phosphorylation impairment, anti-apoptotic |
| PEP-TPP complex [3] | Myocardial ischemia-reperfusion injury | Intravenous naked mitochondria were ineffective; PEP-TPP mitochondria were therapeutic | Reduced apoptosis, inflammation, and infarct size |
| Dextran-TPP coated Mito [3] | Breast and heart cancer cells | ~3-fold increase in internalization vs. uncoated mitochondria | Reduction of reactive oxygen species (ROS) |
The Scientist's Toolkit: Essential Research Reagents
Table 4: Key Reagents for Mitochondrial Delivery Research
| Reagent / Material | Function / Application | Example / Notes |
|---|---|---|
| MitoTracker Deep Red | Fluorescent labeling of live mitochondria for uptake tracking | Preferred for superior photostability; use with flow cytometry or confocal microscopy. |
| JC-1 Dye | Assessment of mitochondrial membrane potential (ΔΨm) | Quality control post-isolation and modification. Ratio of red/green fluorescence indicates health. |
| TAT Peptide | Canonical CPP for enhancing cellular uptake | Sequence: YGRKKRRQRRR; can be chemically synthesized and conjugated to dextran or other carriers. |
| Dextran, Amine-Reactive | Polymer backbone for creating stealth coatings | Conjugate with CPPs; 40-70 kDa is common. Provides shielding without significantly impairing uptake. |
| LLOMe | Inducer of endosomal escape; research tool [29] | Used experimentally to promote release of endosomally trapped cargo, including CPP-conjugates. |
| Antibody Cocktail (for Flow) | Confirm mitochondrial purity post-isolation | Targets: COX IV (mito marker), Calnexin (ER negative), LAMP1 (lysosome negative). |
Diagram 2: Logical relationship between key barriers and CPP-based solution strategies, leading to the desired therapeutic outcome.
Engineering Strategies and Coating Technologies for Mitochondrial Surface Modification
Cell-penetrating peptides (CPPs) represent a promising technological advancement for overcoming the fundamental challenge of cellular membrane impermeability in drug delivery. These short peptides, typically consisting of 5-30 amino acids, possess the unique ability to traverse plasma membranes and facilitate the intracellular delivery of various macromolecular cargoes, including proteins, nucleic acids, and other therapeutic agents [17] [26]. Their application is particularly valuable for mitochondrial research, where delivering compounds across both the plasma membrane and the double mitochondrial membrane presents a significant obstacle. The discovery of CPPs originated with the HIV-1 TAT protein in 1988, followed by the identification of the Drosophila Antennapedia (ANTP) peptide, which paved the way for the development of numerous natural and synthetic peptides with enhanced transduction capabilities [17] [30]. For mitochondrial surface modification and delivery strategies, understanding the structural and functional classifications of CPPs is paramount for selecting appropriate sequences that can not only penetrate cells but also specifically localize to mitochondria.
Classification and Characteristics of CPPs
CPPs are categorized based on their physicochemical properties, origins, and translocation mechanisms. The primary classification system groups CPPs into three major categories: cationic, amphipathic, and hydrophobic peptides, each with distinct structural features and interaction profiles with biological membranes [17] [26].
Table 1: Fundamental Classification of Cell-Penetrating Peptides
| Category | Key Characteristics | Representative Sequences | Primary Translocation Mechanisms |
|---|---|---|---|
| Cationic | Rich in basic residues (Arginine, Lysine); overall positive charge [17] [26]. | TAT (YGRKKRRQRRR) [26], Penetratin (RQIKIWFQNRRMKWKK) [26], Polyarginine (e.g., R9) [17] [26]. | Macropinocytosis, clathrin-mediated endocytosis, caveolae-dependent endocytosis [26]. |
| Amphipathic | Contain segregated hydrophobic and hydrophilic regions; can be primary or secondary [26]. | MPG (GALFLGFLGAAGSTMGAWSQPKKKRKV) [26], Transportan (GWTLNSAGYLLG-K-INLKALAALAKKIL) [26]. | Direct penetration, endocytosis, inverted micelle formation [26]. |
| Hydrophobic | Dominated by non-polar amino acids; low net charge [31]. | PFV (PFVYLI) [31], C105Y (CSIPPEVKFNKPFVYLI) [26]. | Membrane integration and translocation via lipid interactions [31]. |
The translocation efficiency of CPPs is influenced by multiple factors. For cationic CPPs, the number and spatial arrangement of arginine residues are critical, with oligoarginines containing 8-10 residues (R8-R10) generally demonstrating optimal membrane perforation activity [17]. Amphipathic CPPs often rely on their ability to form secondary structures, such as alpha-helices or beta-sheets, upon interacting with phospholipid membranes, which facilitates their uptake [26]. It is important to note that many CPPs utilize multiple, often simultaneous, entry pathways, and their mechanism can be influenced by factors such as cell type, cargo, and extracellular conditions [26].
Quantitative Comparison of CPP Classes for Mitochondrial Targeting
Selecting CPPs for mitochondrial surface modification requires consideration of their intrinsic ability to localize to mitochondria post-cellular internalization. The following table summarizes key properties and mitochondrial targeting efficacy for several well-characterized and novel CPPs, including protein-derived sequences.
Table 2: CPP Profiles for Mitochondrial Targeting Applications
| CPP Name | Class & Origin | Sequence | Mitochondrial Localization Evidence | Key Findings & Notes |
|---|---|---|---|---|
| cAmbly [30] | Protein-derived (C-terminal of Amblyomin-X) | EEQTHFHFESPKLISFKVQDYWILNDIMKKNLTGISLKSEEEDADSGEID | Preferential colocalization with mitochondria in T98G cells within 30 min [30]. | Non-toxic (up to 2 µM); C-terminus is optimal for conjugation; derived from an antitumoral protein [30]. |
| (ChaZZ)₃ [32] | Synthetic Amphipathic Oligoproline | (ChaZZ)₃ (Z = Gup) | Strong colocalization with MitoTracker in MCF-7 cells at 10-20 µM [32]. | Rigid PPII helix with aligned cationic/hydrophobic edges; hydrophobicity tuning is crucial for selectivity [32]. |
| (ChaZZ)₂-Cha [32] | Synthetic Amphipathic Oligoproline (7-mer) | (ChaZZ)₂-Cha (Z = Gup) | Strong mitochondrial colocalization at 20 µM [32]. | Shorter analog with C-terminal Cha; demonstrates the impact of C-terminal hydrophobicity on uptake and targeting [32]. |
| TAT (47-57) [26] | Cationic, Protein-derived (HIV-1) | YGRKKRRQRRR | Not specifically mitochondrial; general cytoplasmic/nuclear localization [17]. | Widely used benchmark; efficient uptake but lacks inherent organelle specificity [17] [26]. |
| Penetratin [26] | Cationic, Protein-derived (Drosophila) | RQIKIWFQNRRMKWKK | Not specifically mitochondrial; general cytoplasmic/nuclear localization [17]. | Classic CPP; substitution of Trp14 with Phe abolishes membrane permeability [17] [26]. |
The data indicate that while classic cationic CPPs like TAT and Penetratin are efficient at cellular entry, they do not inherently target mitochondria. Specific mitochondrial targeting requires deliberate design, such as incorporating amphipathic structures with precisely tuned hydrophobicity [32] or deriving sequences from proteins with natural mitochondrial affinity [30].
Experimental Protocols for Evaluating Mitochondrial-Targeting CPPs
Protocol 1: Assessing Cellular Uptake and Intracellular Localization
This protocol outlines the steps to visualize and confirm the subcellular localization of fluorescently labeled CPPs, a critical first step in validation.
Materials:
- Synthesized CPPs: Conjugated with a fluorophore (e.g., Carboxyfluorescein (CF), FITC) [32] [30].
- Cell Line: Adherent cells relevant to the research context (e.g., MCF-7, T98G, HeLa) [32] [30].
- Live-Cell Imaging Media: Phenol-red free culture medium.
- Mitochondrial Stain: MitoTracker Deep Red or similar dye [32].
- Nuclear Stain: Hoechst 33342 [32].
- Confocal Microscope: Equipped with appropriate lasers and filters.
Procedure:
- Cell Seeding: Plate cells in glass-bottom culture dishes at a suitable density (e.g., 70-90% confluency) and allow them to adhere for 24 hours [30].
- Staining and Treatment:
- Replace the medium with fresh pre-warmed imaging medium.
- Incubate cells with the fluorescent CPP (e.g., 1-20 µM, concentration must be optimized) for a designated time (e.g., 30 minutes to 1 hour) at 37°C [32] [30].
- During the final 15-30 minutes of incubation, add the MitoTracker dye and Hoechst 33342 according to the manufacturer's instructions.
- Washing: After incubation, gently wash the cells three times with PBS or imaging medium to remove any non-internalized peptide and excess dye.
- Image Acquisition: Observe the cells immediately under the confocal microscope. Acquire Z-stack images to accurately determine colocalization.
- Data Analysis: Use image analysis software (e.g., ImageJ/Fiji with colocalization plugins) to calculate Pearson's correlation coefficient or Mander's overlap coefficient between the CPP and MitoTracker signals to quantify mitochondrial targeting [32].
Protocol 2: Functional Validation via Mitochondrial Functional Assay
This protocol assesses the functional consequences of CPP localization, such as disruption of the mitochondrial membrane potential, a key indicator of bioactivity.
Materials:
- CPP of Interest: Unlabeled or fluorescently labeled.
- JC-1 Dye: A cationic dye that exhibits potential-dependent accumulation in mitochondria.
- Cell Line and Culture Reagents.
- Flow Cytometer or Fluorescence Plate Reader.
Procedure:
- Cell Treatment: Seed cells in a multi-well plate. The following day, treat the cells with the desired concentration of the CPP. Include a negative control (vehicle alone) and a positive control (e.g., CCCP, a mitochondrial uncoupler).
- JC-1 Staining: After treatment (e.g., 4-24 hours), incubate cells with the JC-1 dye according to the manufacturer's protocol for 20-30 minutes at 37°C.
- Washing and Analysis: Gently wash the cells with PBS. Analyze the cells immediately.
- Flow Cytometry: Analyze the fluorescence in both the FITC (monomeric, green) and PE (aggregate, red) channels. A decrease in the red/green fluorescence ratio indicates mitochondrial depolarization.
- Fluorescence Microscopy/Plate Reader: Image the cells or read the fluorescence intensities. Healthy mitochondria with high membrane potential display orange/red J-aggregates, while depolarized mitochondria show green monomeric fluorescence.
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Mitochondrial-Targeting CPP Research
| Reagent / Tool | Function / Purpose | Example & Notes |
|---|---|---|
| Fluorophore-Conjugated CPPs | Direct visualization of cellular uptake and subcellular localization via microscopy [32] [30]. | CF-Ahx-(ChaZZ)3-NH2 [32]; cAmbly-FITC [30]. Ahx spacer minimizes fluorophore interference. |
| Live-Cell Mitochondrial Trackers | Counter-staining of mitochondria to assess CPP colocalization [32]. | MitoTracker Deep Red FM; stable for long-term tracking. |
| Cellular Viability Assays | Assessment of CPP cytotoxicity, crucial for therapeutic application [30]. | MTT assay: Measures mitochondrial metabolic activity [30]. |
| Mitochondrial Membrane Potential Probes | Functional analysis of mitochondrial health after CPP treatment. | JC-1, Tetramethylrhodamine (TMRM); detect early apoptosis or dysfunction. |
| pH-Responsive Nanoparticles | Delivery vehicle to shield cationic CPPs, improving tumor selectivity and reducing systemic toxicity [33]. | CA-NPs: Electrostatic self-assembly of cationic MTP with anionic, acid-sensitive polypeptide [33]. |
| Zwitterionic Polypeptides | Dissociation of plant cell wall amorphous layer for delivery in plant research [4]. | Facilitates penetration of CPP/cargo complexes through the physical barrier of cell walls [4]. |
Visualizing the Workflow and Targeting Strategy
The following diagrams illustrate the experimental workflow for evaluating mitochondrial-targeting CPPs and a strategic approach for designing organelle-specific peptides.
Figure 1: A linear workflow for the experimental evaluation of mitochondrial-targeting CPPs, from synthesis to functional validation.
Figure 2: Strategic principles for designing mitochondrial-targeting CPPs and their subsequent journey from cellular entry to mitochondrial localization. MTS: Mitochondrial Targeting Sequence; TOM/TIM: Translocase of the Outer/Inner Membrane.
Innovative Amphipathic Oligoprolines for Enhanced Mitochondrial Selectivity
The surface modification of mitochondria with Cell-Penetrating Peptides (CPPs) represents a frontier in therapeutic delivery strategies for addressing mitochondrial dysfunction. Within this field, amphipathic oligoprolines have emerged as a novel class of CPPs engineered for superior mitochondrial selectivity. These peptides are designed with a rigid, helical polyproline II (PPII) backbone that spatially segregates cationic and hydrophobic moieties, creating an optimal structure for crossing cellular and mitochondrial membranes [34]. This protocol details the application of these peptides for targeted mitochondrial delivery, providing a framework for researchers and drug development professionals to leverage their unique properties in the context of gene therapy, organelle transplantation, and the treatment of mitochondrial diseases.
Structural Design and Mechanism of Action
The enhanced mitochondrial selectivity of amphipathic oligoprolines is not accidental but is rooted in specific, engineered structural features. The design principles are as follows:
- Rigid PPII Helical Backbone: Unlike flexible peptides, the oligoproline backbone adopts a polyproline II (PPII) helix. This rigidity prevents the peptide from folding back on itself, ensuring that the functional groups remain correctly aligned for interaction with mitochondrial membranes [34].
- Aligned Cationic and Hydrophobic Residues: The helical structure allows for the precise spatial separation of cationic guanidinium groups and hydrophobic cyclohexyl groups along its edges. This amphipathic character is critical for interacting with the phospholipid bilayers of mitochondria [34].
- Systematic Hydrophobicity Tuning: The cellular uptake and mitochondrial selectivity can be optimized by introducing C-terminal and backbone modifications that systematically vary the overall hydrophobicity of the peptide [34].
The mechanism begins with cellular internalization, facilitated by the peptide's cell-penetrating properties. Once inside the cell, the specific amphipathic and structural profile of the oligoproline promotes its interaction with the mitochondrial outer membrane, leading to translocation and subsequent time-dependent redistribution that results in prolonged mitochondrial residency [34].
The diagram below illustrates this targeted delivery pathway.
Quantitative Characterization of Oligoproline CPPs
The performance of amphipathic oligoprolines can be quantified through various physicochemical and biological assays. Comparative studies with more flexible peptide analogs consistently demonstrate the benefit of the rigid, aligned design for mitochondrial targeting [34].
Table 1: Key Quantitative Data from Oligoproline CPP Studies
| Parameter | Measurement | Significance |
|---|---|---|
| Cellular Uptake Efficiency | Enhanced uptake compared to flexible analogs | Indicates improved ability to cross the plasma membrane [34]. |
| Mitochondrial Selectivity | High colocalization with mitochondrial markers | Confirms specific targeting to mitochondria over other organelles [34]. |
| Intracellular Kinetics | Time-dependent redistribution to mitochondria | Suggests an active or facilitated process beyond passive diffusion [34]. |
| Retention Time | Prolonged mitochondrial residency | Critical for sustained therapeutic effect [34]. |
Experimental Protocol: Mitochondrial Targeting Using Amphipathic Oligoprolines
This protocol outlines the methodology for evaluating the mitochondrial targeting efficiency of novel amphipathic oligoproline CPPs in a cell culture model.
Materials and Reagents
- Synthesized Amphipathic Oligoproline CPP: Designed with a PPII helix backbone, cationic guanidinium groups, and hydrophobic cyclohexyl groups [34].
- Fluorescent Tag: e.g., FITC or a similar fluorophore for conjugation and tracking.
- Cell Line: Appropriate mammalian cell line (e.g., HEK293, HeLa).
- Culture Media: Standard growth medium (e.g., DMEM) supplemented with fetal bovine serum (FBS) and penicillin/streptomycin [35].
- Mitochondrial Stain: e.g., MitoTracker Red CMXRos [36].
- Fixative: 4% paraformaldehyde (PFA) in PBS.
- Permeabilization Buffer: PBS with 0.1% Triton X-100.
- Mounting Medium: Antifade mounting medium with DAPI.
- Imaging Equipment: Confocal laser-scanning microscope (CLSM) [35].
Step-by-Step Procedure
Step 1: Peptide Preparation
- Reconstitute the lyophilized, fluorescently-labeled oligoproline CPP in sterile ultrapure water or DMSO to create a stock solution.
- Dilute the stock solution in serum-free culture medium to the desired working concentration (typically in the µM range).
Step 2: Cell Culture and Seeding
- Culture the chosen cell line under standard conditions (37°C, 5% CO₂).
- Seed cells onto glass-bottom culture dishes or multi-well plates suitable for microscopy at a density that will reach 60-70% confluence at the time of treatment.
Step 3: Peptide Treatment and Incubation
- Replace the cell culture medium with the prepared peptide-containing serum-free medium.
- Incubate cells for a predetermined time (e.g., 1-4 hours) at 37°C with 5% CO₂ to allow for cellular uptake and mitochondrial localization [34].
Step 4: Mitochondrial Staining
- Following peptide incubation, prepare a working solution of MitoTracker Red CMXRos (e.g., 100-500 nM) in pre-warmed serum-free medium.
- Remove the peptide-containing medium, wash cells gently with PBS, and add the MitoTracker solution.
- Incubate for 15-30 minutes at 37°C under culture conditions.
Step 5: Cell Fixation and Preparation for Imaging
- Aspirate the MitoTracker solution and wash cells gently with PBS.
- Fix cells with 4% PFA for 15 minutes at room temperature.
- Wash cells three times with PBS to remove residual PFA.
- (Optional) Permeabilize cells with 0.1% Triton X-100 for 10 minutes if intracellular immunostaining is required.
- Add a drop of antifade mounting medium with DAPI to stain the nucleus and mount the coverslip.
Step 6: Confocal Microscopy and Image Analysis
- Image the cells using a confocal laser-scanning microscope (CLSM). Acquire images using appropriate laser lines and filter sets for DAPI, FITC (or the chosen CPP fluorophore), and MitoTracker Red [36].
- Use image analysis software (e.g., ImageJ) to quantify the degree of colocalization between the green (CPP) and red (mitochondria) signals. The Pearson's correlation coefficient or Manders' overlap coefficient are standard metrics for this analysis.
The workflow for this protocol is summarized in the diagram below.
The Scientist's Toolkit: Key Research Reagent Solutions
Successful execution of mitochondrial targeting experiments relies on a suite of specialized reagents. The table below catalogs essential materials and their functions.
Table 2: Essential Research Reagents for Mitochondrial Targeting Studies
| Research Reagent | Function / Application | Example & Notes |
|---|---|---|
| Amphipathic Oligoproline CPPs | Core targeting moiety; facilitates cellular and mitochondrial uptake. | Rigid PPII helix with aligned guanidinium/cationic and cyclohexyl/hydrophobic groups [34]. |
| Mitochondrial Vital Dyes | Visualizing the endogenous mitochondrial network in live or fixed cells. | MitoTracker Red CMXRos, MitoTracker Green FM [36]. |
| Triphenylphosphonium (TPP) | A classic mitochondrial-targeting ligand that exploits the mitochondrial membrane potential. | Often conjugated to peptides or nanoparticles as a targeting moiety [35] [4]. |
| DSPE-PEG Polymer | A coating polymer for surface engineering of isolated mitochondria or nanoparticles to enhance stability and enable further functionalization. | Used to create a stealth layer and conjugate targeting peptides (e.g., VCAM-1 binding peptide) for specific delivery [36]. |
| Endosomal pH-Responsive Polymer | A key component of nanoparticle systems to facilitate endosomal escape and release cargo into the cytosol. | e.g., Meo-PEG-b-PDPA, which disrupts endosomes via the "proton sponge" effect [35]. |
| Seahorse XF Analyzer | Instrument for measuring mitochondrial function in live cells (OCR and ECAR). | Assesses bioenergetic restoration after therapeutic intervention [36]. |
Concluding Remarks
Amphipathic oligoprolines provide a robust and selective platform for mitochondrial targeting, underpinned by their unique structural rigidity and amphipathic design. The application notes and detailed protocol provided here serve as a foundation for researchers to incorporate these innovative peptides into their work on mitochondrial medicine. The continued development of such CPPs, especially when combined with other advanced delivery platforms like surface-engineered mitochondria or targeted nanoparticles, holds significant promise for addressing a broad spectrum of mitochondrial-related diseases.
The DSPE-PEG (1,2-distearoyl-sn-glycero-3-phosphoethanolamine-polyethylene glycol) platform represents a cornerstone in the design of advanced drug delivery systems, particularly for challenging subcellular targets like mitochondria. This phospholipid-polymer hybrid combines the biomimetic properties of the phospholipid DSPE with the stealth and functionalization capabilities of the polymer PEG. Within the context of mitochondrial drug delivery, the platform's utility is critical; mitochondria are key regulators of cell survival and death, and their dysfunction is implicated in cancer, neurodegenerative diseases, and metabolic disorders [37] [20]. Effective delivery of therapeutics to mitochondria is fraught with challenges, including the need for deep tissue penetration, cellular internalization, endosomal escape, and finally, traversal across the mitochondrial membranes [38]. The DSPE-PEG platform provides a versatile foundation for constructing nanocarriers that can be engineered with mitochondrial-penetrating peptides (MPPs) and cell-penetrating peptides (CPPs) to overcome these barriers, offering a promising path for enhancing the efficacy of mitochondrially-targeted therapies [38] [4].
Molecular Structure and Functional Design
The molecular architecture of DSPE-PEG is ingeniously composed of three distinct functional domains, each contributing to its overall performance in nanocarrier assembly and biological interaction.
- DSPE (Hydrophobic Anchor): This phospholipid moiety features two C18 saturated fatty acid chains. Its structure is analogous to the phospholipids found in natural cell membranes, allowing it to integrate stably into the hydrophobic bilayers of liposomes, lipid nanoparticles (LNPs), and other nanocarriers. This integration provides a strong anchoring point, ensuring the polymer remains associated with the particle and contributes to overall formulation stability [39].
- PEG (Hydrophilic Stealth Layer): Attached to the polar head of DSPE is a poly(ethylene glycol) chain. This hydrophilic segment extends into the aqueous environment, forming a protective "stealth" layer around the nanoparticle. Through steric hindrance and strong hydration, this PEG corona minimizes opsonization—the adsorption of plasma proteins that would otherwise mark the nanoparticle for rapid clearance by the immune system. This effect significantly extends the nanoparticle's circulation time in the bloodstream, which is essential for allowing sufficient time to accumulate in target tissues like tumors via the Enhanced Permeability and Retention (EPR) effect [40] [39].
- Functional Termini (Reactive Handles): The distal end of the PEG chain can be modified with various functional groups, transforming the platform from a passive carrier to an active targeting and conjugation tool. Common termini include maleimide (MAL), carboxyl (COOH), and amine (NH2). Among these, maleimide (MAL) is particularly valuable for bioconjugation, as it reacts rapidly and specifically with thiol (-SH) groups under mildly acidic to neutral conditions (pH 6.5–7.2) to form stable thioether bonds. This enables the efficient and covalent attachment of targeting ligands, such as mitochondrial signal peptides, antibodies, or other homing molecules [40] [39].
Table 1: Common DSPE-PEG Derivatives and Their Key Characteristics
| Derivative | Terminal Group | Key Properties | Primary Conjugation Chemistry |
|---|---|---|---|
| DSPE-PEG-MAL | Maleimide | Rapid, specific reaction with thiols; ideal for antibodies, peptides | Thioether bond via Michael addition |
| DSPE-PEG-COOH | Carboxyl | Activatable for reaction with amines; versatile for small molecules | Amide bond formation (e.g., via EDC/NHS) |
| DSPE-PEG-NH2 | Amine | Reacts with carboxyls or other electrophiles; useful for small molecules | Amide bond or other covalent linkages |
Quantitative Performance Data
The efficacy of nanocarriers incorporating DSPE-PEG and mitochondrial targeting motifs has been quantitatively demonstrated in recent studies. The data reveals significant advantages over traditional targeting strategies.
Table 2: Quantitative Comparison of Mitochondria-Targeted Liposomal Formulations
| Formulation | Key Targeting Component | Mitochondrial Targeting Efficiency (vs. L-TPP) | Key Experimental Findings | Reference |
|---|---|---|---|---|
| L-G2R | 2nd-gen arginine-rich dendritic lipopeptide (G2R) | ~5.7-fold higher | Superior cellular uptake via macropinocytosis; efficient endosome escape | [38] |
| L-G2R-DA | G2R with pH-sensitive 2,3-dimethylmaleic acid (DA) | ~3.7-fold higher | Charge-reversible ("stealth") property enhances tumor accumulation; high targeting | [38] |
| L-TPP | Triphenylphosphonium cation | 1.0 (Baseline) | Common delocalized lipophilic cation; can cause proton leakage & cytotoxicity at high conc. | [38] [4] |
These studies highlight that dendritic arginine-rich lipopeptides, which mimic natural mitochondrial precursor proteins, can achieve far greater mitochondrial accumulation than conventional lipophilic cations like TPP [38]. Furthermore, the incorporation of pH-sensitive groups (like DA) provides a "stealth" function that shields positive charges during circulation, reducing non-specific interactions, but is cleaved in the acidic tumor microenvironment to re-expose the cationic, mitochondria-penetrating surface [38].
Application Protocols for Mitochondrial Targeting
The following protocols outline key methodologies for leveraging the DSPE-PEG platform in the development of mitochondria-specific drug delivery systems.
Protocol 1: Synthesis and Conjugation of Maleimide-Functionalized DSPE-PEG
This protocol describes the synthesis of the key building block, DSPE-PEG-MAL, and its conjugation to a thiolated mitochondrial-penetrating peptide (MPP).
Materials:
- DSPE-PEG-NH2 (e.g., DSPE-PEG2000-Amine)
- N-succinimidyl 3-(maleimidopropionate) (Mal-PEG-NHS)
- Triethylamine (TEA)
- Anhydrous chloroform/dimethylformamide (DMF) mixture
- Thiolated Mitochondrial-Penetrating Peptide (e.g., peptide with C-terminal cysteine)
- Purification equipment: Dialysis tubing (MWCO 3.5 kDa) or Size-Exclusion Chromatography (SEC) system
Procedure:
- Activation: Dissolve DSPE-PEG-NH2 and a molar excess of Mal-PEG-NHS (1.2:1 ratio) in an anhydrous mixture of chloroform and DMF (2:1 v/v).
- Reaction: Add triethylamine (TEA, 2-3 molar equivalents) as a catalyst to the reaction mixture. Stir the solution vigorously under an inert atmosphere (e.g., nitrogen or argon) for 12-16 hours at room temperature.
- Purification: Isolate the crude DSPE-PEG-MAL product using rotary evaporation to remove solvents. Purify further via dialysis against ultrapure water or size-exclusion chromatography to remove unreacted reagents and catalyst. Lyophilize the pure product for long-term storage.
- Conjugation: Hydrate the purified DSPE-PEG-MAL in a degassed phosphate buffer (pH 7.0). Add a slight molar excess of the thiolated MPP. Allow the conjugation reaction to proceed for 4-6 hours at room temperature.
- Verification: Confirm successful conjugation using analytical techniques such as MALDI-TOF Mass Spectrometry to observe the mass shift, and thin-layer chromatography (TLC) to monitor reaction completion [40] [39].
Protocol 2: Formulation of Mitochondria-Targeted Hybrid Nanoparticles
This protocol details the preparation of lipid-polymer hybrid nanoparticles (LPHNPs) incorporating the DSPE-PEG-MPP conjugate for mitochondrial delivery.
Materials:
- Polymer core material (e.g., PLGA, PCL)
- Lipids: Soy phosphatidylcholine (SPC), Cholesterol, DSPE-PEG-MPP conjugate
- Therapeutic cargo (e.g., drug, gene, photosensitizer like Indocyanine Green - ICG)
- Organic solvent (e.g., dichloromethane, ethanol)
- Equipment: Probe sonicator, Dynamic Light Scattering (DLS)/Zetasizer, Dialysis tubing
Procedure:
- Lipid and Polymer Dissolution: Dissolve the polymer (e.g., PLGA, 50 mg), SPC (20 mg), cholesterol (10 mg), and the DSPE-PEG-MPP conjugate (5-10 mol% of total lipid) in a suitable organic solvent like dichloromethane.
- Formation of Thin Film: Remove the organic solvent using rotary evaporation, forming a thin, uniform lipid-polymer film on the walls of a round-bottom flask.
- Hydration and Nanoparticle Formation: Hydrate the dried film with an aqueous buffer (e.g., PBS, pH 7.4) containing the therapeutic cargo if encapsulating a hydrophilic agent. For hydrophobic drugs, they are co-dissolved in the organic solvent in step 1. Vigorously agitate the mixture above the phase transition temperature of the lipids to form multilamellar vesicles.
- Size Reduction and Homogenization: Subject the hydrated suspension to probe sonication on an ice bath or extrude it through polycarbonate membranes (e.g., 100-200 nm pores) to form small, monodisperse nanoparticles.
- Purification and Characterization: Purify the formulated nanoparticles from non-encapsulated cargo and free ligands using dialysis or gel filtration. Characterize the final formulation for:
Protocol 3: In Vitro Assessment of Mitochondrial Targeting and Colocalization
This protocol is used to validate the mitochondrial targeting efficiency of the developed formulation.
Materials:
- Cell culture (e.g., 4T1 breast cancer cells)
- Nanoparticle formulation labeled with a fluorescent dye (e.g., Coumarin 6, Dil)
- MitoTracker Green FM (or other mitochondrial-specific dyes)
- Inhibitors: Cytochalasin B (macropinocytosis inhibitor)
- Confocal Laser Scanning Microscopy (CLSM), Flow Cytometry
Procedure:
- Cell Seeding: Seed cells onto glass-bottom confocal dishes and culture until they reach 60-70% confluence.
- Treatment and Inhibition:
- Pre-treat a set of cells with cytochalasin B (5 µg/mL) for 1 hour to inhibit macropinocytosis.
- Co-incubate cells with the fluorescently labeled nanoparticle formulation (e.g., L-G2R-DA) and MitoTracker Green FM (100 nM) for a predetermined period (e.g., 2-4 hours).
- Imaging and Analysis:
- After incubation, wash the cells thoroughly with PBS to remove uninternalized nanoparticles.
- Fix the cells with paraformaldehyde (4%) and mount for imaging.
- Acquire high-resolution images using Confocal Laser Scanning Microscopy (CLSM).
- Quantify the colocalization of the nanoparticle signal (red) with the mitochondrial signal (green) using image analysis software (e.g., ImageJ with JaCoP plugin) by calculating Pearson's correlation coefficient or Mander's overlap coefficient [38].
- Independent Validation: Isolate mitochondria from treated cells using a mitochondrial isolation kit and analyze the fluorescence associated with the mitochondrial fraction using flow cytometry to quantitatively determine nanoparticle accumulation, as referenced in Table 2 [38].
Visualizing the Workflow and Mechanism
The following diagrams illustrate the preparation process and the intracellular mechanism of action for mitochondria-targeted DSPE-PEG hybrid nanoparticles.
Figure 1. Schematic workflow for the preparation of mitochondria-targeted lipid-polymer hybrid nanoparticles using the DSPE-PEG platform, from component dissolution to final characterization.
Figure 2. The intracellular trafficking pathway of targeted nanoparticles. The nanoparticles enter cells via energy-dependent macropinocytosis, escape from endosomes, and are guided to mitochondria by the MPP ligand, which interacts with the Tom translocase complex on the mitochondrial membrane [38] [4].
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagents for Developing DSPE-PEG Mitochondrial Delivery Systems
| Reagent / Material | Function / Role | Specific Example(s) |
|---|---|---|
| DSPE-PEG-X Functional Lipids | Amphiphilic anchor for nanoparticle surface functionalization. | DSPE-PEG-MAL, DSPE-PEG-COOH, DSPE-PEG-NH2 [40] [39]. |
| Mitochondrial-Penetrating Peptides (MPPs) | Ligand that mimics natural protein signals to direct carriers to mitochondria. | Arginine-rich dendritic peptides (e.g., G2R), Presequence peptides derived from native mitochondrial proteins [38] [4]. |
| Polymer Core Materials | Forms the structural core of the hybrid nanoparticle for drug encapsulation. | PLGA, PCL, PLA [41] [42]. |
| Auxiliary Lipids | Provide structural integrity and fluidity to the lipid shell of nanoparticles. | Soy Phosphatidylcholine (SPC), Cholesterol, DOPE [38] [42]. |
| Therapeutic Cargos | Active agents to be delivered to mitochondria to exert a therapeutic effect. | Photosensitizers (ICG), Chemotherapeutics, Pro-apoptotic drugs, mtDNA [37] [20] [38]. |
| Characterization Tools | Essential for quantifying nanoparticle properties and biological efficacy. | DLS (Size/PDI), Zetasizer (Zeta Potential), CLSM (Colocalization), Flow Cytometry (Quantitative Uptake) [38]. |
The surface modification of mitochondria with specific targeting ligands represents a cutting-edge strategy in the field of organelle transplantation and targeted drug delivery. Within the broader context of enhancing mitochondrial delivery via surface engineering with cell-penetrating peptides (CPPs), the conjugation of VCAM-1-binding and collagen-binding peptides enables precise targeting to dysfunctional endothelium. This approach addresses a critical limitation of conventional mitochondrial transplantation: the lack of target specificity, which results in inefficient uptake and limited long-term retention of transplanted organelles [43] [36].
Vascular cell adhesion molecule-1 (VCAM-1) is upregulated on endothelial cells during inflammation, providing an excellent target for drug delivery systems in vascular diseases [44] [45]. Similarly, collagen-binding peptides target exposed subendothelial matrix proteins that become accessible upon endothelial injury [36]. The strategic combination of these targeting modalities on mitochondrial surfaces creates a sophisticated delivery platform capable of recognizing both cellular and structural markers of vascular pathology, thereby enhancing delivery precision to damaged tissues.
Ligand Characterization and Targeting Mechanisms
VCAM-1 Binding Peptide (VBP)
The VCAM-1 binding peptide (VHPKQHRGGSKGC) demonstrates specific affinity for VCAM-1, an adhesion molecule significantly overexpressed on activated endothelial cells in inflammatory conditions such as atherosclerosis, rheumatoid arthritis, and diabetic vascular disease [36] [44]. The peptide's binding mechanism involves interaction with the exposed binding domains of VCAM-1, facilitating selective accumulation on inflamed endothelium. This targeting approach has been successfully employed in various nanocarrier systems, including lipopolyplexes for gene delivery and hybrid micelles for anti-inflammatory therapy, demonstrating its versatility and efficacy [44] [45].
Collagen Binding Peptide (CBP)
The collagen binding peptide (CQDSETRTFY) exhibits specific binding affinity for collagen types I and IV, which are major components of the vascular basement membrane and subendothelial matrix [36] [46]. This peptide enables targeting to sites where endothelial denudation has occurred, exposing the underlying collagen matrix. Such targeting is particularly valuable in acute vascular injuries, including those induced by endovascular procedures or ischemia-reperfusion events [36]. The binding mechanism involves specific molecular recognition of collagen fibrils, with studies demonstrating that peptides maintaining a triple-helical structure show enhanced binding affinity [46].
Table 1: Characterization of Targeting Peptides
| Parameter | VCAM-1 Binding Peptide (VBP) | Collagen Binding Peptide (CBP) |
|---|---|---|
| Amino Acid Sequence | VHPKQHRGGSKGC | CQDSETRTFY |
| Target | VCAM-1 on inflamed endothelium | Collagen I/IV in exposed subendothelium |
| Targeting Context | Chronic inflammation (atherosclerosis, diabetes) | Acute injury (vascular procedures, denudation) |
| Binding Mechanism | Protein-protein interaction with VCAM-1 domains | Molecular recognition of collagen fibrils |
| Conjugation Chemistry | Thiol-maleimide click chemistry | Thiol-maleimide click chemistry |
Experimental Protocols
Polymer-Peptide Conjugate Synthesis
Objective: To synthesize DSPE-PEG-peptide conjugates for mitochondrial surface functionalization.
Materials:
- DSPE-PEG-MAL (5,000 Da, Nanosoft Polymers)
- Biotinylated VBP and CBP (ABI Scientifics)
- Slide-a-Lyzer Dialysis Cassette (MWCO 7,000 Da, ThermoFisher)
- Ultrapure water
- Lyophilizer
Procedure:
- Prepare peptide solution in ultrapure water at appropriate concentration.
- React peptides with DSPE-PEG-MAL at a 1:1 thiol:maleimide molar equivalent ratio.
- Incubate reaction mixture at room temperature for 24 hours with gentle agitation.
- Purify reaction products by dialysis against ultrapure water for 24 hours.
- Lyophilize purified conjugates and store at -20°C until use.
- Verify successful conjugation using MALDI mass spectrometry [36].
Mitochondrial Surface Engineering
Objective: To functionalize isolated mitochondria with targeting peptide conjugates.
Materials:
- Freshly isolated mitochondria from iPSC-MSCs
- DSPE-PEG-peptide conjugates (1 mg/mL in Reagent C buffer)
- Mitochondria Isolation Kit (Thermo Fisher Scientific)
- Refrigerated centrifuge
Procedure:
- Isolate mitochondria from iPSC-MSCs using mitochondrial isolation kit according to manufacturer's protocol.
- Quantify mitochondrial protein content using BCA assay.
- Combine mitochondrial aliquots with DSPE-PEG-peptide conjugate solutions at optimized mass ratios (polymer to mitochondrial protein).
- Incubate mixture on ice for 3 hours with continuous shaking.
- Rinse functionalized mitochondria by centrifugation at 12,000×g for 5 minutes.
- Replace supernatant with fresh Reagent C buffer and repeat washing step.
- Resuspend final mitochondrial pellet in Mitochondria Storage Buffer.
- Use immediately for experiments or store at -80°C for long-term preservation (up to 14 days) [36].
Binding Assay for Functionalized Mitochondria
Objective: To validate the binding capacity of peptide-functionalized mitochondria.
Materials:
- Collagen-coated plates (Type I or IV)
- VCAM-1 coated surfaces
- Fluorescence-labeled streptavidin
- Plate reader
- Confocal microscopy equipment
Procedure for Collagen Binding:
- Coat plates with collagen I (10 μg/cm²) or use commercial collagen IV-coated strips.
- Add increasing concentrations of biotinylated CBP and DSPE-PEG-CBP-biotin.
- Incubate for 3 hours at 37°C.
- Rinse with PBS to remove unbound peptides.
- Incubate with streptavidin-594 (4 μg/mL) for 30 minutes.
- Measure fluorescence with plate reader to quantify binding [36].
Procedure for Mitochondrial Uptake Assessment:
- Culture human diabetic aortic endothelial cells (DAECs) under appropriate conditions.
- Treat with functionalized mitochondria labeled with Mitotracker.
- Incubate for predetermined time points (4-24 hours).
- Analyze using flow cytometry and confocal microscopy.
- Quantify uptake efficiency and intracellular distribution [36].
Quantitative Characterization Data
Performance Metrics of Engineered Mitochondria
Table 2: Functional Characterization of Surface-Engineered Mitochondria
| Parameter | Uncoated Mitochondria | DSPE-PEG Coated | VBP-Functionalized | CBP-Functionalized |
|---|---|---|---|---|
| Cellular Uptake in DAECs | Baseline | ~2-3x increase | Significant enhancement | Significant enhancement |
| Cytoplasmic Retention (24h) | Low | Moderate | High | High |
| Colocalization with Endogenous Network | Limited | Moderate | Extensive | Extensive |
| Membrane Potential | Variable | Improved | Significantly improved | Significantly improved |
| Oxygen Consumption Rate | Baseline | Sustained | Enhanced | Enhanced |
| Targeting Specificity | None | Passive | Active (VCAM-1 mediated) | Active (collagen mediated) |
Experimental Workflow
The following diagram illustrates the comprehensive workflow for the conjugation of targeting ligands to mitochondria and subsequent functional validation:
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Research Reagents for Mitochondrial Ligand Conjugation
| Reagent/Kit | Supplier | Function/Application |
|---|---|---|
| DSPE-PEG-MAL (5,000 Da) | Nanosoft Polymers | Polymer backbone for peptide conjugation via thiol-maleimide chemistry |
| Custom Peptides (VBP, CBP) | ABI Scientifics | Targeting ligands for specific endothelial recognition |
| Mitochondria Isolation Kit | Thermo Fisher Scientific | Isolation of intact, functional mitochondria from source cells |
| STEMdiff Mesenchymal Progenitor Kit | Stemcell Technologies | Differentiation of iPSCs to MSCs for mitochondrial isolation |
| Seahorse XF Analyzer Reagents | Agilent Technologies | Assessment of mitochondrial bioenergetics (OCR, ATP production) |
| MitoTracker Probes | Thermo Fisher Scientific | Fluorescent labeling of mitochondria for tracking and visualization |
| Slide-a-Lyzer Dialysis Cassettes | ThermoFisher | Purification of DSPE-PEG-peptide conjugates |
| Collagen-Coated Plates (I/IV) | Sigma-Aldrich | Substrates for binding assays with collagen-targeting constructs |
The conjugation of VCAM-1 and collagen-binding peptides to mitochondrial surfaces represents a significant advancement in targeted organelle delivery systems. The detailed protocols presented herein provide researchers with a robust methodology for engineering mitochondria with enhanced targeting capabilities toward dysfunctional endothelium. The quantitative data demonstrates clear improvements in cellular uptake, retention, and functional integration compared to unmodified mitochondria.
This approach establishes a foundation for developing precision mitochondrial therapies for vascular diseases characterized by endothelial dysfunction, including atherosclerosis, diabetic vasculopathy, and ischemia-reperfusion injury. The modular nature of the surface engineering platform further allows for adaptation to other targeting ligands and therapeutic applications beyond vascular repair.
Combining CPPs with Mitochondrial Targeting Agents like TPP
Application Notes
The strategic combination of Cell-Penetrating Peptides (CPPs) and Triphenylphosphonium (TPP) cations creates a powerful dual-targeting system for subcellular drug delivery. This approach is designed to overcome the multiple biological barriers that impede conventional therapeutics, enhancing the precision and efficacy of treatments for cancer, neurodegenerative diseases, and primary mitochondrial disorders [47] [48].
The rationale for this hybrid system hinges on a sequential targeting mechanism: CPPs mediate efficient cellular internalization across the plasma membrane, while the TPP moiety exploits the elevated mitochondrial membrane potential (ΔΨm) to achieve final organelle accumulation [49] [48]. This is particularly effective in cancer cells, which often exhibit a higher ΔΨm than healthy cells, allowing for selective accumulation of TPP-conjugated compounds and a resulting therapeutic advantage [50] [51].
This platform is highly versatile and can deliver diverse cargoes, including small molecule drugs, nucleic acids (DNA, RNA), and probes for diagnostic imaging [47] [51]. The successful application of this strategy requires careful design considerations of the nanocarrier system, the choice of CPP and linker chemistry, and rigorous validation of subcellular localization [48].
Table 1: Key Characteristics of Mitochondrial Targeting Agents
| Targeting Agent | Targeting Mechanism | Key Advantages | Reported Efficacy/Performance |
|---|---|---|---|
| Triphenylphosphonium (TPP) | Accumulates due to negative mitochondrial membrane potential (ΔΨm) [51] | High stability, simple synthesis, proven clinical safety (e.g., MitoQ) [51] | Can achieve >1000-fold higher concentration in mitochondria versus medium [51] |
| Cell-Penetrating Peptides (CPPs) | Energy-dependent or independent internalization across plasma membrane [48] | Enhances cellular uptake of cargo, overcomes first cellular barrier [48] | Performance is highly sequence-dependent; essential for initial cellular entry [48] |
| TPP-Functionalized Nanocarrier | Combines EPR effect (passive tumor targeting) and mitochondrial targeting [49] | Reduces required drug dosage, minimizes off-target effects [49] | Superior to non-targeted nanocarriers in inducing cancer cell death [50] [49] |
| Multi-TPP BODIPY Probe (5TPP) | Multiple TPP groups enhance mitochondrial affinity [52] | Improved water solubility, photostability, and "turn-on" fluorescence sensing [52] | Superior mitochondrial staining and permeation compared to single-TPP analogs [52] |
Table 2: Design Parameters for CPP-TPP Hybrid Systems
| Parameter | Impact on Delivery Efficiency | Recommendations & Considerations |
|---|---|---|
| Linker Chemistry | Influences stability, cargo release, and overall pharmacokinetics [51] | Use cleavable linkers (e.g., disulfide bonds, ester linkages) for controlled intracellular release [48]. |
| Number of TPP Moieties | Directly correlates with mitochondrial uptake and targeting specificity [52] | Multi-valent TPP presentation (e.g., 5TPP) can significantly enhance accumulation over mono-valent TPP [52]. |
| Nanocarrier Selection | Affects drug loading, circulation time, and biocompatibility [49] [53] | Common choices: Liposomes, polymeric NPs, dendrimers. Selection should be based on cargo type (hydrophobic/hydrophilic, size) [49] [53]. |
| Surface Functionalization | Mitigates nonspecific interactions, enhances stability, and enables active targeting [49] [48] | PEGylation ("stealth" coating) is common. Additional ligands (e.g., folic acid) can enable active cellular targeting [49]. |
Experimental Protocols
Protocol: Formulation and Preparation of CPP-TPP-Functionalized Liposomes
This protocol details the synthesis of a dual-targeted liposomal system for mitochondrial delivery of a hydrophobic agent (e.g., Curcumin).
I. Materials
- Lipids: 1,2-distearoyl-sn-glycero-3-phosphocholine (DSPC), Cholesterol, 1,2-distearoyl-sn-glycero-3-phosphoethanolamine (DSPE)
- Functional Lipids: DSPE-PEG2000, DSPE-PEG2000-Maleimide (for conjugation)
- Targeting Ligands: TPP-Amine (commercially available), CPP peptide with a C-terminal cysteine (e.g., TAT, R8)
- Cargo: Curcumin (or other hydrophobic drug)
- Solvents: Chloroform, Methanol, Isopropanol
- Buffers: HEPES Buffered Saline (HBS, pH 7.4), Calibration Buffer (pH 7.4)
II. Equipment
- Round-bottom flask
- Rotary evaporator
- Water bath sonicator
- Extruder with polycarbonate membranes (100 nm pore size)
- Size and Zeta Potential Analyzer
III. Procedure Step 1: Pre-conjugation of TPP to DSPE-PEG2000
- Dissolve DSPE-PEG2000-COOH (1.5 μmol) and TPP-NH2 (1.8 μmol) in anhydrous DMF.
- Add N,N'-Dicyclohexylcarbodiimide (DCC, 3.0 μmol) and N-Hydroxysuccinimide (NHS, 3.0 μmol).
- React for 24 hours at room temperature with stirring.
- Precipitate the product, DSPE-PEG2000-TPP, in cold diethyl ether and purify by dialysis against methanol/water.
Step 2: Liposome Preparation via Thin-Film Hydration & Extrusion
- Prepare Lipid Mixture: Dissolve DSPC (65 μmol), Cholesterol (30 μmol), DSPE-PEG2000 (4.25 μmol), and the synthesized DSPE-PEG2000-TPP (0.75 μmol) in chloroform in a round-bottom flask. Add the drug (e.g., Curcumin, 5 mol% relative to total lipid).
- Form Thin Film: Remove solvent using a rotary evaporator (40°C) to form a thin, dry lipid film.
- Hydration: Hydrate the film with 5 mL HBS (pH 7.4) at 60°C (above lipid phase transition temperature) for 1 hour with occasional vortexing to form multilamellar vesicles (MLVs).
- Size Reduction: Extrude the MLV suspension 21 times through two stacked 100 nm polycarbonate membranes using a heated extruder (60°C) to form small, unilamellar vesicles (SUVs).
Step 3: Conjugation of CPP to Liposome Surface
- Activate the maleimide groups on the liposomes by ensuring they are in HBS (pH 7.4).
- Incubate the liposomes with the CPP peptide (bearing a C-terminal cysteine) at a 1.5:1 (peptide:maleimide) molar ratio for 4 hours at room temperature under gentle stirring.
- Purify the final CPP-TPP-functionalized liposomes from unreacted peptide by size-exclusion chromatography (e.g., Sephadex G-25) using HBS as the eluent.
Step 4: Characterization
- Size and PDI: Determine the hydrodynamic diameter and polydispersity index (PDI) of the liposomes using Dynamic Light Scattering (DLS). Target: ~100-120 nm, PDI < 0.2.
- Zeta Potential: Measure the surface charge in HBS (pH 7.4) using a Zeta Potential Analyzer.
- Drug Encapsulation Efficiency (EE): Separate unencapsulated drug by ultracentrifugation (100,000 g, 1 h). Lyse an aliquot of liposomes with 1% Triton X-100 and quantify drug content via HPLC or UV-Vis spectroscopy. Calculate EE% = (Amount of drug in liposomes / Total amount of drug used) * 100.
Protocol: Validation of Mitochondrial Targeting and Function
I. Materials
- Cell line (e.g., HeLa, MCF-7)
- MitoTracker Deep Red (or Green)
- Tetramethylrhodamine, Methyl Ester (TMRM) or JC-1 dye (for ΔΨm measurement)
- Commercial MitoSOX Red reagent (for mitochondrial ROS)
- Hoechst 33342 (nuclear stain)
- Paraformaldehyde (4% in PBS)
- Confocal microscopy dishes
II. Equipment
- Confocal Laser Scanning Microscope (CLSM)
- Flow cytometer
- Cell culture incubator
III. Procedure Step 1: Colocalization Studies via Confocal Microscopy
- Seed cells in confocal dishes and incubate until 60-70% confluent.
- Treat cells with the CPP-TPP nanocarrier loaded with a fluorescent cargo (e.g., Coumarin 6, FITC) for 4 hours.
- Stain mitochondria with MitoTracker Deep Red (100 nM) for 30 minutes.
- Stain nuclei with Hoechst 33342 (5 μg/mL) for 10 minutes.
- Wash cells with PBS and image immediately using a CLSM. Use sequential scanning to avoid bleed-through.
- Analysis: Quantify colocalization using Manders' overlap coefficient or Pearson's correlation coefficient (R) with image analysis software (e.g., ImageJ with JACoP plugin). An R value > 0.7 indicates strong colocalization.
Step 2: Assessing Impact on Mitochondrial Membrane Potential (ΔΨm)
- Seed cells in a multi-well plate and treat with drug-loaded CPP-TPP nanocarriers for 12-24 hours.
- Load cells with JC-1 dye (2 μM) or TMRM (100 nM) for 30 minutes at 37°C.
- For JC-1: Analyze by flow cytometry or fluorescence microscopy. Healthy mitochondria (high ΔΨm) show red J-aggregates (emission ~590 nm). Depolarized mitochondria (low ΔΨm) show green J-monomers (emission ~529 nm). Calculate the red/green fluorescence intensity ratio.
- A decrease in the red/green ratio in treated cells indicates a loss of ΔΨm, a hallmark of early apoptosis induced by successful mitochondrial targeting.
Step 3: Measuring Mitochondrial Reactive Oxygen Species (ROS)
- Treat cells as in Step 2.
- Incubate with MitoSOX Red reagent (5 μM) for 30 minutes at 37°C, protected from light.
- Wash gently with warm buffer and measure fluorescence intensity by flow cytometry (excitation/emission ~510/580 nm) or fluorescence microscopy.
- An increase in MitoSOX fluorescence indicates elevated mitochondrial superoxide production, which can be a desired outcome for pro-oxidant therapies or an indicator of toxicity for protective therapies.
Visualization Diagrams
Synthesis and Mechanism of CPP-TPP-Nanocarrier
Experimental Workflow for Validation
The Scientist's Toolkit
Table 3: Essential Research Reagents and Materials
| Category / Item | Specifications / Examples | Primary Function in Protocol |
|---|---|---|
| Targeting Moieties | ||
| TPP-NH2 / TPP-Br | Alkyltriphenylphosphonium salts (C3-C12 alkyl chain) [51] | Mitochondrial targeting unit; conjugated to polymers or lipids. |
| CPP Peptides | TAT (GRKKRRQRRRPQ), R8 (RRRRRRRR), or custom sequences with Cys [48] | Enhances cellular uptake; conjugated via thiol-maleimide chemistry. |
| Nanocarrier Components | ||
| Phospholipids | DSPC, DOPE, DSPE [49] [48] | Main structural components of liposomal/nanoparticle systems. |
| PEGylated Lipids | DSPE-PEG2000, DSPE-PEG2000-Maleimide [49] | Provides "stealth" properties and a handle for ligand conjugation. |
| Characterization Dyes | ||
| MitoTracker | Deep Red FM, Green FM, etc. | Standard dye for staining intact mitochondria in colocalization studies. |
| ΔΨm Sensors | JC-1, TMRM, TMRE | Fluorescent probes to measure mitochondrial membrane potential. |
| MitoSOX Red | - | Mitochondria-targeted fluorescent probe for detecting superoxide. |
| Critical Equipment | ||
| Confocal Microscope | With 405, 488, 561, 640 nm laser lines | High-resolution imaging for subcellular colocalization analysis. |
| Dynamic Light Scattering (DLS) Instrument | - | Measures nanoparticle hydrodynamic size and polydispersity (PDI). |
| Extruder | With polycarbonate membranes (50-200 nm) | Produces uniform, small unilamellar vesicles (SUVs) of defined size. |
Mitochondria, the powerhouses of eukaryotic cells, are increasingly recognized as critical targets for therapeutic intervention in diseases ranging from cancer to neurodegenerative disorders [54]. A groundbreaking approach in this field involves the surface functionalization of isolated mitochondria with cell-penetrating peptides (CPPs) to enhance their cellular delivery and therapeutic potential. This application note details a comprehensive workflow, from isolating high-integrity mitochondria to their functionalization with engineered CPPs, providing researchers with a robust framework for mitochondrial transplantation and therapy development. The protocols are framed within a broader research context aiming to leverage CPP-modified mitochondria to restore mitochondrial function in diseased cells, a promising frontier in precision medicine.
Mitochondrial Isolation Protocols
The foundation of successful functionalization is the isolation of intact, functional mitochondria. Below are two detailed protocols offering different advantages.
Standard Isolation from Cultured Fibroblasts via Differential Centrifugation
This protocol, adapted from the Seifert Lab, is a reliable method for obtaining mitochondrial fractions from mammalian cells [55].
Key Materials:
- Mitochondria Isolation Buffer (MIB): 215 mM mannitol, 75 mM sucrose, 0.1% BSA, 20 mM HEPES, 1 mM EGTA, pH 7.2 with KOH. The osmolarity and EGTA are crucial for preserving structural integrity and preventing permeability transition.
- Ficoll Gradient: Prepared with STE buffer (Sucrose, Tris, EDTA) and Ficoll stock to purify the mitochondrial fraction from other organelles.
- Nitrogen Cell Disruptor: Provides a controlled, shear-free method for plasma membrane rupture.
Detailed Workflow:
- Cell Harvesting: Culture fibroblasts to near-confluence. Dislodge cells using trypsin and collect by centrifugation at 1,500 rpm for 5 minutes. Discard the supernatant and keep the cell pellet on ice.
- Cell Washing: Resuspend the cell pellet in 1 mL of ice-cold MIB. Centrifuge at 400 RCF for 5 minutes to wash the cells. Discard the supernatant.
- Cell Disruption: Resuspend the pellet in 450 µL of EDTA. Use a nitrogen cell disruptor at 1,250 psi for 10 minutes to selectively rupture the plasma membrane while leaving mitochondria intact.
- Crude Mitochondria Isolation: Pass the lysate through a 26-gauge syringe topped with 1.5 mL MIB. Centrifuge at 1,300 x g at 4°C for 3 minutes. Repeat this step to remove large debris and unbroken cells.
- Mitochondrial Purification: Transfer the supernatant to a new tube and centrifuge at 1,300 x g at 4°C for 10 minutes to pellet nuclei and other contaminants. Apply the resulting supernatant to a pre-formed Ficoll gradient.
- Gradient Centrifugation: Centrifuge the Ficoll gradient at 3,200 rpm for 30 minutes. This step separates mitochondria from lighter membranous contaminants.
- Final Pellet: Discard the supernatant and resuspend the purified mitochondrial pellet in 1 mL of MIB. Perform a final wash by centrifuging at 13,000 RCF for 10 minutes. The final pellet consists of highly purified, intact mitochondria, ready for functionalization or cryopreservation [55].
Advanced Isolation and Cryopreservation with Improved Integrity
This method, developed by Okutani et al., emphasizes mitochondrial integrity and enables long-term storage, which is vital for practical experimental workflows [56].
Key Innovation: Selective plasma membrane weakening using low-concentration digitonin, under conditions that do not increase mitochondrial membrane permeability. This causes mitochondria to contract within the cell before plasma membrane rupture, facilitating gentler extraction.
Detailed Workflow:
- Plasma Membrane Weakening: Incubate cells with a carefully titrated concentration of digitonin on ice.
- Selective Rupture: Apply gentle mechanical stress to rupture the weakened plasma membrane, releasing the pre-contracted mitochondria.
- Integrity Check: Assess mitochondrial integrity by measuring the polarization of the inner membrane (e.g., using TMRE or JC-1 dyes). This protocol achieves ~90% polarization in the isolated population [56].
- Cryopreservation Protocol:
- Isolate mitochondria using the digitonin method.
- Resuspend in a cryopreservation medium containing appropriate cryoprotectants.
- Rapid Thawing: For thawing, rapidly warm cryovials in a 37°C water bath until the last ice crystal disappears, completing the process in under 1.5 minutes. This rapid thawing is critical, resulting in a decrease of polarized mitochondria by only about 10% post-thaw [56].
Table 1: Quantitative Comparison of Mitochondrial Isolation Methods
| Parameter | Differential Centrifugation [55] | Digitonin-Based Isolation [56] |
|---|---|---|
| Typical Starting Material | Cultured fibroblasts | Various cell cultures |
| Key Reagent | Nitrogen disruptor, Ficoll gradient | Digitonin |
| Outer Membrane Integrity | Standard | Improved |
| Inner Membrane Polarization | Not specified | ~90% |
| Intermembrane Space Protein Retention | Standard | High |
| Cryopreservation Compatibility | Not specified | Excellent (with rapid thawing) |
| Best For | Standard purification & fractionation | Applications requiring maximum functionality |
Functionalization with Cell-Penetrating Peptides (CPPs)
Once isolated, mitochondria can be engineered with CPPs to create "enhanced delivery vehicles." Recent research provides clear design principles for mitochondria-targeting CPPs.
CPP Design Principles for Mitochondrial Targeting
The work of Schmitt and Wennemers establishes that successful mitochondrial targeting requires more than just cellular uptake; it demands specific structural features [32] [57].
- Rigid Polyproline II (PPII) Helical Backbone: A stable secondary structure is essential. Peptides adopting a PPII helix organize their functional groups along defined edges, promoting correct interaction with mitochondrial membranes.
- Amphipathic Patterning: The PPII helix must be engineered with a specific amphipathic pattern: guanidinium groups (from Gup residues) aligned on two edges and hydrophobic residues (e.g., cyclohexylalanine, Cha) on the third edge. This pattern mimics natural mitochondrial-targeting signals.
- Hydrophobicity Tuning: Hydrophobicity is a critical lever for optimizing performance.
- C-Terminal Modification: Adding a hydrophobic Cha residue at the C-terminus of a 9-mer peptide (e.g., creating
(ChaZZ)3-Cha) significantly increases cellular uptake and mitochondrial colocalization [32]. - Backbone Engineering: Incorporating (4S)-cyclohexyl-proline (ChPro) directly into the backbone creates highly hydrophobic peptides with strong membrane association and eventual mitochondrial redistribution over 24 hours.
- The Hydrophobicity Balance: Excessive hydrophobicity leads to endosomal trapping, strong plasma membrane binding, and cytotoxicity. The goal is a subtle increase that promotes mitochondrial localization without off-target accumulation [32] [57].
- C-Terminal Modification: Adding a hydrophobic Cha residue at the C-terminus of a 9-mer peptide (e.g., creating
Table 2: Impact of CPP Sequence Modifications on Mitochondrial Targeting
| CPP Sequence Feature | Effect on Cellular Uptake | Effect on Mitochondrial Localization | Key Findings |
|---|---|---|---|
| Base PPII Helix (e.g., (ChaZZ)3) | Moderate | Yes, at higher concentrations (≥10 µM) | Establishes the minimal requirement for mitochondrial targeting [32]. |
| C-terminal Cha (e.g., (ChaZZ)3-Cha) | Significantly enhanced | Enhanced, with localization at lower concentrations (5 µM) | Increases hydrophobicity, improving uptake and selectivity for mitochondria over the nucleus [32]. |
| ChPro in Backbone | Greatly enhanced | Enhanced, but with time-dependent redistribution from membranes/endosomes | Higher hydrophobicity can cause initial endosomal entrapment; mitochondrial localization improves over 24h [32]. |
| Flexible Arg-based control (e.g., (ChaRR)3) | High | No | Confirms the necessity of the rigid PPII helical structure for mitochondrial targeting, not just amphipathicity [32]. |
Workflow for Mitochondrial Surface Functionalization
The functionalization process involves conjugating the designed CPPs to the outer mitochondrial membrane.
- CPP Synthesis and Purification: Synthesize the desired amphipathic CPP (e.g.,
CF-Ahx-(ChaZZ)3-Cha-NH2) using solid-phase peptide synthesis (SPPS). Purify via Reverse-Phase High-Performance Liquid Chromatography (RP-HPLC) and confirm structure and PPII helicity via Circular Dichroism (CD) spectroscopy [32]. - Mitochondrial Incubation: Resuspend the freshly isolated or thawed mitochondrial pellet in a suitable incubation buffer (e.g., MIB). Incubate with the synthesized CPP at a optimized concentration (e.g., 5-20 µM) for a defined period at 37°C with gentle agitation.
- Removal of Unbound CPP: Purify the CPP-coated mitochondria from excess, unbound peptide using a series of gentle centrifugations and washes with ice-cold MIB.
- Validation of Functionalization:
- Flow Cytometry: Quantify the uptake of fluorescently-labeled CPPs on the mitochondrial surface.
- Confocal Microscopy: Confirm the co-localization of the CPP signal with a mitochondrial-specific dye (e.g., MitoTracker Deep Red) after the functionalized mitochondria are introduced to recipient cells.
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagents for Mitochondrial Isolation and CPP Functionalization
| Reagent / Material | Function / Application | Example / Specification |
|---|---|---|
| Mitochondria Isolation Buffer (MIB) | Maintains osmolarity and integrity during isolation; contains mannitol, sucrose, BSA, HEPES, EGTA [55]. | 215 mM mannitol, 75 mM sucrose, 0.1% BSA, 20 mM HEPES, 1 mM EGTA, pH 7.2 [55]. |
| Digitonin | Selective permeabilization of the cholesterol-rich plasma membrane without damaging mitochondria [56]. | Requires careful concentration titration for different cell types. |
| Ficoll Gradient | Purification of mitochondria by density from other cellular components [55]. | Prepared in STE buffer (Sucrose, Tris, EDTA) [55]. |
| Nitrogen Cell Disruptor | Shear-free, controlled rupture of the plasma membrane to release intact organelles [55]. | Operated at 1,250 psi for 10 minutes [55]. |
| Amphipathic CPPs | Functionalization of isolated mitochondria to enhance delivery to recipient cells. | PPII-helical peptides with sequence (XZZ)n, where Z is Gup and X is a hydrophobic residue like Cha [32] [57]. |
| MitoTracker Deep Red | Fluorescent staining of mitochondria in live cells for imaging and colocalization studies. | Far-red fluorescent dye (ex/em ~644/665 nm), stable in live cells. |
| Chloroquine (CQ) / Bafilomycin A1 (BAF) | Lysosomotropic agents that inhibit lysosomal proteolytic activity; used to track cargo degradation [58]. | Useful for confirming lysosomal involvement in the degradation pathway of internalized cargo. |
Workflow Visualization
The following diagrams summarize the core experimental and conceptual workflows described in this application note.
Diagram 1: Integrated workflow for mitochondrial isolation and CPP functionalization. The CPP design node incorporates critical structural principles for successful mitochondrial targeting [32] [55] [57].
Diagram 2: Post-functionalization cellular pathway of CPP-enhanced mitochondria. The pathway highlights the challenge of endosomal entrapment and a potential degradation route that can be probed with inhibitors like chloroquine (CQ) [58].
Overcoming Technical Challenges: From Cellular Uptake to Functional Integration
Optimizing Hydrophobicity and C-Terminal Modifications for Improved Uptake
The efficacy of cell-penetrating peptides (CPPs) in therapeutic delivery is profoundly influenced by their physicochemical properties. Within the broader context of surface-modifying mitochondria with CPPs for enhanced delivery, fine-tuning hydrophobicity and implementing specific C-terminal modifications have emerged as critical strategies for optimizing cellular uptake and achieving precise subcellular targeting. Mitochondria, being key organelles in cellular metabolism and apoptosis, represent a prime target for treating a wide range of diseases, from neurodegenerative disorders to cancer [32] [59]. However, the dense, hydrophobic proteins and high negative electrochemical gradient of the inner mitochondrial membrane present a formidable barrier [32]. Amphipathic CPPs, which feature distinct cationic and hydrophobic regions, can overcome this barrier. Their cationic moieties facilitate initial electrostatic interactions with the negatively charged mitochondrial membrane, while their hydrophobic segments integrate into the lipid bilayer, promoting translocation. This application note details protocols for systematically modulating these properties, providing a structured approach to enhance the delivery efficiency of CPP-based mitochondrial carriers.
Quantitative Data: Correlating Modifications with Functional Outcomes
Strategic modifications at the C-terminus and along the peptide backbone directly impact key performance parameters. The data below summarize findings from systematic studies on how hydrophobicity and specific C-terminal changes influence peptide behavior and function.
Table 1: Impact of Hydrophobic Residues on Peptide Properties and Localization
| Hydrophobic Residue | Relative Hydrophobicity (RP-HPLC) | Cellular Uptake | Mitochondrial Localization | Key Findings |
|---|---|---|---|---|
| Valine (Val) | Lowest | Low (minimal at 20 µM) | No | Insufficient hydrophobicity for significant uptake. |
| Phenylalanine (Phe) | Moderate | Yes | No | Localizes in cytosol and nucleoli. |
| Tryptophan (Trp) | High | Yes | No | Localizes in cytosol and nucleoli. |
| Cyclohexylalanine (Cha) | Highest | High (at 10 µM) | Yes | Key for mitochondrial targeting; requires PPII helix structure. |
Table 2: Effects of C-Terminal and Backbone Hydrophobicity Modifications
| Modification Type | Peptide Example | Impact on Hydrophobicity | Cellular Uptake & Mitochondrial Localization | Notes & Considerations |
|---|---|---|---|---|
| Additional C-Terminal Cha | CF-Ahx-(ChaZZ)₃-Cha-NH₂ | Significantly increased (↑ retention time ~3 min) | Enhanced uptake (visible at 5 µM); strong mitochondrial localization. | Can increase endosomal entrapment. |
| Short Peptide + C-Terminal Cha | CF-Ahx-(ChaZZ)₂-Cha-NH₂ (7-mer) | Increased vs. non-C-terminal Cha 6-mer | Strong mitochondrial colocalization at 20 µM; limited endosomal entrapment. | Highlights role of hydrophobicity in shorter sequences. |
| Cyclohexyl at Cγ (ChPro) | CF-Ahx-(ChProZZ)₃-NH₂ | Higher than Cha analogs | Modulated uptake and targeting; increased PPII helix propensity. | All-tertiary amide bonds increase backbone hydrophobicity. |
| N-Terminal Aromatic/Dabcyl | Dabcyl-AMBA-(Arg)ₙ [60] | Increased | Dramatically enhanced cellular uptake for short oligoarginines. | Synergistic effect of Dabcyl and aromatic AMBA group. |
Experimental Protocols
Protocol 1: Solid-Phase Peptide Synthesis (SPPS) and C-Terminal Modification
This protocol describes the synthesis of amphipathic oligoprolines and the introduction of C-terminal hydrophobic residues.
- Key Reagents: Rink amide MBHA resin, Fmoc-protected amino acids (including Fmoc-Gup(OH) and Fmoc-Cha-OH), Fmoc-AMBA, 4-((4-(dimethylamino)phenyl)azo)benzoic acid (Dabcyl), coupling reagents (DIC, OxymaPure), cleavage cocktail (TFA, phenol, water, thioanisole, 1,2-ethanedithiol) [32] [60].
- Procedure:
- Resin Preparation: Load the synthesizer with Rink amide MBHA resin (0.1 mmol scale). Swell the resin in DCM for 30 minutes, then in DMF.
- Fmoc Deprotection: Treat the resin with a deprotection solution (2% piperidine, 2% DBU in DMF) in cycles of 2, 2, 5, and 10 minutes. Wash extensively with DMF (8 x 1 min) after each deprotection [60].
- Chain Elongation:
- For a sequence like (ChaZZ)₃, couple Fmoc-Cha-OH, then Fmoc-Gup(OH) twice, and repeat this pattern.
- For each coupling, use a 3x molar excess of Fmoc-amino acid, DIC, and OxymaPure in DMF. React for 60 minutes at room temperature.
- After coupling, wash with DMF (2 x 1 min) and DCM (3 x 1 min). Perform a Kaiser test to confirm coupling completion [60].
- C-Terminal Modification:
- For the standard C-terminal amide, proceed to final Fmoc deprotection after the last amino acid.
- For an additional C-terminal Cha, after incorporating the final Cha, proceed to final Fmoc deprotection.
- Conjugation with Functional Groups:
- For fluorophore labeling (e.g., with 5(6)-carboxyfluorescein, CF), couple CF to the N-terminus of the peptide on-resin using DIC/OxymaPure [32] [60].
- For Dabcyl modification, couple Dabcyl (2 eq) to the N-terminus or the ε-amino group of a C-terminal lysine using DIC/OxymaPure in DMF with DIEA, reacting overnight [60].
- Cleavage and Deprotection: Cleave the peptide from the resin using a TFA-based cocktail (e.g., 94% TFA, 2.5% water, 2.5% thioanisole, 1% phenol) for 2-4 hours. Precipitate the crude peptide in cold diethyl ether.
- Purification and Analysis: Purify the peptide by semi-preparative RP-HPLC. Characterize the final product using analytical RP-HPLC and ESI-MS [32] [60].
Protocol 2: Assessing Hydrophobicity and Secondary Structure
- Key Reagents: Purified peptides, HPLC-grade water, acetonitrile, trifluoroacetic acid (TFA), phosphate buffer (for CD).
- Procedure:
- Reverse-Phase HPLC (RP-HPLC) Analysis:
- Column: C4 column.
- Mobile Phases: A: H₂O/MeCN/TFA (1000:10:1); B: MeCN.
- Gradient: 30% to 55% B over 20 minutes.
- Analysis: Measure the retention time of each peptide. A longer retention time indicates greater hydrophobicity [32].
- Circular Dichroism (CD) Spectroscopy:
- Prepare a peptide solution in a compatible buffer (e.g., 10 mM phosphate buffer) at a concentration of 0.1-0.2 mg/mL.
- Record spectra in the far-UV range (190-250 nm) using a quartz cuvette with a 1 mm path length.
- Data Interpretation: A characteristic PPII helix shows a minimum at ~206 nm and a maximum at ~224 nm. Compare the intensity and shape of spectra to confirm the secondary structure is maintained after modification [32].
- Reverse-Phase HPLC (RP-HPLC) Analysis:
Protocol 3: Cellular Uptake and Mitochondrial Colocalization Assay
- Key Reagents: MCF-7 cells (or other relevant cell lines), cell culture media, peptides, Hoechst 33342, MitoTracker Deep Red, serum-free medium, confocal microscope.
- Procedure:
- Cell Preparation: Plate MCF-7 cells (or EBC-1 cells for flow cytometry) on glass-bottom dishes at a density of 10⁵ cells per well. Incubate for 24 hours at 37°C to achieve 70-80% confluence.
- Peptide Treatment: Incubate cells with the peptide (e.g., at 5, 10, and 20 µM) in serum-free medium for 1-1.5 hours at 37°C [32] [60].
- Staining:
- For mitochondrial staining, add MitoTracker Deep Red (according to manufacturer's instructions) during the last 30 minutes of peptide incubation.
- For nuclear staining, add Hoechst 33342 (1-2 µg/mL) for the final 10-15 minutes.
- Washing and Imaging: Remove the peptide solution and wash the cells 3x with PBS. Image immediately using a confocal microscope.
- Excitation/Emission: CF (~492/~517 nm), MitoTracker Deep Red (~644/~665 nm), Hoechst (~350/~461 nm).
- Image Analysis: Use image analysis software (e.g., ImageJ) to quantify fluorescence intensity and calculate Pearson's correlation coefficient for peptide and MitoTracker signals to assess mitochondrial colocalization.
- (Optional) Flow Cytometry: For quantitative uptake, after peptide incubation and trypsinization to remove membrane-bound peptides, analyze cell-associated fluorescence using flow cytometry [60].
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for CPP Development and Mitochondrial Targeting Studies
| Reagent / Tool | Function / Application | Example in Context |
|---|---|---|
| Fmoc-Gup(OH) | Building block for introducing guanidinium groups directly on the proline backbone, creating cationic edges. | Used in (XZZ)₃ sequences to form the rigid, cationic PPII helix [32]. |
| Fmoc-Cha-OH | Incorporates a highly hydrophobic cyclohexylalanine side chain to enhance membrane interaction. | Key residue in mitochondrial-targeting peptides like (ChaZZ)₃ [32]. |
| Dabcyl Group | Acts as an internalization enhancer when conjugated to the N-terminus, dramatically boosting uptake. | Modification of oligoarginines like Dabcyl-Arg₆ significantly improved cellular uptake [60]. |
| 4-(Aminomethyl)benzoic Acid (AMBA) | Aromatic unnatural amino acid that synergizes with Dabcyl to further enhance cellular internalization. | Used in Dabcyl-AMBA-Argₙ constructs, which were more effective than octaarginine [60]. |
| 5(6)-Carboxyfluorescein (CF) | Fluorescent dye for labeling peptides to enable tracking of cellular uptake and localization via microscopy/flow cytometry. | Conjugated to peptides via an Ahx spacer for intracellular visualization [32] [60]. |
| MitoTracker Deep Red | Far-red fluorescent dye that accumulates in active mitochondria, used as a counterstain for colocalization studies. | Standard dye for confirming mitochondrial targeting of CPPs in confocal microscopy [32]. |
| Cell-Penetrating Peptides (CPPs) | Facilitate the cellular uptake of cargo molecules. Can be engineered for specific organelle targeting. | HIV-1 TAT peptide and Pep-1 have been used to facilitate mitochondrial uptake and release [59]. |
Visualizing the Workflow and Mechanism
The following diagrams illustrate the logical workflow for optimizing CPPs and the mechanism by which engineered peptides target mitochondria.
Diagram 1: CPP Optimization Workflow - This flowchart outlines the iterative process of designing, synthesizing, characterizing, and testing CPPs to achieve optimal mitochondrial targeting.
Diagram 2: Mitochondrial Targeting Mechanism - This diagram depicts the multi-step mechanism by which an engineered amphipathic CPP targets and translocates across the mitochondrial membrane.
Preventing Endosomal Entrapment and Ensuring Cytosolic Release
The successful delivery of therapeutics to subcellular organelles, such as mitochondria, represents a significant challenge in targeted drug development. A primary bottleneck in this process is endosomal entrapment, where cargo becomes sequestered within endosomes and is ultimately degraded in lysosomes rather than reaching its intended cytosolic or organellar destination. For mitochondrial-targeted therapies using cell-penetrating peptides (CPPs), achieving efficient endosomal escape is critical for therapeutic efficacy. This application note examines validated experimental approaches to prevent endosomal entrapment and promote cytosolic release within the specific context of mitochondrial delivery systems, providing researchers with practical methodologies to enhance their intracellular delivery platforms.
Mechanisms and Chemical Promoters of Endosomal Escape
Understanding Endosomal Entrapment and Escape Pathways
Cell-penetrating peptides primarily enter cells via endocytosis, but subsequently remain trapped within endosomes and lysosomes without intervention [29]. The degradative environment of these compartments destroys biological cargoes before they can reach mitochondrial targets. Effective endosomal escape strategies work by disrupting the endosomal membrane through various mechanisms, including the proton sponge effect, membrane pore formation, or lipid bilayer disruption.
The small molecule L-Leucyl-L-Leucine methyl ester (LLOMe) has been identified as a potent endosomal escape promoter that functions by permeabilizing endosomal and lysosomal membranes [29]. This property makes it a valuable experimental tool for studying and enhancing the cytosolic delivery of CPP-conjugated cargoes destined for mitochondrial localization.
Experimental Optimization of Endosomal Escape Assessment
Accurate assessment of endosomal escape requires careful consideration of multiple experimental parameters that can significantly influence results:
- Amino acid chirality: CPPs constructed with D-amino acids demonstrate enhanced resistance to proteolytic degradation within endolysosomal compartments, potentially increasing the window for successful escape [29].
- Fluorophore selection: The choice of fluorophore conjugated to CPPs critically affects behavior and interpretation, as some fluorophores may be released from CPPs in the degradative environment of endosomes and lysosomes, creating false positives for cytosolic localization [29].
- Temporal monitoring: Endosomal escape is time-dependent, requiring measurements at multiple time points to accurately capture the dynamics of vesicular release and mitochondrial accumulation [29] [32].
Table 1: Key Parameters for Assessing Endosomal Escape
| Parameter | Impact on Endosomal Escape Assessment | Optimization Guidelines |
|---|---|---|
| Amino Acid Chirality | D-amino acids enhance metabolic stability | Use D-amino acids in CPP sequence to prolong functional availability for escape |
| Fluorophore Choice | Affects CPP behavior and signal interpretation | Select fluorophores with stable CPP linkages; verify signal correlates with cargo location |
| Timing of Recording | Escape is a dynamic, time-dependent process | Conduct measurements at multiple time points (e.g., 1h, 4h, 24h) to capture escape kinetics |
| Escape Promoters | Chemicals like LLOMe artificially enhance escape | Use as positive controls; titrate concentration to balance efficacy with cytotoxicity |
Mitochondrial-Targeting CPP Design Strategies
Structural Features for Enhanced Mitochondrial Targeting
Beyond overcoming endosomal barriers, effective mitochondrial delivery requires specific structural features that promote organelle-specific targeting:
- Amphipathic patterning: CPPs with rigid polyproline II (PPII) helical backbones displaying aligned cationic and hydrophobic residues along separate edges demonstrate superior mitochondrial targeting compared to flexible analogs [32]. This structural organization appears critical for mitochondrial interaction and uptake.
- Hydrophobicity optimization: Systematic modulation of peptide hydrophobicity through C-terminal modifications or backbone engineering significantly enhances both cellular uptake and mitochondrial selectivity [32]. For example, adding a C-terminal cyclohexylalanine (Cha) residue to amphipathic oligoprolines increased mitochondrial localization while maintaining PPII helicity.
- Cationic character: The presence of guanidinium groups in periodic arrangements along the peptide backbone facilitates interaction with the negatively charged mitochondrial membrane, a key step in mitochondrial targeting [32].
Comparative Performance of Mitochondrial-Targeting Moieties
Table 2: Mitochondrial Targeting Strategies and Their Applications
| Targeting Moiety | Mechanism of Action | Experimental Applications | Advantages/Limitations |
|---|---|---|---|
| Triphenylphosphonium (TPP) | Utilizes mitochondrial membrane potential (-160 to -180 mV) for electrophoretic accumulation [49] [4] | Conjugated to nanocarriers, antioxidants, and probes for mitochondrial delivery [49] | Advantages: Well-characterized, simple conjugation chemistryLimitations: Can cause proton leakage at high concentrations, limited hydrophilicity [4] |
| Amphipathic Proline-Rich Peptides | PPII helix with aligned cationic/hydrophobic edges interacts with mitochondrial membranes [32] | Mitochondrial delivery of therapeutics, molecular imaging | Advantages: Rigid structure minimizes endosomal entrapment, tunable hydrophobicityLimitations: More complex synthesis, sequence optimization required |
| Carbon Quantum Dots | Intrinsic mitochondrial targeting without additional ligands [61] | Mitochondrial imaging, photodynamic therapy | Advantages: Excellent photostability, low cytotoxicity, facile synthesisLimitations: Limited drug loading capacity, relatively new technology |
| Mitochondrial Presequences | Natural protein targeting signals recognized by TOM/TIM complexes [4] | Delivery of protein cargoes to mitochondria | Advantages: High specificity, biological relevanceLimitations: Large size, potential immunogenicity |
Experimental Protocols for Evaluating Endosomal Escape and Mitochondrial Delivery
Protocol: Quantitative Assessment of Endosomal Escape Using LLOMe
Purpose: To measure and enhance the endosomal escape efficiency of mitochondrial-targeted CPPs using the chemical promoter LLOMe.
Materials:
- CPPs conjugated to appropriate fluorophores (e.g., 5(6)-Carboxyfluorescein)
- L-Leucyl-L-Leucine methyl ester (LLOMe)
- Cell culture appropriate for experiment (e.g., MCF-7 cells)
- Confocal microscopy with live-cell imaging capabilities
- Mitochondrial stain (e.g., MitoTracker Deep Red)
- Nuclear stain (e.g., Hoechst 33342)
Procedure:
- Culture cells on glass-bottom confocal dishes until 60-70% confluent.
- Pre-treat cells with LLOMe (typical concentration range: 10-100 μM) for 30 minutes in serum-free medium [29].
- Incubate cells with CPP-fluorophore conjugates (5-20 μM) for 1 hour at 37°C.
- Replace with fresh medium to remove extracellular peptides.
- Counterstain with MitoTracker Deep Red (according to manufacturer protocol) and Hoechst 33342.
- Image using confocal microscopy with appropriate filter sets.
- Analyze images for:
- Punctate vs. diffuse fluorescence pattern (punctate indicates endosomal entrapment)
- Colocalization coefficients with mitochondrial markers
- Cytosolic fluorescence intensity
Interpretation: Successful endosomal escape is indicated by a shift from punctate to diffuse fluorescence pattern and increased colocalization with mitochondrial markers. LLOMe-treated samples should show enhanced mitochondrial localization compared to untreated controls.
Protocol: Evaluating Mitochondrial Targeting Efficiency of Amphipathic CPPs
Purpose: To design, synthesize, and validate mitochondrial-targeting CPPs with optimized endosomal escape properties.
Materials:
- Solid-phase peptide synthesis equipment and reagents
- Amino acids including (4S)-guanidiniumproline (Gup) and cyclohexylalanine (Cha)
- 5(6)-Carboxyfluorescein (CF) for labeling
- Reverse-phase HPLC system with C4 column
- Circular dichroism (CD) spectrometer
- Cell culture and confocal microscopy setup
Procedure: Peptide Design and Synthesis:
- Design peptides using the pattern (XZZ)n where Z = Gup and X = hydrophobic amino acids (e.g., Cha, Phe, Trp).
- Synthesize peptides using solid-phase peptide synthesis with CF label connected via aminohexanoic acid (Ahx) spacer [32].
- Purify using RP-HPLC and characterize by mass spectrometry.
Structural Confirmation:
- Analyze secondary structure by circular dichroism (CD).
- Confirm characteristic PPII helix with minimum at ~206 nm and maximum at ~224 nm [32].
- Determine relative hydrophobicity using RP-HPLC retention times.
Cellular Uptake and Localization:
- Incubate MCF-7 cells with peptides (5-20 μM) for 1 hour at 37°C.
- Counterstain with MitoTracker Deep Red and Hoechst 33342.
- Image using confocal microscopy with appropriate filter sets.
- Quantify mitochondrial colocalization using Pearson's correlation coefficient.
Interpretation: Successful mitochondrial targeting peptides will show high Pearson's correlation with MitoTracker signal, while maintaining minimal endosomal punctate staining. Peptides with optimized hydrophobicity (e.g., CF-Ahx-(ChaZZ)3-Cha-NH2) should demonstrate enhanced mitochondrial localization compared to less hydrophobic variants.
The Scientist's Toolkit: Essential Reagents for Mitochondrial Delivery Research
Table 3: Key Research Reagent Solutions for Endosomal Escape and Mitochondrial Delivery Studies
| Reagent/Category | Specific Examples | Primary Function | Application Notes |
|---|---|---|---|
| Endosomal Escape Promoters | LLOMe (L-Leucyl-L-Leucine methyl ester) | Permeabilizes endosomal/lysosomal membranes [29] | Use as positive control; optimize concentration to balance efficacy and toxicity |
| Cell-Penetrating Peptides | R9, TAT, penetratin, amphipathic oligoprolines [29] [32] | Facilitate cellular uptake and mitochondrial targeting | D-amino acid versions enhance stability; fluorophore choice critical for accurate tracking |
| Mitochondrial Stains | MitoTracker Deep Red | Visualize mitochondria for colocalization studies | Validate new CPP mitochondrial targeting; use fresh solutions for optimal performance |
| Structural Analysis Tools | Circular Dichroism (CD) Spectroscopy | Confirm PPII helical structure of CPPs [32] | Characteristic spectrum: minimum at ~206 nm, maximum at ~224 nm |
| Hydrophobicity Assessment | Reverse-Phase HPLC (C4 column) | Estimate relative hydrophobicity of CPP variants [32] | Longer retention times indicate higher hydrophobicity; correlates with uptake efficiency |
| Imaging Tools | Confocal Microscopy with live-cell capability | Visualize subcellular localization and endosomal escape | Distinguish punctate (endosomal) vs. diffuse (cytosolic) fluorescence patterns |
Visualization of Experimental Workflows
Endosomal Escape Assessment Workflow
Workflow for Assessing Endosomal Escape
Mitochondrial-Targeted CPP Design Logic
Strategies to Overcome Endosomal Entrapment
Preventing endosomal entrapment represents a critical challenge in developing effective mitochondrial-targeted delivery systems. The integrated approaches presented here—combining chemical promoters like LLOMe with structurally optimized amphipathic CPPs—provide researchers with validated methodologies to enhance cytosolic release and mitochondrial delivery. The rigorous assessment protocols, focusing on temporal dynamics, fluorophore selection, and structural characterization, enable accurate evaluation of endosomal escape efficiency. Implementation of these strategies will advance the development of mitochondrial-targeted therapies with improved intracellular bioavailability and therapeutic efficacy.
Mitigating Cytotoxicity and Unwanted Immune Activation
Mitochondrial transplantation has emerged as a promising therapeutic strategy for restoring cellular function in diseases characterized by mitochondrial dysfunction. However, the clinical application of this approach faces significant challenges, including potential cytotoxic responses and unwanted immune activation following the introduction of exogenous mitochondria. Surface engineering of mitochondria with cell-penetrating peptides (CPPs) represents an innovative solution to these challenges, enhancing delivery efficiency while minimizing adverse reactions. This Application Note provides detailed protocols and experimental frameworks for mitigating cytotoxicity and immune activation through mitochondrial surface modification, supporting the broader thesis that CPP-enhanced delivery systems can revolutionize mitochondrial therapeutics.
The imperative for these refined protocols stems from recognized limitations in current mitochondrial transfer and transplantation (MTT) methods. As noted in recent literature, "Injected mitochondria have to cope with an inhospitable extracellular environment" and "non-viable or damaged mitochondria can release damage-associated molecular patterns that lead to activation of the immune system" [11]. Furthermore, studies indicate that only a small proportion (approximately 10%) of injected mitochondria typically reach target cells, and the transfer process lacks specificity [11]. Surface engineering strategies, particularly those utilizing CPPs, directly address these limitations by enhancing targeting precision and cellular uptake while reducing immune recognition.
Key Challenges in Mitochondrial Transplantation
Immune Recognition and Activation
Exogenous mitochondria are susceptible to immune recognition through multiple pathways. Mitochondrial DNA containing unmetabolized CpG motifs can activate Toll-like receptor 9 (TLR9) signaling, while mitochondrial transcription factor A (TFAM) functions as a damage-associated molecular pattern (DAMP) [62]. Additionally, the externalization of cardiolipin on the outer mitochondrial membrane can trigger immune responses [11]. These pathways collectively contribute to unwanted immune activation following mitochondrial transplantation.
Limitations in Uptake Efficiency
Recent quantitative studies reveal that mitochondrial uptake is relatively inefficient, with only 1-2% of administered mitochondria typically internalized by recipient cells under standard conditions [6]. This low efficiency necessitates higher transplantation doses, which in turn increases the risk of immune reactions and cytotoxicity. The primary mechanism of uptake appears to be fluid-phase endocytosis rather than specific receptor-mediated processes, as evidenced by the virtual elimination of uptake at 4°C [6].
Surface Engineering Strategies with CPPs
CPP Selection and Mechanisms
Cell-penetrating peptides facilitate mitochondrial internalization through interactions with cell membranes, bypassing traditional endocytic pathways that often lead to lysosomal degradation. The table below summarizes characterized CPPs for mitochondrial delivery:
Table 1: Cell-Penetrating Peptides for Mitochondrial Delivery
| CPP | Sequence/Characteristics | Mechanism of Action | Efficiency Findings | Applications Demonstrated |
|---|---|---|---|---|
| Pep-1 | Tryptophan-rich, amphipathic | Non-covalent complex formation; endocytosis-independent cellular entry | Significant enhancement compared to cell-free mitochondria [11] | Parkinson's disease models, mitochondrial myopathy models [11] |
| HIV TAT | Arginine-rich, cationic | Covalent coupling; direct membrane translocation | Restoration of complex I activity [11] | Delivery of mitochondrial enzymes [11] |
| DSPE-PEG-Pep | Polymer-peptide conjugate | Enhanced stability and targeting via lipid anchoring | Significantly enhanced uptake in diabetic aortic endothelial cells [36] | Vascular endothelial targeting [36] |
Surface Modification Workflow
The following diagram illustrates the comprehensive workflow for mitochondrial surface modification with CPPs:
Experimental Protocols
Mitochondrial Isolation and Surface Modification
Protocol 1: Mitochondrial Isolation from iPSC-MSCs
- Cell Source: Use induced pluripotent stem cell-derived mesenchymal stem cells (iPSC-MSCs) from healthy donors [36].
- Isolation Procedure:
- Dissociate approximately 15×10⁶ cells using trypsin and resuspend in 800 µL of Buffer A with 10 µL of Mitochondria Isolation Reagent B.
- Incubate on ice for 5 minutes, vortexing every minute.
- Add 800 µL of Reagent C and centrifuge at 700×g for 10 minutes at 4°C.
- Transfer supernatant to a new tube and centrifuge at 3,000×g for 15 minutes at 4°C.
- Resuspend mitochondrial pellet in 500 µL Reagent C and centrifuge at 12,000×g for final purification [36].
- Quality Assessment: Determine protein content using BCA assay and confirm membrane integrity via JC-1 staining for membrane potential.
Protocol 2: DSPE-PEG-Peptide Conjugate Synthesis
- Materials:
- DSPE-PEG-MAL (5,000 MW)
- Biotinylated CPPs (VHPKQHRGGSKGC for VCAM-1 targeting or CQDSETRTFY for collagen targeting) [36]
- Procedure:
- React peptides with DSPE-PEG-MAL at thiol:maleimide molar equivalent in ultrapure water.
- Incubate at room temperature for 24 hours with gentle agitation.
- Purify conjugates by dialysis using Slide-a-Lyzer Cassette (MWCO 7,000 kDa) for 24 hours.
- Lyophilize and confirm conjugation using MALDI mass spectrometry [36].
Protocol 3: Mitochondrial Surface Functionalization
- Coating Procedure:
- Combine freshly isolated mitochondria with DSPE-PEG-peptide conjugate (1 mg/mL in Reagent C) at varying mass ratios (polymer to mitochondrial protein).
- Incubate on ice for 3 hours with continuous shaking.
- Purify by centrifugation at 12,000×g for 5 minutes.
- Replace supernatant with fresh Reagent C and repeat centrifugation.
- Resuspend in Mitochondria Storage Buffer for immediate use or store at -80°C [36].
- Coating Validation: Assess coating efficiency using flow cytometry with Mitotracker-labeled mitochondria and AlexaFluor-488 streptavidin.
Uptake and Internalization Assays
Protocol 4: Quantitative Uptake Measurement
- Cell Model: Use human diabetic aortic endothelial cells (DAECs) for disease-relevant assessment [36].
- Labeling Strategy:
- Pre-label mitochondria with Mitotracker CmxRos (500 nM for 45 minutes at 37°C) before isolation.
- Alternatively, engineer donor cells to express NLuc-HA-OMP25 or COX8a-NLuc-HA constructs for highly sensitive luminescence-based tracking [6].
- Uptake Procedure:
- Incubate recipient cells with engineered mitochondria for 24 hours.
- For luminescence assays: Measure NLuc activity in cell lysates using standard protocols.
- For fluorescence assays: Analyze internalization via flow cytometry or confocal microscopy.
- Include temperature controls (4°C) to confirm energy-dependent uptake [6].
Protocol 5: Intracellular Fate Tracking
- Experimental Setup:
- Culture recipient cells on glass-bottom confocal dishes.
- Incubate with Mitotracker Red-labeled engineered mitochondria for 4-24 hours.
- Stain cells with Lysotracker Green (endosomes/lysosomes) and MitoTracker Green (host mitochondria).
- Perform live-cell imaging or fix for immunostaining.
- Analysis: Quantify colocalization coefficients using ImageJ or similar software with appropriate plugins.
Immune Activation Assessment
Protocol 6: Cytokine Profiling
- Experimental Design:
- Co-culture engineered mitochondria with peripheral blood mononuclear cells (PBMCs) or macrophage cell lines.
- Use unmodified mitochondria and lipopolysaccharide as controls.
- Collect supernatants at 6, 24, and 48 hours.
- Analysis: Measure pro-inflammatory cytokines (IL-1β, IL-6, TNF-α, IFN-γ) using ELISA or multiplex immunoassays.
Protocol 7: Immune Cell Activation Markers
- Procedure:
- Isolate primary immune cells (monocytes, T cells) from human blood.
- Treat with engineered mitochondria for 24 hours.
- Stain for activation markers (CD69, CD25, CD86, HLA-DR).
- Analyze via flow cytometry with appropriate isotype controls.
Data Analysis and Validation
Quantitative Uptake and Function Metrics
Table 2: Key Performance Metrics for CPP-Engineered Mitochondria
| Parameter | Assessment Method | Expected Outcome with CPP | Significance |
|---|---|---|---|
| Uptake Efficiency | Flow cytometry, luminescence assays | 2-4 fold increase vs. unmodified [36] [6] | Determines therapeutic dosage |
| Colocalization with Host Network | Confocal microscopy, Manders' coefficients | >60% colocalization after 24h [36] | Induces functional integration |
| Membrane Potential | JC-1 staining (red/green ratio) | Maintained >80% of initial potential [36] | Preserves bioenergetic capacity |
| Oxygen Consumption | Seahorse metabolic analyzer | Sustained basal and maximal respiration [36] | Confirms functional integration |
| Inflammatory Cytokines | ELISA/multiplex assays | >50% reduction vs. unmodified [11] | Demonstrates immune evasion |
Mitochondrial Uptake and Intracellular Trafficking Pathway
The intracellular journey of engineered mitochondria involves specific pathways that can be modulated by surface modifications:
The Scientist's Toolkit
Table 3: Essential Research Reagent Solutions for Mitochondrial Engineering
| Reagent/Category | Specific Examples | Function/Application | Protocol References |
|---|---|---|---|
| Mitochondrial Isolation | Mitochondria Isolation Kit (Thermo Fisher); Buffer A, B, C reagents | Maintain structural integrity and function during isolation | Protocol 1 [36] |
| CPP Conjugates | DSPE-PEG-MAL; Biotinylated VBP, CBP, TAT, Pep-1 peptides | Enable mitochondrial surface functionalization and targeting | Protocol 2 [36] [11] |
| Tracking & Labeling | MitoTracker CmxRos/Green; JC-1 dye; NLuc-HA-OMP25/COX8a-NLuc-HA constructs | Visualize and quantify uptake, localization, and membrane potential | Protocol 4 [36] [6] |
| Uptake Inhibition | Heparin; sodium chlorate; 4°C temperature block | Characterize uptake mechanisms (heparan sulfate dependence) | [6] |
| Functional Assays | Seahorse XF Analyzer kits; ATP luminescence assays; OROBOROS O2k | Quantify bioenergetic functional improvement | [36] |
| Immune Profiling | ELISA kits for IL-1β, IL-6, TNF-α; flow cytometry antibodies for CD69, CD86, HLA-DR | Assess immune activation and inflammatory responses | Protocol 6 & 7 |
Troubleshooting and Optimization
Common Technical Challenges
- Low Coating Efficiency: Optimize mass ratio of polymer to mitochondrial protein through systematic titration (0.5:1 to 5:1). Confirm conjugation efficiency using MALDI mass spectrometry before mitochondrial coating [36].
- Reduced Membrane Potential: Limit processing time and maintain cold temperatures during isolation. Assess immediately after preparation using JC-1 staining [11].
- High Batch Variability: Standardize iPSC-MSC differentiation protocols and use consistent cell passage numbers (recommended P4-P8) for mitochondrial donors [36].
- Aggregation Issues: Sonicate mitochondrial suspensions briefly (5-10 seconds at low amplitude) before surface modification to disrupt aggregates.
The surface engineering of mitochondria with CPPs represents a significant advancement in mitochondrial transplantation therapeutics, directly addressing the dual challenges of cytotoxicity and unwanted immune activation. The protocols detailed in this Application Note provide researchers with standardized methodologies for developing enhanced mitochondrial delivery systems with improved uptake efficiency and reduced immunogenicity. As the field progresses, these approaches will facilitate the clinical translation of mitochondrial transplantation for a range of diseases characterized by mitochondrial dysfunction, from neurodegenerative disorders to cardiovascular diseases. The integration of surface engineering with mitochondrial biology holds particular promise for developing next-generation cellular therapies that can restore bioenergetic capacity without triggering detrimental immune responses.
Strategies for Prolonging Mitochondrial Lifespan and Stability Ex Vivo
The therapeutic potential of mitochondrial transfer and transplantation (MTT) has garnered significant attention for treating diseases linked to mitochondrial dysfunction, such as neurodegenerative and cardiovascular disorders [3]. A critical bottleneck in the clinical translation of this technology is the rapid decline in mitochondrial function once isolated from their cellular environment. Isolated mitochondria significantly lose respiratory function after approximately 2 hours, and they must cope with an inhospitable extracellular milieu characterized by high calcium concentration and reactive oxygen species (ROS) [3]. This application note details standardized protocols, developed within the broader research context of enhancing mitochondrial delivery via surface modification with Cell-Penetrating Peptides (CPPs), to effectively prolong mitochondrial viability and stability ex vivo.
Key Challenges in Ex Vivo Mitochondrial Stability
The following table summarizes the primary obstacles to maintaining functional mitochondria outside the cell and the consequent implications for therapeutic applications.
Table 1: Key Challenges in Maintaining Mitochondrial Stability Ex Vivo
| Challenge | Impact on Isolated Mitochondria | Therapeutic Consequence |
|---|---|---|
| Short Lifespan | Significant loss of respiratory function after ~2 hours [3]. | Narrow window for administration, limiting practical application. |
| Extracellular Stress | Susceptibility to high Ca²⁺ and ROS in the extracellular environment [3]. | Rapid functional decline before reaching target cells. |
| Structural Instability | Tendency to undergo aggregation, swelling, and structural changes [3]. | Loss of membrane potential and integrity, rendering mitochondria ineffective. |
| Immune Recognition | Potential to be recognized as foreign entities, triggering immune responses [3]. | Reduced therapeutic efficacy and potential for adverse inflammatory reactions. |
| Inefficient Cellular Uptake | Only a small proportion (~10%) of injected mitochondria reach target cells [3]. | Low dose delivery to the site of pathology, requiring high initial quantities. |
Core Strategies and Quantitative Comparison
Biotechnological strategies focusing on surface engineering and encapsulation have been developed to address these challenges. The primary goal is to create a protective microenvironment that shields mitochondria from degradation and immune detection while enhancing their uptake by target cells.
Table 2: Biotechnological Strategies for Enhancing Mitochondrial Stability and Delivery
| Strategy | Mechanism of Action | Reported Efficacy | Key Advantages |
|---|---|---|---|
| CPP Conjugation (Pep-1) | Direct fusion with mitochondrial membrane, enhancing cellular internalization [3]. | 60.5% of cells received mitochondria vs. 14.5% with naked mitochondria [3]. | Simple conjugation; significantly boosts cellular uptake. |
| Polymer Coating (TAT-dextran) | Forms a protective hydrophilic layer on mitochondrial surface, preventing aggregation [3]. | 182.8% increase in transfer efficiency over free mitochondria [3]. | Enhances stability in physiological buffers; improves uptake. |
| Ligand Targeting (PEP-TPP complex) | Uses mitochondrial targeting signal (PEP) and triphenylphosphonium (TPP) for directed delivery [3]. | Enabled therapeutic effect via intravenous injection where free mitochondria failed [3]. | Increases specificity and delivery efficiency to target tissues. |
| Extracellular Vesicle (EV) Encapsulation | Natural lipid bilayer shields mitochondria from immune system and degradation [3]. | Data not provided in search results; based on mechanism. | High biocompatibility; leverages natural delivery pathways. |
| Nanomotorized Systems | Propulsion system enhances mobility and tissue penetration. | 7.9% of system reached cardiac tissue vs. 1.0% for free mitochondria [3]. | Actively penetrates tissues, overcoming diffusion barriers. |
Detailed Experimental Protocols
Protocol 1: Mitochondrial Isolation and Baseline Quality Assessment
This protocol outlines the standard procedure for obtaining functional mitochondria from tissue, which serves as the starting point for all subsequent stabilization procedures.
Reagents and Materials:
- Isolation Buffer: 250 mM sucrose, 10 mM HEPES, 1 mM EGTA, pH 7.4 (pre-cooled to 4°C).
- Protease Inhibitor Cocktail: Added to isolation buffer immediately before use.
- Dounce Homogenizer with loose and tight-fitting pestles.
- Centrifuges: Refrigerated benchtop and high-speed centrifuges.
- BCA Protein Assay Kit: For quantifying mitochondrial yield.
Procedure:
- Tissue Homogenization: Minced tissue (e.g., liver) is placed in 10 mL of ice-cold Isolation Buffer and homogenized with 10-15 strokes using a loose-fitting Dounce pestle.
- Differential Centrifugation:
- Centrifuge the homogenate at 1,000 × g for 10 min at 4°C to remove nuclei and unbroken cells.
- Transfer the supernatant to a new tube and centrifuge at 12,000 × g for 15 min at 4°C to pellet the mitochondrial fraction.
- Washing: Gently resuspend the mitochondrial pellet in 10 mL of fresh Isolation Buffer and repeat the high-speed centrifugation (12,000 × g for 15 min).
- Resuspension: Finally, resuspend the purified mitochondrial pellet in a suitable storage or experimental buffer (e.g., MAS buffer). Keep on ice at all times.
- Quality Assessment:
- Protein Yield: Determine mitochondrial concentration using a BCA assay.
- Membrane Potential: Assess using JC-1 or TMRE dyes via fluorescence measurement. A high red/green (JC-1) or high TMRE fluorescence indicates a healthy membrane potential.
- Oxygen Consumption Rate (OCR): Measure using a Clark-type oxygen electrode or Seahorse Analyzer in states of basal respiration, ADP-stimulated (State 3), and oligomycin-inhibited (State 4).
Protocol 2: Surface Functionalization of Mitochondria with TAT-Dextran
This protocol describes a method to coat mitochondria with a cell-penetrating peptide (CPP)-polymer conjugate to enhance stability and cellular uptake.
Reagents and Materials:
- TAT-Dextran Conjugate: Synthesized by conjugating the TAT peptide (YGRKKRRQRRR) to amino-dextran (e.g., 10 kDa) via NHS-ester chemistry.
- Functionalization Buffer: 250 mM sucrose, 10 mM HEPES, pH 7.4.
- Purification Columns: Zeba Spin Desalting Columns (7K MWCO).
Procedure:
- Mitochondrial Preparation: Isolate mitochondria as described in Protocol 1 and resuspend in Functionalization Buffer to a final concentration of 2 mg/mL protein.
- Conjugation Reaction: Add the TAT-Dextran conjugate to the mitochondrial suspension at a 1:10 (w/w) ratio (TAT-Dextran:Mitochondrial protein).
- Incubation: Incubate the mixture for 30 minutes at 4°C with gentle rotation. The reaction is performed at a cold temperature to preserve mitochondrial function while allowing the TAT peptide to interact with the membrane.
- Purification: Load the reaction mixture onto a pre-equilibrated desalting column. Centrifuge at 1,000 × g for 2 minutes to separate the TAT-Dextran-coated mitochondria from the unbound conjugate.
- Quality Control: The purified, coated mitochondria can be assessed for:
- Colloidal Stability: Monitor aggregation by dynamic light scattering (DLS) over 4 hours.
- Membrane Potential: Compare to uncoated controls using TMRE fluorescence.
- In Vitro Uptake: Co-culture with recipient cells (e.g., cardiomyocytes) and quantify transfer efficiency via flow cytometry or confocal microscopy.
Diagram 1: Mitochondrial Surface Functionalization Workflow.
Assessment of Functional Longevity
Rigorous assessment is crucial for validating the success of any stabilization strategy. The following parameters should be monitored over a time course (e.g., 0, 2, 4, 6 hours) while storing mitochondria in a simulated physiological buffer at room temperature or 37°C.
Table 3: Key Parameters for Assessing Ex Vivo Mitochondrial Longevity
| Parameter | Assessment Method | Performance Metric | Target for Viable Mitochondria |
|---|---|---|---|
| Membrane Integrity | JC-1 assay (flow cytometry) | Ratio of red (aggregate) to green (monomer) fluorescence. | High Red/Green ratio (>80% of time-zero value). |
| Respiratory Function | Clark-type oxygen electrode | Oxygen Consumption Rate (OCR) in State 3 (ADP-stimulated). | OCR > 60% of initial rate after 4 hours. |
| Structural Stability | Dynamic Light Scattering (DLS) | Hydrodynamic diameter and polydispersity index (PDI). | Consistent size; PDI < 0.2 indicates low aggregation. |
| Membrane Potential | Tetramethylrhodamine ethyl ester (TMRE) fluorescence | Fluorescence intensity measured via plate reader or flow cytometry. | High fluorescence intensity (>70% of time-zero value). |
| ATP Production | Luciferase-based ATP assay | Luminescence output proportional to ATP concentration. | ATP output > 50% of initial level after 4 hours. |
The Scientist's Toolkit: Essential Research Reagents
Table 4: Key Reagents for Mitochondrial Stabilization and Functionalization Experiments
| Reagent / Material | Function / Purpose | Example / Notes |
|---|---|---|
| TAT Peptide | Cell-Penetrating Peptide (CPP) that enhances cellular uptake of conjugated mitochondria [3]. | Sequence: YGRKKRRQRRR; can be conjugated to dextran for a "shielding" effect. |
| Dextran Polymer | Provides a hydrophilic protective coating, reducing mitochondrial aggregation and immune recognition [3]. | 10 kDa amino-dextran is commonly used as a backbone for TAT conjugation. |
| Triphenylphosphonium (TPP) | A hydrophobic cation that drives accumulation in mitochondria due to the negative mitochondrial membrane potential [4]. | Used in conjunction with targeting peptides (e.g., PEP) for specific delivery. |
| JC-1 Dye | Fluorescent probe for assessing mitochondrial membrane potential. | Emits red fluorescence in healthy mitochondria (high potential) and green in depolarized mitochondria. |
| Zeba Spin Columns | Rapidly desalts and purifies functionalized mitochondria from excess, unbound reagents. | Critical for removing free TAT-Dextran after the conjugation reaction. |
| Protease Inhibitor Cocktail | Prevents proteolytic degradation of mitochondrial surface proteins during isolation and functionalization. | Added fresh to all isolation and storage buffers. |
Integrated Workflow from Isolation to Application
The entire process, from isolating mitochondria to applying them in a therapeutic context, requires a seamless, integrated workflow to maximize the chances of in vivo success. The following diagram illustrates this comprehensive pipeline and the logical decision points within it.
Diagram 2: Integrated R&D Pipeline for Mitochondrial Therapeutics.
The strategies outlined herein—particularly surface functionalization with CPP-polymer conjugates like TAT-dextran—provide a robust framework for prolonging the functional lifespan of mitochondria ex vivo. The provided protocols for isolation, quality control, and stabilization are designed to be integrated into a broader research program focused on developing mitochondrial therapies. By systematically implementing these strategies and rigorously assessing the key parameters of mitochondrial health, researchers can significantly improve the yield, stability, and efficacy of mitochondria for therapeutic transplantation, thereby accelerating the path toward clinical application.
The therapeutic transplantation of functional mitochondria into damaged cells represents a promising strategy for treating a wide range of diseases associated with mitochondrial dysfunction, including cardiovascular disorders, neurodegenerative conditions, and metabolic syndromes [59]. Despite this potential, a significant challenge hindering clinical translation is the lack of target specificity in current delivery approaches, leading to off-target organelle localization and suboptimal therapeutic outcomes [43] [63].
The surface modification of mitochondria with cell-penetrating peptides (CPPs) has emerged as a powerful strategy to enhance delivery precision. However, the inherent physicochemical properties of CPPs—such as their charge distribution, hydrophobicity, and structural conformation—can inadvertently promote accumulation in non-target organelles like endosomes, nucleoli, and lysosomes [32] [21]. This application note details specific protocols and design principles to engineer mitochondrial specificity, thereby minimizing off-target effects and maximizing bioenergetic restoration in recipient cells.
Quantitative Analysis of CPP Properties and Mitochondrial Targeting Efficiency
The design of CPPs for specific mitochondrial targeting requires careful consideration of multiple physicochemical parameters. The following table summarizes key quantitative findings from recent studies on CPP-mitochondria interactions.
Table 1: Quantitative Analysis of CPP Properties and Mitochondrial Targeting Efficiency
| CPP Sequence / Strategy | Hydrophobicity (Retention Time) | Cellular Uptake Efficiency | Mitochondrial Colocalization | Key Structural Features |
|---|---|---|---|---|
| (ChaZZ)₃ [32] | ~18.5 minutes | Moderate (at 10-20 μM) | Moderate | PPII helix with aligned cationic/hydrophobic edges |
| (ChaZZ)₃-Cha [32] | ~21.5 minutes | High (even at 5 μM) | High, with some endosomal entrapment | Additional C-terminal Cha residue |
| (ChaZZ)₂-Cha [32] | ~19.2 minutes | Moderate (at 20 μM) | Strong, limited endosomal entrapment | Shorter 7-mer with C-terminal Cha |
| DSPE-PEG-VBP/CBP [43] [63] | N/A | Significantly enhanced vs uncoated | Improved cytoplasmic retention & network integration | Lipid-PEG coating with targeting peptides |
| TAT-Pep-1 [59] | N/A | High | Context-dependent | Well-characterized CPPs for mitochondrial delivery |
Table 2: Functional Outcomes of Surface-Engineered Mitochondria in Disease Models
| Therapeutic Approach | Disease Model | Mitochondrial Function Restoration | Cellular Outcomes |
|---|---|---|---|
| Pep-1-mediated delivery [59] | Parkinson's disease (6-OHDA-induced PC12 cells) | Improved ATP production, restored redox balance | Attenuated neurotoxicity, improved cell survival |
| DSPE-PEG-VBP/CBP mitochondria [43] [63] | Human diabetic aortic endothelial cells (DAECs) | Sustained oxygen consumption, improved membrane potential | Enhanced bioenergetic function, vascular repair |
| iPSC-MSC derived mitochondria [63] | Endothelial injury | Maintained polarization, bioenergetic competence | Improved cellular integration and retention |
Experimental Protocols
Protocol 1: Mitochondrial Isolation from iPSC-Mesenchymal Stem Cells
Principle: Isolate bioenergetically competent mitochondria from a reproducible source while maintaining structural integrity and function [63].
Materials:
- iPSC-derived mesenchymal stem cells (≥15×10⁶ cells)
- Mitochondria Isolation Kit (e.g., Thermo Fisher Scientific)
- Reagents A, B, and C from kit
- BCA protein assay kit
- Refrigerated centrifuge capable of 12,000×g
- MitoTracker dyes (e.g., CMXRos) for visualization
Procedure:
- Culture iPSC-MSCs to 80-90% confluency in appropriate growth media.
- For visualization, incubate cells with 500 nM MitoTracker CMXRos for 45 minutes at 37°C before isolation.
- Dissociate cells using trypsin-EDTA and collect by centrifugation at 300×g for 5 minutes.
- Resuspend cell pellet in 800 μL of Buffer A with 10 μL of Mitochondria Isolation Reagent B.
- Incubate on ice for 5 minutes, vortexing every minute to ensure complete lysis.
- Add 800 μL of Reagent C and centrifuge at 700×g for 10 minutes at 4°C.
- Transfer supernatant to a new tube and centrifuge at 3,000×g for 15 minutes at 4°C.
- Resuspend mitochondrial pellet in 500 μL Reagent C and centrifuge at 12,000×g for final purification.
- Resuspend final mitochondrial pellet in 1 mL Reagent C and quantify protein content using BCA assay.
- Use immediately for surface engineering or store at -80°C for up to 14 days.
Validation: Assess mitochondrial membrane potential using JC-1 staining and oxygen consumption rate using Seahorse metabolic analyzer [43].
Protocol 2: Surface Engineering of Mitochondria with CPP-Modified DSPE-PEG
Principle: Create a versatile phospholipid-based coating platform that enables peptide functionalization while enhancing mitochondrial stability and targeting specificity [43] [63].
Materials:
- Freshly isolated mitochondria (from Protocol 1)
- DSPE-PEG-MAL (5,000 Da, Nanosoft Polymers)
- Biotinylated targeting peptides (VCAM-1 binding peptide: VHPKQHRGGSKGC; Collagen binding peptide: CQDSETRTFY)
- Slide-a-Lyzer Dialysis Cassettes (MWCO 7,000 Da)
- Mitochondria Storage Buffer
Procedure: A. Polymer-Peptide Conjugate Synthesis
- React DSPE-PEG-MAL with thiol-containing peptides at a 1:1 molar ratio in ultrapure water.
- Incubate reaction at room temperature for 24 hours with gentle agitation.
- Purify conjugates by dialysis for 24 hours using Slide-a-Lyzer Cassettes.
- Lyophilize purified DSPE-PEG-peptide conjugates and store at -20°C.
B. Mitochondrial Surface Functionalization
- Resuspend freshly isolated mitochondria in Reagent C buffer at 1 mg/mL protein concentration.
- Prepare DSPE-PEG-peptide conjugate solution at 1 mg/mL in Reagent C.
- Combine mitochondria and DSPE-PEG-peptide solutions at optimal mass ratio (determined empirically, typically 1:1 to 1:4 polymer:mitochondria protein).
- Incubate mixture on ice for 3 hours with continuous shaking.
- Remove unbound polymer by centrifugation at 12,000×g for 5 minutes at 4°C.
- Wash twice by replacing supernatant with fresh Reagent C.
- Resuspend functionalized mitochondria in Mitochondria Storage Buffer for immediate use or storage.
Coating Validation: Determine coating efficiency using flow cytometry with AlexaFluor-488 labeled streptavidin to detect biotinylated peptides [43]. Characterize particle size and zeta potential using dynamic light scattering.
Protocol 3: Assessment of Targeting Specificity and Off-Tocalization
Principle: Quantitatively evaluate mitochondrial targeting efficiency and identify off-target localization using confocal microscopy and image analysis [32].
Materials:
- Target cells (e.g., human diabetic aortic endothelial cells)
- CPP-engineered mitochondria (from Protocol 2)
- Organelle-specific dyes: MitoTracker Deep Red (mitochondria), LysoTracker (lysosomes), Hoechst 33342 (nucleus)
- Confocal microscopy system with image analysis software
- Flow cytometer
Procedure:
- Culture target cells in appropriate media on glass-bottom confocal dishes to 70-80% confluency.
- Pre-stain cellular organelles with specific dyes according to manufacturer's protocols.
- Incubate cells with CPP-engineered mitochondria (10-50 μg/mL) for 1-4 hours at 37°C.
- Remove uninternalized mitochondria by gentle washing with PBS.
- Fix cells with 4% paraformaldehyde for 15 minutes at room temperature.
- Image cells using confocal microscopy with consistent settings across experimental groups.
- Quantify colocalization using Pearson's correlation coefficient or Mander's overlap coefficient.
- For flow cytometry analysis, analyze cells after mitochondrial incubation without fixation.
Data Analysis: Calculate specificity ratios by comparing mitochondrial signal colocalization with target organelles versus non-target organelles. Statistically compare engineered mitochondria versus unmodified controls.
Signaling Pathways and Experimental Workflows
Diagram 1: Mitochondrial Delivery Specificity Workflow. This flowchart illustrates the critical decision points in achieving targeted mitochondrial delivery versus off-target localization, highlighting where surface engineering strategies intervene to enhance specificity.
Diagram 2: CPP Design Logic for Mitochondrial Specificity. This diagram outlines the structural and chemical principles governing CPP specificity for mitochondrial targeting, contrasting optimal design strategies with characteristics that lead to off-target localization.
The Scientist's Toolkit: Essential Research Reagents
Table 3: Essential Research Reagents for Mitochondrial Surface Engineering Studies
| Reagent/Category | Specific Examples | Function & Application |
|---|---|---|
| Source Cells | iPSC-derived mesenchymal stem cells (iPSC-MSCs) | Provide bioenergetically competent mitochondria with therapeutic potential [63] |
| CPP Platforms | (ChaZZ)₃, (ChaZZ)₃-Cha, TAT, Pep-1 | Facilitate cellular uptake and mitochondrial targeting; tunable properties enhance specificity [59] [32] |
| Coating Polymers | DSPE-PEG-MAL (5,000 Da) | Enable surface functionalization and provide conjugation platform for targeting ligands [43] [63] |
| Targeting Ligands | VCAM-1 binding peptide (VBP), Collagen binding peptide (CBP) | Direct mitochondria to specific cell types or injury sites [43] [63] |
| Isolation Kits | Mitochondria Isolation Kit (Thermo Fisher Scientific) | Standardized protocols for obtaining functional mitochondrial fractions [63] |
| Validation Dyes | MitoTracker CMXRos, JC-1, LysoTracker | Visualize mitochondrial localization, membrane potential, and assess off-target accumulation [43] [32] |
| Analytical Instruments | Seahorse Analyzer, Confocal Microscope, Zetasizer | Assess metabolic function, visualize intracellular localization, measure particle characteristics [43] [63] |
The strategic surface engineering of mitochondria with selectively designed CPPs represents a transformative approach to overcome the critical challenge of off-target organelle localization. By implementing the protocols and design principles outlined in this document—including the use of structurally rigid PPII helices, optimized hydrophobicity through C-terminal modifications, and targeted ligand conjugation via DSPE-PEG platforms—researchers can significantly enhance mitochondrial delivery specificity. These advanced methodologies provide a foundation for developing precision mitochondrial therapies with improved efficacy and reduced off-target effects, ultimately accelerating the translation of mitochondrial transplantation into viable clinical applications.
Addressing Scalability and GMP Production for Clinical Translation
The surface modification of mitochondria with Cell-Penetrating Peptides (CPPs) represents a promising frontier in therapeutic delivery for a range of diseases linked to mitochondrial dysfunction, including neurodegenerative, cardiovascular, and metabolic disorders [64]. While research demonstrates the potential of engineered mitochondria to restore bioenergetic function in recipient cells, the transition from laboratory proof-of-concept to clinically applicable therapies hinges on overcoming significant challenges in scalable and reproducible manufacturing [36]. This document outlines application notes and detailed protocols designed to address the critical gaps in scaling up and establishing Good Manufacturing Practice (GMP)-compliant production of CPP-surface-modified mitochondria, framing them within a broader thesis on advancing mitochondrial delivery systems.
The following tables consolidate key quantitative findings from recent studies on mitochondrial surface engineering and CPP-mediated delivery, providing a basis for benchmarking and process development.
Table 1: Characterization and Functional Outcomes of Surface-Engineered Mitochondria
| Parameter | Uncoated Mitochondria | DSPE-PEG-Coated Mitochondria | Measurement Technique |
|---|---|---|---|
| Cellular Uptake | Baseline (Reference) | Significantly enhanced [36] | Flow cytometry, confocal microscopy [36] |
| Intracellular Retention (24h) | Lower | Increased cytoplasmic retention & colocalization [36] | Confocal imaging, quantitative analysis [36] |
| Mitochondrial Membrane Potential | Impaired (in dysfunctional models) | Improved [36] | JC-1 staining [36] |
| Oxygen Consumption Rate (OCR) | Reduced (in dysfunctional models) | Sustained / Improved [36] | Seahorse metabolic analysis [36] |
| Primary Cell Model | Human Diabetic Aortic Endothelial Cells (DAECs) [36] |
Table 2: Properties of Mitochondria-Targeting CPPs and Ligands
| Peptide/Ligand | Sequence / Composition | Targeting Motif | Key Functional Property |
|---|---|---|---|
| Amphipathic Oligoproline (e.g., (ChaZZ)₃) | Cha = cyclohexylalanine; Z = (4S)-guanidiniumproline (Gup) [32] | Mitochondria (via PPII helix & amphipathicity) [32] | Rigid PPII helical backbone with aligned cationic/hydrophobic edges; mitochondrial selectivity enhanced by hydrophobicity tuning [32] |
| VCAM-1 Binding Peptide (VBP) | VHPKQHRGGSKGC [36] | VCAM-1 (upregulated in inflamed endothelium) [36] | Enables targeting to dysfunctional/diseased vasculature [36] |
| Collagen Binding Peptide (CBP) | CQDSETRTFY [36] | Collagen IV (exposed upon endothelial injury) [36] | Enables targeting to sites of vascular injury [36] |
| Szeto-Schiller (SS) Peptides | Alternating aromatic-cationic motif (e.g., SS-31) [64] | Mitochondrial Inner Membrane [64] | Antioxidant activity; localizes to inner mitochondrial membrane [64] |
| Triphenylphosphonium (TPP) | Lipophilic cation [4] | Mitochondrial Matrix (driven by membrane potential) [4] | Not a CPP; facilitates accumulation but can cause proton leakage at high concentrations [4] |
Experimental Protocols for Key Processes
Protocol: Isolation of Mitochondria from iPSC-Mesenchymal Stem Cells (iPSC-MSCs)
This protocol is optimized for sourcing mitochondria from a therapeutically relevant and scalable cell line [36].
Materials:
- Cell Source: Induced Pluripotent Stem Cell-derived Mesenchymal Stem Cells (iPSC-MSCs) [36].
- Reagents: Mitochondria Isolation Kit (e.g., Thermo Fisher Scientific), Trypsin, Buffer A, Mitochondria Isolation Reagent B, Reagent C, Mitochondria Storage Buffer, BCA assay kit [36].
Procedure:
- Cell Harvesting: Culture iPSC-MSCs to approximately 80% confluence. Dissociate cells using trypsin and collect ~15 x 10⁶ cells by centrifugation [36].
- Cell Lysis: Resuspend the cell pellet in 800 µL of Buffer A. Add 10 µL of Mitochondria Isolation Reagent B and mix by vortexing. Incubate the suspension on ice for 5 minutes, vortexing briefly every minute to ensure thorough lysis [36].
- Homogenate Clarification: Add 800 µL of Reagent C to the lysate. Centrifuge at 700 × g for 10 minutes at 4°C. Carefully transfer the resulting supernatant (containing the crude mitochondria) to a new, pre-chilled tube [36].
- Mitochondrial Pelletting: Centrifuge the supernatant at 3,000 × g for 15 minutes at 4°C. Discard the supernatant. Resuspend the mitochondrial pellet in 500 µL of Reagent C [36].
- Final Purification: Perform a final centrifugation at 12,000 × g to pellet the purified mitochondria. Discard the supernatant [36].
- Resuspension and Quantification: Resuspend the final mitochondrial pellet in 1 mL of Mitochondria Storage Buffer. Determine the protein concentration using a BCA assay. Store isolated mitochondria at 4°C for immediate use or at -80°C for long-term storage (up to 14 days) [36].
Protocol: DSPE-PEG-Peptide Conjugation and Mitochondrial Surface Functionalization
This protocol describes a scalable phospholipid-based coating method for attaching targeting peptides to the mitochondrial surface [36].
Materials:
- Polymers: 1,2-distearoyl-sn-glycero-3-phosphoethanolamine-N-[maleimide(polyethylene glycol)-5000] (DSPE-PEG-MAL) [36].
- Peptides: Biotinylated VBP or CBP (or other cysteine-containing peptides) [36].
- Equipment: Slide-a-Lyzer Dialysis Cassette (MWCO 7,000 kDa), lyophilizer, MALDI mass spectrometer [36].
Procedure: Part A: Synthesis of DSPE-PEG-Peptide Conjugate
- Reaction Setup: React the cysteine-terminated peptide (e.g., VBP or CBP) with DSPE-PEG-MAL in ultrapure water. Use a 1:1 molar ratio of peptide thiol groups to polymer maleimide groups [36].
- Conjugation: Allow the reaction to proceed at room temperature for 24 hours with gentle agitation [36].
- Purification: Transfer the reaction product to a dialysis cassette (MWCO 7,000) and dialyze against ultrapure water for 24 hours to remove unreacted peptides and small molecules [36].
- Product Isolation: Lyophilize the purified solution to obtain the DSPE-PEG-Peptide conjugate as a solid. Verify successful conjugation using MALDI mass spectrometry [36].
Part B: Surface Engineering of Isolated Mitochondria
- Coating Incubation: Combine an aliquot of freshly isolated mitochondria with a solution of the DSPE-PEG-peptide conjugate (1 mg/mL in Reagent C buffer). Use a defined mass ratio of polymer to mitochondrial protein (e.g., 1:1 to 4:1) and incubate on ice for 3 hours with shaking [36].
- Washing and Storage: After incubation, rinse the functionalized mitochondria twice by centrifuging at 12,000 × g for 5 minutes and replacing the supernatant with fresh Reagent C. Finally, resuspend the pellet in Mitochondria Storage Buffer for immediate use or storage [36].
- Quality Control (Coating Efficiency): Label mitochondria with MitoTracker (e.g., Red CMXRos) prior to isolation. After coating with biotinylated conjugates, incubate with AlexaFluor-488 conjugated streptavidin. Analyze by flow cytometry; coating efficiency is calculated as the percentage of MitoTracker-positive events that are also AlexaFluor-488 positive [36].
Protocol: In Vitro Functional Validation in a Disease-Relevant Model
Materials:
- Cell Model: Human Diabetic Aortic Endothelial Cells (DAECs) or other relevant cell lines [36].
- Stains/Dyes: MitoTracker Deep Red, Hoechst 33342, JC-1 dye [36].
- Equipment: Confocal microscope, flow cytometer, Seahorse XF Analyzer [36].
Procedure:
- Cellular Uptake and Localization:
- Seed DAECs in confocal imaging dishes.
- Treat cells with functionalized mitochondria (e.g., 5-20 µg/mL protein concentration) for 1-4 hours at 37°C.
- Co-stain with MitoTracker Deep Red (for endogenous mitochondria) and Hoechst 33342 (for nuclei).
- Image using confocal microscopy and quantify mitochondrial uptake and colocalization using image analysis software [36].
Assessment of Mitochondrial Membrane Potential (ΔΨm):
- Treat DAECs with engineered mitochondria.
- Incubate with JC-1 dye according to manufacturer's instructions.
- Analyze by fluorescence microscopy or flow cytometry. A shift from red (aggregates) to green (monomers) fluorescence indicates mitochondrial depolarization, while maintained red fluorescence indicates healthy ΔΨm [36].
Bioenergetic Functional Rescue (Seahorse Assay):
- Seed DAECs in a Seahorse XF cell culture microplate.
- Treat with engineered mitochondria 24 hours prior to the assay.
- Measure the Oxygen Consumption Rate (OCR) under basal conditions and in response to stressors (e.g., oligomycin, FCCP, rotenone/antimycin A) using a Seahorse XF Analyzer. Improved OCR indicates restored mitochondrial function [36].
Visualization of Workflows and Pathways
Mitochondria Surface Engineering Workflow
Diagram Title: Mitochondria Surface Engineering and Functionalization Process
CPP-Mediated Mitochondrial Targeting Logic
Diagram Title: Design Logic for Mitochondria-Targeting Motifs
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Mitochondrial Surface Engineering Research
| Item Name | Function / Application | Specific Example / Notes |
|---|---|---|
| iPSC-MSCs | Therapeutic Mitochondria Source: Provides a scalable, clinically relevant source of bioenergetically competent mitochondria [36]. | Differentiated from iPSCs using commercial kits; confirm phenotype with CD73, CD90, CD105 markers [36]. |
| DSPE-PEG-MAL | Coating & Conjugation Platform: Phospholipid-PEG polymer spontaneously inserts into mitochondrial outer membrane; maleimide group enables covalent thiol coupling to peptides [36]. | 5000 Da PEG common; forms a versatile platform for attaching various targeting ligands [36]. |
| Targeting Peptides (VBP, CBP) | Active Targeting Moieties: Direct surface-modified mitochondria to specific cell types or disease sites (e.g., inflamed/damaged endothelium) [36]. | Custom synthesized with a C-terminal cysteine for conjugation and optional biotin tag for tracking [36]. |
| Mitotracker Dyes | Mitochondrial Labeling: Fluorescent dyes for visualizing isolated mitochondria and tracking uptake in recipient cells. | CMXRos (red fluorescent) is retained after fixation; use pre-isolation staining for particle tracking [36]. |
| Seahorse XF Analyzer | Functional Bioenergetic Assay: Measures mitochondrial function in live cells (OCR, ECAR) to validate functional restoration by engineered mitochondria [36]. | Key for demonstrating therapeutic efficacy beyond mere uptake. |
| JC-1 Dye | Membrane Potential Assay: Ratiometric fluorescent probe to assess the health and polarization of mitochondria in recipient cells [36]. | Shift from red to green fluorescence indicates loss of ΔΨm, a key marker of dysfunction. |
Functional Assays and Comparative Analysis of Engineered Mitochondria
The therapeutic potential of mitochondrial transplantation is increasingly recognized for treating diseases rooted in mitochondrial dysfunction, such as neurodegenerative, cardiovascular, and metabolic disorders [11] [62]. A significant bottleneck in the clinical translation of this approach is achieving efficient and targeted delivery of functional mitochondria to damaged cells. Current methods often suffer from nonspecific distribution and poor cellular uptake, with studies suggesting that as little as 10% of injected mitochondria typically reach their intended cellular targets [11]. Surface engineering of mitochondria, particularly using cell-penetrating peptides (CPPs), has emerged as a promising strategy to overcome these delivery barriers [43] [17]. CPPs are short peptides (typically 5-30 amino acids) rich in cationic residues like arginine and lysine, which facilitate efficient translocation across cell membranes [17]. The development of robust, quantitative methods to evaluate the success of these engineering strategies is therefore paramount. This application note details the integrated use of flow cytometry and confocal microscopy to precisely quantify the cellular uptake efficiency of CPP-enhanced mitochondria, providing researchers with a reliable framework for optimizing next-generation mitochondrial therapeutics.
Key Principles of Uptake Quantification
Accurately quantifying the uptake of engineered mitochondria requires an understanding of both the biological process and the technical capabilities of the instrumentation used to measure it. Isolated mitochondria can be internalized by recipient cells primarily through energy-dependent endocytic pathways, such as macropinocytosis [65]. A critical subsequent step for therapeutic efficacy is the escape of internalized mitochondria from endosomal compartments into the cytosol, where they may integrate into the host's existing mitochondrial network [65]. Flow cytometry and confocal microscopy offer complementary insights into this dynamic process. Flow cytometry excels at providing rapid, quantitative, and statistically significant data on the proportion of a cell population that has internalized labeled mitochondria and the relative amount taken up per cell [66]. In contrast, confocal microscopy provides high-resolution spatial information, confirming intracellular localization, verifying endosomal escape, and observing potential integration with the endogenous mitochondrial network [43] [65]. Used in tandem, these methods provide a comprehensive assessment of uptake efficiency and fate, which is essential for validating the functional benefit of CPP-based surface modifications.
Experimental Protocols
Mitochondrial Isolation, Labeling, and Surface Engineering
Key Reagents:
- Source of Mitochondria: Induced pluripotent stem cell-derived mesenchymal stem cells (iPSC-MSCs) are a preferred source due to their bioenergetic competence [43].
- Isolation Kit: Mitochondrial Isolation Kit for Cultured Cells.
- Fluorescent Dyes: MitoTracker Green FM (for flow cytometry) or MitoTracker Deep Red FM (for confocal microscopy to avoid autofluorescence). Alternatively, lipophilic carbocyanine dyes like DiOC6(3) can be used [65].
- Surface Engineering Reagents: DSPE-PEG phospholipid-based coating platform, conjugated with VCAM-1-binding or collagen-binding peptides for targeting, and/or CPPs such as TAT or Pep-1 [43] [11].
Procedure:
- Isolation: Isolate mitochondria from donor cells using a standard differential centrifugation protocol or a commercial kit to ensure the harvest of a pure and functionally intact mitochondrial fraction [43].
- Fluorescent Labeling: Resuspend the mitochondrial pellet in incubation buffer containing a low concentration (e.g., 20-100 nM) of the chosen MitoTracker dye. Incubate for 15-30 minutes at 37°C. Pellet the mitochondria by centrifugation and wash twice with fresh buffer to remove unincorporated dye.
- Surface Functionalization: Incubate the labeled mitochondria with the DSPE-PEG-CPP conjugate (e.g., at a weight ratio of 1750:1 for Pep-1) for 30-60 minutes at 37°C to form the engineered complex [11]. Use uncoated mitochondria as a negative control.
- Characterization: Validate the size, charge (zeta potential), and concentration of the engineered mitochondria using dynamic light scattering (DLS) and flow cytometry [43]. Confirm bioenergetic function via Seahorse XF Analyzer.
Cell Culture and Mitochondrial Uptake
Key Reagents:
- Recipient Cells: Human diabetic aortic endothelial cells (DAECs) for vascular repair models, or other relevant cell lines (e.g., HeLa, A431) [43] [65].
- Cell Culture Medium: Appropriate complete growth medium (e.g., DMEM with 10% FBS).
Procedure:
- Seed recipient cells in appropriate culture vessels (e.g., 12-well plates for flow cytometry, glass-bottom dishes for confocal microscopy) and culture until they reach 60-80% confluency.
- Gently wash the cells with pre-warmed PBS.
- Uptake Incubation: Add the suspension of fluorescently labeled, CPP-engineered mitochondria to the cells in a serum-free or low-serum medium. A typical mitochondria-to-cell ratio can be optimized but often ranges from 100:1 to 500:1.
- Incubate the cells for a defined period (e.g., 4-24 hours) at 37°C in a 5% CO₂ incubator [43] [65].
- Wash and Trypsinization: After incubation, thoroughly wash the cells with PBS to remove non-internalized mitochondria. Treat the cells with trypsin-EDTA (e.g., for 5-10 minutes) to dissociate adherent cells and remove any mitochondria bound to the external cell surface.
- Terminate trypsinization with complete medium, collect the cells by centrifugation, and resuspend in flow cytometry buffer (PBS with 1% BSA or FBS) for analysis.
Quantitative Analysis via Flow Cytometry
Instrument Setup:
- Use a standard flow cytometer equipped with lasers and filters appropriate for your fluorescent dye (e.g., 488 nm laser and 530/30 nm filter for MitoTracker Green).
- Establish forward scatter (FSC) and side scatter (SSC) settings to gate on the live, single-cell population.
- Create a fluorescence histogram and dot plot to analyze the signal from the mitochondrial label.
Data Acquisition and Analysis:
- Run control samples (unstained cells, cells incubated with unlabeled mitochondria) to establish autofluorescence baseline and set the positive signal gate.
- Acquire a minimum of 10,000 events per sample.
- Quantify Uptake:
- Percentage of Positive Cells: Determine the proportion of cells exhibiting fluorescence above the threshold set by the control sample.
- Mean Fluorescence Intensity (MFI): Measure the average fluorescence intensity of the positive cell population, which correlates with the average mitochondrial load per cell.
- Use the following table to summarize key quantitative findings from the literature, which can serve as a benchmark for expected outcomes:
Table 1: Quantitative Uptake Data from Representative Studies
| Mitochondrial Type | Target Cell | Incubation Time | Uptake Efficiency (Flow Cytometry) | Key Finding |
|---|---|---|---|---|
| CPP-coated (Pep-1) [11] | PC12 cells | 30 min | Significantly higher than uncoated | Pep-1 conjugation enhances mitochondrial transfer efficiency. |
| DSPE-PEG-coated [43] | Diabetic Aortic Endothelial Cells | 24 h | Significantly enhanced vs. uncoated | Coating improves cytoplasmic retention and network integration. |
| Free isolated mitochondria (F4') [65] | Various adherent cell lines (HeLa, A431, SKOV3) | 24 h | 1-2% of loaded NLuc activity | A small but measurable fraction is internalized by macropinocytosis. |
Diagram 1: Flow cytometry data analysis workflow.
Spatial Analysis and Validation via Confocal Microscopy
Instrument Setup:
- Use a confocal laser scanning microscope with high-resolution objectives (60x or 100x oil immersion).
- Configure laser lines and emission filters for your mitochondrial label (e.g., 640 nm laser and 670 nm LP filter for MitoTracker Deep Red) and any counterstains.
Staining and Imaging Procedure:
- After the uptake incubation, wash the cells grown on glass-bottom dishes with PBS.
- Counterstaining:
- Nucleus: Stain with Hoechst 33342 (1-2 µg/mL) for 10 minutes.
- Endogenous Mitochondria: Stain with a different MitoTracker dye (e.g., MitoTracker Green for cells that took up Deep Red-labeled mitochondria) for 30 minutes.
- Lysosomes/Endosomes: Stain with LysoTracker or immunostain for LAMP1 protein to investigate endosomal escape.
- Acquire Z-stack images (0.5 µm slices) to capture the full 3D volume of the cells.
- Image Analysis:
- Colocalization Analysis: Use software (e.g., ImageJ with JaCoP plugin or Imaris) to calculate Pearson's or Manders' coefficients between the signal of internalized mitochondria and markers for lysosomes/endosomes or the endogenous mitochondrial network.
- Cytoplasmic Localization: Visually confirm and quantify the presence of labeled mitochondria in the cell cytoplasm, outside of membrane-bound organelles.
Table 2: Key Confocal Microscopy Findings from Literature
| Analysis Type | Experimental Setup | Observation | Interpretation |
|---|---|---|---|
| Cytoplasmic Retention [43] | DAECs incubated with DSPE-PEG-mitochondria | Increased cytoplasmic signal after 24h | Surface coating promotes mitochondrial stability and retention inside the cell. |
| Network Integration [43] | DAECs incubated with engineered mitochondria | Greater colocalization with host mitochondrial network | Internalized mitochondria can functionally integrate. |
| Endosomal Escape [65] | Cells incubated with free extracellular mitochondria | Subset of mitochondria escape protease-rich compartments | Essential step for functional integration; occurs in <10% of internalized organelles. |
Diagram 2: Confocal microscopy validation workflow.
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagent Solutions for Mitochondrial Uptake Studies
| Reagent / Kit | Supplier Examples | Function in the Protocol |
|---|---|---|
| MitoTracker Probes | Thermo Fisher Scientific, Abcam | Fluorescently label isolated mitochondria for tracking and quantification. |
| Mitochondrial Isolation Kit | Abcam, Sigma-Aldrich, Miltenyi Biotec | Isulate pure and functionally intact mitochondria from donor cells. |
| Cell-Penetrating Peptides (TAT, Pep-1) | Genscript, AnaSpec, Bachem | Engineer mitochondrial surface to enhance cellular uptake and delivery. |
| DSPE-PEG Reagents | Nanocs, Creative PEGWorks | Provide a biocompatible platform for conjugating targeting peptides/CPPs to mitochondria. |
| Seahorse XF Kits | Agilent Technologies | Analyze mitochondrial bioenergetic function pre- and post-engineering. |
| LysoTracker Probes | Thermo Fisher Scientific | Label acidic endo-lysosomal compartments to monitor mitochondrial escape. |
Critical Considerations and Methodological Pitfalls
- Fluorescence Quenching: Tryptophan-rich CPPs can quench the fluorescence of certain dyes (like TAMRA) when in close proximity, potentially leading to an underestimation of uptake in flow cytometry. This quenching can be reversed upon cell lysis, creating a discrepancy between flow cytometry and spectroscopic measurements [66]. Choosing bright, photostable dyes (e.g., MitoTracker) and validating results with multiple methods is crucial.
- Specificity of Uptake Signal: Thorough washing and trypsinization are critical to ensure that the quantified fluorescence represents internalized mitochondria, not merely surface-adhered particles [65]. A 4°C temperature block can be used as a control to inhibit energy-dependent endocytosis and confirm the specificity of the uptake mechanism.
- Functional Correlation: Quantifying uptake is only the first step. Researchers must correlate increased uptake with functional improvements in recipient cells, such as enhanced mitochondrial membrane potential, increased ATP production, and improved oxygen consumption rates, as measured by JC-1 staining and Seahorse metabolic analysis [43].
Within the rapidly advancing field of mitochondrial medicine, the therapeutic potential of surface-modified mitochondria, particularly those engineered with cell-penetrating peptides (CPPs), is gaining significant traction. These modified mitochondria are designed for enhanced cellular delivery to rescue cells with dysfunctional mitochondrial networks, showing promise for treating a range of diseases from neurodegeneration to cardiovascular disorders [59]. However, the development of such therapies is critically dependent on robust, quantitative methods to validate mitochondrial function post-modification and post-delivery. This document provides detailed application notes and protocols for two cornerstone techniques in metabolic functional validation: the Seahorse XF Analyzer, which provides a real-time measurement of cellular bioenergetics, and JC-1 staining, a sensitive fluorescent probe for assessing mitochondrial membrane potential (ΔΨM) [67] [68] [69]. When applied to CPP-enhanced mitochondrial transplantation studies, these assays are indispensable for confirming that the delivered mitochondria are not only taken up by cells but are also functionally active and capable of improving the host's bioenergetic health [36].
Metabolic Profiling with the Seahorse XF Analyzer
The Seahorse XF Analyzer measures the Oxygen Consumption Rate (OCR) and Extracellular Acidification Rate (ECAR) of live cells in real-time, providing a window into fundamental metabolic processes. The OCR is a direct indicator of mitochondrial oxidative phosphorylation (OXPHOS), while the ECAR primarily reflects glycolytic activity [67] [48]. This platform is especially powerful for evaluating the bioenergetic consequences of introducing CPP-modified mitochondria into recipient cells, allowing researchers to quantify the functional contribution of the transplanted organelles to the host's energy metabolism.
Key Application: Validating CPP-Mitochondria Bioenergetic Rescue
Recent studies highlight the utility of Seahorse technology in proving the efficacy of mitochondrial transplantation. For instance, one investigation demonstrated that iPSC-derived mitochondria, once surface-engineered for targeted delivery, could be taken up by human diabetic aortic endothelial cells. Subsequent Seahorse metabolic analysis confirmed that cells receiving these engineered mitochondria showed a significant improvement in their OCR, underscoring a successful restoration of bioenergetic function [36]. This type of data is critical for moving mitochondrial transplantation toward clinical application.
A standard method to probe mitochondrial function in depth is the Mitochondrial Stress Test, which uses sequential injections of metabolic modulators to dissect the individual components of the respiratory chain.
Table: Reagents for the Seahorse XF Cell Mito Stress Test
| Reagent | Final Working Concentration | Function in the Assay |
|---|---|---|
| Oligomycin | 1.0 - 2.0 µM | ATP synthase inhibitor. Measures ATP-linked respiration and reveals proton leak. |
| FCCP | 0.5 - 2.0 µM | Mitochondrial uncoupler. Collapses the proton gradient, forcing the ETC to operate at maximum capacity to reveal maximal respiratory capacity. |
| Antimycin A & Rotenone | 0.5 µM each | Inhibitors of Complex III and I, respectively. Shut down mitochondrial respiration, allowing quantification of non-mitochondrial oxygen consumption. |
Table: Key Bioenergetic Parameters from Mitochondrial Stress Test Data
| Parameter | Description | Biological Interpretation |
|---|---|---|
| Basal Respiration | OCR measured before any drug injection. | The combined energy demand of the cell under baseline conditions. |
| ATP-linked Respiration | The drop in OCR after Oligomycin injection. | The portion of basal respiration used to drive ATP production. |
| Maximal Respiration | The OCR measured after FCCP injection. | The maximum respiratory capacity of the cell, indicating its ability to respond to energetic stress. |
| Spare Respiratory Capacity | Maximal Respiration - Basal Respiration. | A bioenergetic reserve; higher values indicate better fitness and health. |
| Non-Mitochondrial Respiration | OCR remaining after Antimycin A & Rotenone. | Oxygen consumption from cellular processes outside the mitochondria. |
Detailed Protocol: Mitochondrial Stress Test for Cells Receiving CPP-Modified Mitochondria
Materials:
- Seahorse XFp/XFe96 Analyzer (Agilent Technologies)
- XFp/XFe96 Cell Culture Microplate (Agilent Technologies)
- XF Calibrant (Agilent Technologies, Part 100840-000)
- XF Cell Mito Stress Test Kit (Agilent Technologies, contains oligomycin, FCCP, antimycin A, and rotenone) [67]
Procedure:
- Cell Seeding: 24 hours prior to the assay, seed recipient cells (e.g., human diabetic aortic endothelial cells) in the Seahorse microplate at an optimized density (e.g., 20,000-30,000 cells/well for a 96-well plate). Include control groups (untreated cells and cells treated with CPP-modified mitochondria).
- Mitochondrial Transplantation: On the day of the assay, treat cells with the prepared CPP-modified mitochondria. A typical protocol might involve incubating cells with 50-100 µg/mL of isolated mitochondria for 1-4 hours [36].
- Assay Medium Preparation: Prepare Seahorse XF Base Medium supplemented with 1 mM pyruvate, 2 mM glutamine, and 10 mM glucose. Adjust pH to 7.4 ± 0.1 and warm to 37°C.
- Sensor Cartridge Hydration: Hydrate the Seahorse sensor cartridge with XF Calibrant in a non-CO₂ incubator at 37°C for at least 12 hours before the assay.
- Drug Loading: Load the ports of the hydrated sensor cartridge with the Mito Stress Test modulators:
- Port A: Oligomycin (1.5 µM final concentration in well)
- Port B: FCCP (1.0 µM final concentration in well)
- Port C: Antimycin A & Rotenone (0.5 µM final concentration each in well)
- Cell Preparation: After the mitochondrial incubation period, carefully replace the cell culture medium with the pre-warmed assay medium. Incubate the cell culture microplate for 45-60 minutes in a non-CO₂ incubator at 37°C.
- Run Assay: Place the calibrated sensor cartridge and cell culture microplate into the Seahorse Analyzer and start the pre-programmed Mito Stress Test protocol. The standard protocol involves a 2-3 minute mix, a 2-3 minute wait, and a 3-5 minute measurement period for each cycle [67].
- Data Normalization: Upon assay completion, normalize the OCR and ECAR data to total cellular protein content per well (determined by a BCA assay) to account for any differences in cell number or mitochondrial uptake.
Assessing Mitochondrial Health with JC-1 Staining
The mitochondrial membrane potential (ΔΨM) is a critical indicator of mitochondrial health and functional integrity, driving ATP synthesis. The JC-1 dye is a cationic, lipophilic fluorescent probe that serves as a sensitive and quantitative measure of ΔΨM [68] [69]. Its unique property of forming potential-dependent "J-aggregates" allows for a ratiometric measurement, which is less susceptible to artifacts related to mitochondrial size, shape, or dye concentration.
Key Application: Confirming Membrane Polarization after CPP-Mitochondria Uptake
In the context of mitochondrial transplantation, a key question is whether the delivered mitochondria retain a healthy ΔΨM. JC-1 staining is perfectly suited to answer this. For example, research on surface-engineered mitochondria has used JC-1 staining to demonstrate that recipient cells show a significantly higher red/green fluorescence ratio, indicating better-preserved ΔΨM, compared to cells treated with unmodified mitochondria [36]. This confirms that the transplanted organelles are not only present but are also polarized and functionally competent.
The principle of the assay is straightforward: in healthy, polarized mitochondria, JC-1 accumulates and forms aggregates that emit red fluorescence (~590 nm). In depolarized mitochondria, the dye cannot accumulate and remains in its monomeric form, emitting green fluorescence (~529 nm). Therefore, a decrease in the red/green fluorescence intensity ratio is a direct indicator of mitochondrial depolarization, a common early event in apoptosis and cellular stress [69].
Detailed Protocol: JC-1 Staining for Flow Cytometry
Materials:
- MitoProbe JC-1 Assay Kit (Thermo Fisher Scientific, Catalog No. M34152) [69]
- Flow cytometer equipped with 488 nm laser and FITC (530/30) & PE (585/42) detectors
- Carbonyl cyanide m-chlorophenyl hydrazone (CCCP, from kit), as a depolarization control
- Phosphate-buffered saline (PBS)
- DMSO
Procedure:
- Cell Preparation: Harvest recipient cells (e.g., 1x10⁶ cells/mL) that have been co-incubated with CPP-modified mitochondria. Include two control samples: an untreated control (healthy) and a control treated with 50 µM CCCP for 10-20 minutes at 37°C to fully depolarize mitochondria [68].
- JC-1 Staining Solution: Prepare a 2 µM JC-1 working solution in pre-warmed PBS or culture medium from the 200 µM JC-1 stock (prepared in DMSO).
- Staining: Pellet the cells and resuspend them in 1 mL of the JC-1 working solution. Incubate for 30-45 minutes at 37°C in the dark.
- Washing: Centrifuge the cells at 400 x g for 5 minutes. Carefully aspirate the supernatant and wash the cell pellet once with 2 mL of warm PBS.
- Resuspension and Analysis: Resuspend the final cell pellet in 500 µL of warm PBS. Keep the samples on ice and in the dark until immediate analysis on the flow cytometer.
- Flow Cytometry Setup:
- Use 488 nm excitation.
- Detect green JC-1 monomers with a FITC/GFP channel (e.g., 530/30 nm BP filter).
- Detect red JC-1 aggregates with a PE channel (e.g., 585/42 nm BP filter).
- Collect at least 10,000 events per sample.
- Data Analysis:
- Create a density plot of PE (Red J-aggregates) vs. FITC (Green Monomers).
- Gate on the viable cell population.
- Calculate the geometric mean fluorescence intensity (MFI) for both red and green channels.
- Determine the Red/Green Fluorescence Ratio (MFI-PE / MFI-FITC) for each sample. A higher ratio in the test group compared to a dysfunctional control indicates improved mitochondrial membrane potential.
The Scientist's Toolkit: Essential Reagents for Mitochondrial Functional Validation
Table: Key Research Reagent Solutions
| Reagent / Kit | Vendor Examples | Primary Function in Validation |
|---|---|---|
| Seahorse XF Cell Mito Stress Test Kit | Agilent Technologies | Provides optimized concentrations of oligomycin, FCCP, and antimycin A/rotenone for standardized profiling of mitochondrial function in live cells. |
| MitoProbe JC-1 Assay Kit | Thermo Fisher Scientific | A complete kit containing JC-1 dye and the control uncoupler CCCP, optimized for flow cytometry to assess mitochondrial membrane potential. |
| Cell-Penetrating Peptides (CPPs) | Custom synthesis vendors (e.g., ABI Scientific) | Synthetic peptides (e.g., Pep-1, TAT) used to coat isolated mitochondria to enhance their cellular uptake in transplantation studies [59]. |
| Mitotracker Probes (e.g., Deep Red) | Thermo Fisher Scientific | Fixable mitochondrial stains used for tracking mitochondrial localization, uptake, and colocalization within recipient cells via microscopy. |
| DSPE-PEG-Based Polymers | Nanosoft Polymers | Used for surface engineering of mitochondria to improve stability, prevent aggregation, and provide a conjugation point for targeting peptides (e.g., VCAM-1 binding peptide) [36]. |
| Mitochondria Isolation Kits | Thermo Fisher Scientific, Millipore Sigma | Used to isolate intact, functional mitochondria from donor cells (e.g., iPSC-MSCs) prior to surface modification and transplantation. |
The integration of Seahorse Analyzer metabolic profiling and JC-1 staining for membrane potential assessment provides a powerful, complementary framework for the functional validation of therapeutic mitochondria. In the specific context of enhancing mitochondrial delivery via CPP-based surface modification, these assays move beyond simple proof-of-uptake to deliver quantitative evidence of bioenergetic rescue. The Seahorse assay captures the global, functional outcome of successful transplantation through improved oxygen consumption, while JC- staining offers a direct, organelle-level confirmation of the health and integrity of the newly integrated mitochondrial network. By adhering to these detailed protocols, researchers can robustly quantify the efficacy of novel mitochondrial delivery strategies, thereby accelerating the development of mitochondrial transplantation as a viable therapeutic modality.
Mitochondria, the powerhouses of eukaryotic cells, are critical targets for treating a spectrum of diseases, including neurodegenerative disorders, cancer, and metabolic syndromes, due to their central role in apoptosis, metabolism, and redox signaling [62] [70] [71]. The primary challenge in mitochondrial medicine is the efficient and selective delivery of therapeutic agents across the double-membrane structure of the organelle. Two dominant chemical strategies have emerged to overcome this barrier: Cell-Penetrating Peptides (CPPs) and Triphenylphosphonium (TPP) cations. CPPs are short peptides (typically 5-30 amino acids) capable of traversing cellular membranes and can be engineered for organelle specificity [17]. TPP is a lipophilic cation that exploits the mitochondrial membrane potential (ΔΨm) for accumulation [49] [70]. This application note provides a comparative analysis and detailed protocols for these two approaches, supporting research for the surface modification of mitochondria to enhance delivery.
Comparative Analysis: Mechanisms and Properties
The following table summarizes the core characteristics of CPP and TPP-based mitochondrial targeting strategies, highlighting their distinct advantages and challenges.
Table 1: Comparative Analysis of CPP-Modified and TPP-Targeted Mitochondrial Delivery Systems
| Feature | CPP-Based Targeting | TPP-Based Targeting |
|---|---|---|
| Chemical Nature | Peptides (e.g., oligoprolines, Szeto-Schiller peptides) [32] [70] | Small lipophilic cation (delocalized positive charge) [70] [71] |
| Primary Targeting Mechanism | Amphipathicity; specific sequence motifs (e.g., alternating cationic/hydrophobic residues) [32] [4] | Driven by mitochondrial membrane potential (ΔΨm; ~180-200 mV) [70] [71] |
| Typical Cargo Linked | Proteins, nucleic acids, peptides [4] [17] | Small molecule drugs, antioxidants, nanoparticles [49] [70] [72] |
| Key Structural Features | Rigid PPII helix with aligned cationic/hydrophobic edges; +3 net charge; log P > -1.7 [32] [70] | Delocalized positive charge; lipophilicity [70] |
| Cellular Uptake & Specificity | Can achieve selective mitochondrial residency over time; tunable via hydrophobicity [32] | General targeting; accumulates 100-500x in mitochondria vs. cytosol; higher uptake in cancer cells (higher ΔΨm) [70] |
| Major Advantage(s) | High tunability, ability to deliver macromolecules, potential for high selectivity [32] [17] | Simplicity, well-established chemistry, broad applicability, high accumulation factor [49] [70] |
| Major Challenge(s) | Potential endosomal entrapment, stability, optimizing sequence for specificity [32] [17] | Lack of inherent specificity for mitochondrial sub-compartments, dependence on ΔΨm [70] |
Experimental Protocols for Mitochondrial Targeting
Protocol: Evaluating Amphipathic CPPs for Mitochondrial Delivery
This protocol outlines the synthesis and evaluation of amphipathic, proline-rich CPPs based on the (XZZ)₃ motif (Z = Gup; X = hydrophobic amino acid like Cha), as described in recent literature [32].
1. Peptide Synthesis and Characterization
- Solid-Phase Peptide Synthesis (SPPS): Synthesize the (ChaZZ)₃ sequence using Fmoc/t-Bu chemistry. Incorporate 5(6)-Carboxyfluorescein (CF) via an aminohexanoic acid (Ahx) spacer at the N-terminus for visualization [32].
- Purification and Analysis: Purify the crude peptide using Reverse-Phase High-Performance Liquid Chromatography (RP-HPLC) on a C4 column. Use a gradient of 30% to 55% mobile phase B (MeCN) in A (H2O/MeCN/TFA) over 20 minutes. The retention time provides an estimate of peptide hydrophobicity [32].
- Circular Dichroism (CD) Spectroscopy: Confirm the formation of a polyproline II (PPII) helix. Characteristic spectra show a minimum at ~206 nm and a maximum at ~224 nm. This rigid helical structure is crucial for proper alignment of functional groups and mitochondrial targeting [32].
2. Cellular Uptake and Localization Assay
- Cell Culture and Staining: Culture MCF-7 breast cancer cells on glass-bottom confocal dishes. Prior to imaging, incubate cells with the CF-labeled peptide (e.g., 10 µM) for 1 hour at 37°C [32].
- Mitochondrial Counterstaining: Co-incubate cells with MitoTracker Deep Red (e.g., 50-100 nM) to visualize mitochondria. Use Hoechst 33342 (e.g., 1-2 µg/mL) to stain the nucleus [32].
- Confocal Microscopy Imaging: Image live or fixed cells using a confocal laser scanning microscope. Use appropriate laser lines and emission filters to distinguish CF (green), MitoTracker (far-red), and Hoechst (blue) signals.
- Colocalization Analysis: Quantify the degree of overlap between the peptide (green) and mitochondrial (red) signals using software such as ImageJ (with JACoP plugin) or Imaris. Report Manders' or Pearson's correlation coefficients to objectively assess targeting efficiency.
Protocol: Conjugating and Testing TPP-Modified Nanocarriers
This protocol details the construction of a TPP-modified, biomimetic nanocarrier for mitochondrial delivery of small molecules, such as the anti-atherosclerosis drug Empagliflozin (EMP) [72].
1. Synthesis of TPP-Polymer Conjugate and Nanoparticle Formulation
- Synthesis of TPP-PCL-PEG Copolymer:
- Synthesize α,ω-dicarboxyl polyethylene glycol (HOOC-PEG-COOH) and poly(ε-caprolactone) (PCL) separately.
- Couple PEG and PCL to form the amphiphilic block copolymer HOOC-PCL-PEG.
- Synthesize (3-Hydroxypropyl)triphenylphosphonium bromide (TPP-OH) by reacting triphenylphosphine with 3-bromo-1-propanol.
- Conjugate TPP-OH to the HOOC-PCL-PEG copolymer via a DMAP/DCC-mediated esterification reaction to form the final TPP-PCL-PEG polymer [72].
- Nanoparticle Preparation and Drug Loading: Prepare empty TPP-PCL-PEG micelles using a solvent evaporation method. Load Empagliflozin (EMP) by incubating the drug with the micelles. Purify the drug-loaded nanoparticles (EPPT) via dialysis or centrifugation [72].
- Platelet Membrane Coating:
- Isolate platelet membranes from fresh blood by repeated freeze-thaw cycles and differential centrifugation.
- Fuse the isolated platelet membranes onto the pre-formed EPPT nanoparticles by co-extrusion through a porous membrane (e.g., 400 nm polycarbonate membrane) to create the final dual-targeted system, PM@EPPT [72].
2. In Vitro Functional Validation
- Cellular Uptake and Mitochondrial Colocalization: Treat relevant cells (e.g., macrophages) with PM@EPPT nanoparticles. Use a similar protocol as in 3.1, with MitoTracker staining, to confirm mitochondrial targeting via confocal microscopy [72].
- Assessment of Mitochondrial Functional Recovery:
- Reactive Oxygen Species (ROS) Scavenging: Use the CellROX Deep Red reagent to measure intracellular ROS levels via flow cytometry after treatment with oxidative stress inducers (e.g., ox-LDL) and the nanocarrier.
- Mitochondrial Membrane Potential (ΔΨm) Assay: Use the JC-1 dye. A decrease in the red/green fluorescence intensity ratio indicates mitochondrial membrane depolarization. Treatment with functional PM@EPPT should restore a high red/green ratio.
- Mitophagy Activation: Monitor mitophagy levels using immunofluorescence for markers like LC3 and TOM20 to observe mitochondrial clearance [72].
The Scientist's Toolkit: Essential Reagents and Materials
Table 2: Key Research Reagent Solutions for Mitochondrial Targeting Studies
| Reagent/Material | Function/Application | Example Use Case |
|---|---|---|
| MitoTracker Probes (e.g., Deep Red) | Fluorescent staining of live-cell mitochondria; counterstain for colocalization studies. | Visualizing mitochondrial network and confirming subcellular localization of delivered cargo in Protocol 3.1. [32] |
| JC-1 Dye | A cationic dye that exhibits potential-dependent accumulation in mitochondria, indicated by a fluorescence shift from green (~529 nm) to red (~590 nm). Used to measure ΔΨm. | Assessing the health of mitochondria and whether a therapeutic restores depolarized membrane potential in Protocol 3.2. [72] [71] |
| CellROX Reagents | Cell-permeant fluorogenic probes that become fluorescent upon oxidation by ROS. Measure oxidative stress in live cells. | Quantifying the antioxidant effect of a mitochondria-targeted drug like EMP in Protocol 3.2. [72] |
| TPP (Triphenylphosphonium) | A small, lipophilic cation used as a mitochondrial-targeting moiety. Can be conjugated to drugs, probes, or nanocarriers. | Synthesizing the mitochondrial-targeting copolymer TPP-PCL-PEG for nanoparticle construction in Protocol 3.2. [49] [72] |
| Szeto-Schiller (SS) Peptides | A class of mitochondria-penetrating peptides (MPPs) with alternating aromatic and basic residues. | Used as a positive control for peptide-based mitochondrial delivery or as an alternative CPP scaffold. [70] |
| PCL-PEG Copolymer | A biodegradable, amphiphilic block copolymer used to form polymeric micelles for drug encapsulation. | Serves as the core structural material for constructing the nanocarrier in Protocol 3.2. [72] |
Visualization of Signaling Pathways and Workflows
Mitochondrial Targeting Mechanisms
Experimental Workflow for CPP Evaluation
Evaluating Long-Term Retention and Network Integration
Application Note
This document provides detailed application notes and protocols for evaluating the long-term retention and functional integration of exogenous mitochondria delivered into mammalian cells via surface modification with Cell-Penetrating Peptides (CPPs). The methodologies outlined are designed for researchers aiming to validate and quantify the success of mitochondrial transfer and transplantation (MTT) experiments, with a specific focus on persistence and network integration over extended periods (up to 4 weeks).
The surface modification of mitochondria or mitochondrial components with CPPs, such as RD3AD, TAT, or Pep-1, is a key biotechnological strategy to enhance cellular uptake and mitochondrial targeting [59] [73] [74]. Successful integration is defined not only by the physical presence of the delivered material but also by its functional activity in restoring or enhancing cellular bioenergetics. The protocols below are framed within a broader research context aiming to develop CPP-mediated delivery as a pan-therapy for mitochondrial diseases arising from mtDNA mutations or protein deficiencies [75] [73] [74].
Experimental Protocols
Protocol 1: Evaluating Long-Term mtDNA Retention and Function
This protocol is adapted from Wilson et al. (2025) and is designed to assess the delivery and long-term persistence of intact mitochondrial DNA (mtDNA) using a CPP-based nanocomplex [75] [73].
1. Fabrication of CPP-mtDNA Nanocomplex
- Objective: To create a stable nanoparticle for delivering intact, functional mtDNA into cells.
- Materials:
- Methodology:
- Complex the isolated mtDNA with Rho123 via co-precipitation [73].
- Add the mtDNA/Rho123 complex to the RD3AD peptide at varying Nitrogen:Phosphate (N:P) molar ratios (e.g., 1.125 to 11.25) [73].
- Incubate the mixture to allow for self-assembly into nanoparticles.
- Characterization: Use gel shift assays to confirm complete complexation. Utilize Nanoparticle Tracking Analysis (NTA) and Atomic Force Microscopy (AFM) to determine particle size, zeta potential, and morphology [73]. A completely bound complex at an N:P ratio of 11.25 should yield particles of approximately 150-200 nm with a near-neutral zeta potential [73].
2. Cell Treatment and Long-Term Culture
- Objective: To introduce the nanocomplex into target cells and monitor retention over time.
- Cell Model: Mitochondria-depleted cells (e.g., Rho0 ARPE-19 cells) are ideal for tracking exogenous mtDNA without background interference [73].
- Methodology:
- Treat cells with the fabricated CPP-mtDNA nanocomplex.
- Culture the treated cells for an extended period (at least 4 weeks), passaging as normal [73].
- Analyze cells at regular intervals (e.g., weekly) for the presence and functionality of the delivered mtDNA.
3. Assessment of Long-Term Retention and Integration
- Key Metrics and Techniques:
- Exogenous mtDNA Retention: Use quantitative PCR (qPCR) with species-specific primers to detect and quantify the delivered mtDNA over the 4-week culture period [73].
- mtDNA Functionality:
- Mitochondrial RNA (mtRNA) Production: Detect the expression of mitochondrial-encoded transcripts (e.g., ND1, ND4) via RT-qPCR or RNA-Seq [73].
- Mitochondrial Protein Production: Assess the restoration of mitochondrial-encoded proteins (e.g., subunits of the electron transport chain) via Western blotting or immunofluorescence [73].
- Functional Bioenergetic Rescue: Measure the oxygen consumption rate (OCR), ATP production, and lactate secretion using a Seahorse Analyzer and biochemical assays to confirm the functional integration of the delivered genome [73] [74].
Table 1: Key Quantitative Data from CPP-mtDNA Delivery Studies
| Parameter | Measurement Technique | Reported Outcome | Timeline |
|---|---|---|---|
| mtDNA Retention | Species-specific qPCR | Exogenous mtDNA maintained in recipient cells [73] | At least 4 weeks [73] |
| mtRNA Production | RT-qPCR | Restoration of mitochondrial transcripts [73] | At least 4 weeks [73] |
| Protein Production | Western Blot | Detection of mitochondrial-encoded proteins [73] | At least 4 weeks [73] |
| Particle Size | Nanoparticle Tracking Analysis | 151.6 ± 9.7 nm (at N:P 11.25) [73] | Post-fabrication [73] |
| Zeta Potential | Nanoparticle Tracking Analysis | -3.39 ± 1.7 mV (at N:P 11.25) [73] | Post-fabrication [73] |
Protocol 2: Assessing Integration of Transplanted Whole Mitochondria
This protocol focuses on the delivery of entire mitochondria, whose surface has been modified with CPPs (e.g., Pep-1 or TAT), to evaluate their uptake and integration into the endogenous mitochondrial network.
1. Surface Modification and Staining of Isolated Mitochondria
- Objective: To label and enhance the delivery efficiency of intact mitochondria.
- Materials:
- CPP: Pep-1 peptide [59].
- Mitochondrial Stain: MitoTracker Green (for live tracking) or other fluorescent dyes.
- Methodology:
- Isolate functional mitochondria from donor cells via differential centrifugation [59].
- Label the mitochondria with a fluorescent dye (e.g., MitoTracker) following the manufacturer's protocol.
- Conjugate the stained mitochondria with Pep-1 by incubating at a weight ratio of 1750:1 (Pep-1:mitochondria) at 37°C for 30 minutes to form the Pep-1 Mediated Delivery (PMD) complex [59].
2. Mitochondrial Transplantation and Co-Culture
- Objective: To deliver modified mitochondria into recipient cells.
- Cell Model: Can be applied to various disease models, such as neurotoxin-induced PC12 cells or patient-derived fibroblasts [59].
- Methodology:
- Add the PMD complex to the recipient cell culture.
- Co-culture for 24-48 hours to allow for mitochondrial uptake.
3. Evaluation of Network Integration
- Key Metrics and Techniques:
- Physical Integration: Use high-resolution imaging (e.g., confocal microscopy, time-lapse imaging) to visualize the co-localization of fluorescently labeled exogenous mitochondria with the endogenous network (stained with a different color). Look for evidence of mitochondrial fusion events [59].
- Functional Integration:
- Metabolic Rescue: Measure the restoration of oxidative phosphorylation (OXPHOS) function by assessing OCR, ATP levels, and a reduction in lactate secretion [59] [74].
- Restoration of Redox Balance: Quantify the reduction in intracellular reactive oxygen species (ROS) using fluorescent probes like H2DCFDA [59].
- Persistence of Functional Effect: Track the longevity of the rescued metabolic phenotype through multiple cell passages to determine if the effect is transient or stable [59].
Table 2: Key Quantitative Data from Whole Mitochondria Transplantation Studies
| Parameter | Measurement Technique | Reported Outcome | Timeline |
|---|---|---|---|
| Transfer Efficiency | Flow Cytometry / Imaging | ~10% of injected mitochondria reach target cells [59] | Post 24-48h co-culture |
| Bioenergetic Impact | Seahorse Analyzer (OCR) | Improved OCR and ATP production [59] [74] | Post 24-48h co-culture |
| Metabolic Rescue | Lactate Assay | Reduction in lactate secretion [74] | Post-treatment |
| Functional Stability | Long-term metabolic assays | Short-term effects reported; long-term stability requires investigation [59] | Varies (hours to days) |
Visualization of Experimental Workflow
The following diagram illustrates the core experimental workflow for evaluating long-term retention and integration, from nanoparticle fabrication to functional assessment.
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for CPP-Mediated Mitochondrial Delivery Studies
| Reagent / Material | Function / Application | Specific Example |
|---|---|---|
| Cell-Penetrating Peptides (CPPs) | Enhances cellular uptake and mitochondrial targeting of cargo. | RD3AD (for mtDNA delivery) [73]; TAT (for protein delivery) [74]; Pep-1 (for whole mitochondria delivery) [59]. |
| Mitochondrial Targeting Dyes | Fluorescent labeling for tracking localization and uptake. | Rhodamine 123 (complexed with mtDNA) [73]; MitoTracker Green/Red (for whole mitochondria) [59]. |
| Mitochondria-Depleted Cell Line | Creates a background-free model for tracking exogenous mtDNA. | Rho0 (ρ0) ARPE-19 cells [73]. |
| Nanoparticle Characterization Tools | Measures size, concentration, and surface charge of nanocomplexes. | Nanoparticle Tracking Analysis (NTA) [73]. |
| Seahorse Bioanalyzer | Measures mitochondrial respiration (OCR) and glycolytic function (ECAR) in live cells. | XF Cell Mito Stress Test [74]. |
| Species-Specific Primers | qPCR primers designed to uniquely amplify delivered (exogenous) mtDNA without amplifying host mtDNA. | Human-specific mtDNA primers for cross-species delivery or primers distinguishing wild-type from mutant mtDNA [73]. |
Mitochondrial dysfunction is a hallmark of numerous pathologies, including neurodegenerative diseases and endothelial cell damage. The strategic modification of mitochondria with cell-penetrating peptides (CPPs) represents a promising therapeutic avenue to restore cellular function in these conditions. CPPs, such as Penetratin and TAT, facilitate the delivery of cargo across cellular membranes, including the blood-brain barrier (BBB), enabling targeted intervention in the central nervous system [76] [64]. This document details the application of in vitro and preclinical models essential for evaluating the efficacy of CPP-enhanced mitochondrial delivery, providing structured protocols and data analysis tools for researchers developing these novel therapeutics.
The following tables consolidate key quantitative findings from recent studies on mitochondrial transfer and CPP-mediated delivery, providing a reference for experimental design and outcome evaluation.
Table 1: Efficacy Outcomes of Mitochondrial Transfer/Transplantation in Disease Models
| Disease Model | Intervention | Delivery Method | Key Efficacy Outcomes | Citation |
|---|---|---|---|---|
| Pediatric Post-Cardiotomy Cardiogenic Shock | Autologous Mitochondria | Direct myocardial injection (10 injections of 1x10^7 mitochondria) | 4 of 5 patients successfully removed from ECMO support | [77] |
| Neuropathic Pain & Cognitive Defect (Chemotherapy-induced) | Dextran-TPP coated Mitochondria | Nasal administration | Reversal of cognitive defects and neuropathic pain at 55x lower dose vs. bare mitochondria | [77] |
| Parkinson's Disease (PD) Rat Model | Pep-1-mediated Mitochondria Delivery (PMD) | Not Specified | Improved functional outcomes in neurotoxin-induced model | [11] |
| Ischemia-Reperfusion Injury (Cardiomyocytes) | SS-31 peptide conjugated NPs (CsA@PLGA-PEG-SS31) | Targeted delivery | Significantly increased cell viability; improved cardiac function in vivo | [64] |
Table 2: Performance Metrics of Mitochondrial Delivery and Targeting Systems
| Delivery/Modification System | Key Performance Metric | Result/Value | Citation |
|---|---|---|---|
| Dextran-TPP Coating | Change in Mitochondrial Surface Charge | From -44 mV (bare) to -4 mV (coated) | [77] |
| Direct Injection (General) | Estimated Mitochondria Reaching Target Cells | ~10% of injected dose | [11] |
| Isolated Mitochondria | Functional Lifespan Post-Isolation | Significant loss of respiratory function after ~2 hours | [11] |
| Pep-1/Mitochondria Complex | Preparation Ratio (Weight) | Pep-1 : Mitochondria = 1750 : 1 | [11] |
Experimental Protocols for Key Assays
Protocol 1: Assessing CPP-Modified Mitochondrial Binding to Model Membranes
This protocol uses fluorescence anisotropy to determine the binding affinity (equilibrium dissociation constant, K_D) of CPP-tagged mitochondria to lipid membranes mimicking the BBB [76].
Materials:
- Lipids: 1-palmitoyl-2-oleoyl-sn-glycero-3-phosphocholine (POPC), 1-palmitoyl-2-oleoyl-sn-glycero-3-phospho-(1'-rac-glycerol) (POPG), Cholesterol, GM1 Ganglioside (Porcine), Total Brain Lipid Extract (TLBE) (e.g., from Avanti Polar Lipids).
- Peptide: DK17 (DRQIKIWFQNRRMKWKK-NH2) as a Penetratin-derived model CPP.
- Equipment: Fluorescence spectrophotometer with polarizer (e.g., Hitachi F-7000FL), mini-extruder, 100 nm polycarbonate filters.
Procedure:
- Vesicle Preparation: Create large unilamellar vesicles (LUVs).
- For BBB model membrane: Combine POPC, POPG, Cholesterol, and GM1 in a molar ratio of 7.65:0.85:1:0.5.
- Prepare a lipid film by drying chloroform stocks under nitrogen and lyophilizing overnight.
- Rehydrate the film in 10 mM phosphate buffer (pH 6.0) and vortex.
- Extrude the solution through a 100 nm polycarbonate filter at least 23 times to form homogeneous 100 nm LUVs.
- Sample Preparation: Dilute the fluorescently labeled CPP-modified mitochondrial preparation to 10 μM in 10 mM phosphate buffer (pH 6.0).
- Titration: Successively add small volumes of the LUV stock solution to the peptide/mitochondria sample.
- Anisotropy Measurement: After each addition, measure the fluorescence anisotropy (
r).- Use an excitation wavelength of 280 nm and record the tryptophan emission intensity.
- Obtain intensities with vertical excitation and vertical emission (
I_VV) and with vertical excitation and horizontal emission (I_VH). - Calculate the G-factor (
G = I_HV / I_HH). - Compute anisotropy using:
r = (I_VV - G * I_VH) / (I_VV + 2 * G * I_VH)
- Data Analysis: Plot anisotropy (
r) versus lipid concentration. Fit the binding curve to determine the equilibrium dissociation constant (K_D).
Protocol 2: Functional Uptake of CPP-Mitochondria Complex In Vitro
This methodology evaluates the cellular internalization and functional integration of Pep-1-mediated mitochondrial delivery (PMD) in target cells, such as neurotoxin-induced PC12 cells or cybrid cell models of disease [11].
Materials:
- Cells: Relevant cell line (e.g., PC12 cells for neurodegeneration models).
- Mitochondria Isolation Kit: e.g., Miltenyi Biotec magnetic bead-based isolation kit (for research) or differential filtration materials for clinical translation.
- CPP: Pep-1 peptide.
- Assessment Tools: Fluorescence microscope, flow cytometer, ATP/ROS assay kits.
Procedure:
- Mitochondria Isolation: Isolate functional mitochondria from donor tissue or cells using a chosen method (differential centrifugation, differential filtration, or kit-based). Maintain isolates on ice and use within 2 hours to preserve function.
- Complex Formation: Form the Pep-1/mitochondria complex by incubating isolated mitochondria with Pep-1 at a weight ratio of 1:1750 (mitochondria:Pep-1) at 37°C for 30 minutes.
- Cell Culture and Treatment: Culture target cells under standard conditions. Introduce the prepared Pep-1/mitochondria complex to the cell culture medium.
- Uptake and Integration Analysis:
- Visual Confirmation: Use fluorescence microscopy (if mitochondria are fluorescently labeled) to confirm cellular uptake.
- Flow Cytometry: Quantify the percentage of cells that have taken up the labeled mitochondria.
- Functional Assays: 24-48 hours post-transplantation, measure key functional metrics:
- ATP Production: Using a luminescent ATP assay kit.
- ROS Levels: Using fluorescent probes like H2DCFDA.
- Mitochondrial Membrane Potential (ΔΨm): Using JC-1 or TMRE dyes.
- Data Analysis: Compare functional assay results between treatment groups (e.g., untreated, damaged, PMD-treated) to confirm restorative effects.
Signaling Pathways and Experimental Workflows
The following diagrams illustrate the experimental workflow for developing CPP-modified mitochondria and the subsequent cellular uptake and functional integration process.
CPP-Mitochondria Modification Workflow
Cellular Uptake and Functional Integration
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for CPP-Mediated Mitochondrial Delivery Research
| Item/Category | Specific Examples | Function & Application Note |
|---|---|---|
| Cell-Penetrating Peptides (CPPs) | Penetratin (DK17), TAT, Pep-1, SS-31 (Elamipretide) | Facilitate cargo translocation across cell membranes; SS-31 specifically targets and protects the inner mitochondrial membrane. |
| Model Membrane Lipids | POPC, POPG, Cholesterol, GM1 Ganglioside, Total Brain Lipid Extract (TLBE) | Form synthetic vesicles (LUVs) to mimic eukaryotic cell membranes or the Blood-Brain Barrier for binding studies. |
| Mitochondria Isolation Tools | Differential Centrifugation protocols; Differential Filtration kits; Miltenyi Biotec magnetic beads | Isolate pure and functional mitochondria from tissue or cellular sources. Filtration methods are faster (~30 min). |
| Biomaterials for Enhancement | Dextran-TPP, Pluronic F127, Hyaluronic Acid, Chitosan | Coat mitochondria to improve stability, uptake efficiency, and biocompatibility; Dextran-TPP allows for 55x lower dosing. |
| Functional Assessment Assays | Luminescent ATP Assay Kits, H2DCFDA/JC-1 dyes, Fluorescence Anisotropy, Seahorse XF Analyzer | Quantify bioenergetic recovery (ATP), oxidative stress (ROS), mitochondrial membrane potential (ΔΨm), and real-time respiration. |
| Key Disease Models | 6-OHDA-induced PC12 cells (PD), Cybrid cell models, Cardiac Ischemia-Reperfusion models, Mouse stroke models | Preclinical systems for validating the therapeutic efficacy of mitochondrial delivery strategies. |
The strategic goal of enhancing mitochondrial delivery through surface modification with Cell-Penetrating Peptides (CPPs) exists within a broader landscape of advanced drug delivery systems. Among these, lipid-based nanoparticles, particularly liposomes and Extracellular Vesicles (EVs), represent two of the most prominent strategies for transporting therapeutic cargoes, each with distinct advantages and limitations. Liposomes, as synthetic vesicles, offer unparalleled control over their physicochemical properties, while EVs, as biologically derived nanocarriers, provide innate biocompatibility and natural targeting potential [78]. Benchmarking the CPP-mediated mitochondrial surface modification approach against these established systems is crucial for identifying its relative merits, potential application niches, and paths for further development. This application note provides a structured comparison and detailed protocols to facilitate this benchmarking process for researchers and drug development professionals.
Comparative Analysis: Liposomes vs. Extracellular Vesicles
The choice between liposomes and EVs as drug delivery vehicles involves a multi-factorial decision-making process. A direct comparison of their core characteristics, capabilities, and limitations is essential. The following table synthesizes quantitative and qualitative data to summarize these aspects.
Table 1: Benchmarking Liposomes against Extracellular Vesicles as Drug Delivery Systems
| Characteristic | Liposomes (Synthetic) | Extracellular Vesicles (Biological) |
|---|---|---|
| Composition | Defined phospholipid bilayer, often including cholesterol for stability [78]. | Complex lipid bilayer enriched with glycosphingolipids, cholesterol, and phosphatidylserine; includes transmembrane proteins [78]. |
| Surface Charge | Tunable (Cationic, Anionic, Neutral) based on lipid headgroup selection [78]. | Generally negative zeta potential due to anionic lipids and surface glycans [78]. |
| Size Range | Highly controllable: Small Unilamellar Vesicles (SUVs, 20-100 nm), Large Unilamellar Vesicles (LUVs, 100-1000 nm) [78]. | Biologically defined: Exosomes (30-150 nm), Microvesicles (100-1000 nm) [78]. |
| Cargo Loading | Encapsulates both hydrophilic (in aqueous core) and hydrophobic (in lipid bilayer) drugs [78]. | Capable of loading nucleic acids, proteins, and small molecules; mechanisms can be more complex [78] [73]. |
| Engineering & Functionalization | High potential; surface can be modified with targeting ligands, PEG for stealth, and other moieties [78]. | High potential; can be engineered to display targeting peptides or proteins, though complexity is greater [78]. |
| Biocompatibility & Immunogenicity | Can trigger innate immune response; "stealth" liposomes (PEGylated) reduce clearance [78]. | Generally high biocompatibility and low immunogenicity due to endogenous origin [78]. |
| Pharmacokinetics & Biodistribution | Can be engineered for long circulation, but may accumulate in clearance organs (liver, spleen) [78]. | Innate biological properties can promote tissue-specific homing and deeper tissue penetration [78]. |
| Scalability & Manufacturing | Well-established, scalable, and reproducible manufacturing processes [78]. | Difficult to obtain in therapeutic quantities and quality; upstream production and purification are challenging [73]. |
| Regulatory Path | Multiple FDA-approved products, creating a clear regulatory precedent [79]. | Emerging field; regulatory pathway for EV-based therapies is still being defined. |
| Mitochondrial Targeting Potential | Requires explicit engineering with mitochondrial-penetrating peptides or ligands (e.g., TPP) [4]. | Demonstrated capability for mitochondrial delivery and transfer of functional mtDNA [73] [59]. |
Experimental Protocols for Benchmarking Studies
To objectively evaluate a novel mitochondrial delivery strategy against liposomes and EVs, standardized in vitro and in vivo assays are required. The following protocols outline key experiments for comparative analysis.
Protocol: Preparation of Functionalized Liposomes
This protocol describes the preparation of mitochondrial-targeted liposomes using the thin-film hydration and extrusion method, with subsequent functionalization of a CPP-conjugated lipid [78].
Key Research Reagent Solutions:
- Lipid Stock Solutions: 1,2-dipalmitoyl-sn-glycero-3-phosphocholine (DPPC, structural lipid) in chloroform, Cholesterol (stability agent) in chloroform, 1,2-distearoyl-sn-glycero-3-phosphoethanolamine-N-[amino(polyethylene glycol)-2000] (DSPE-PEG(2000)-Amine, for functionalization) in chloroform.
- CPP Conjugate: CPP (e.g., TAT, Pep-1, or custom sequence) chemically conjugated to DSPE-PEG(2000)-NHS ester.
- Buffers: 300 mM Sucrose solution (for hydration), HEPES Buffered Saline (HBS, pH 7.4, for dialysis and size exclusion), Calcein solution (1 mg/mL for encapsulation efficiency).
Methodology:
- Lipid Film Formation: In a glass vial, combine DPPC, cholesterol, and DSPE-PEG(2000)-Amine in a molar ratio (e.g., 55:40:5) to a total of 10 μmol lipids. Evaporate the organic solvent under a stream of nitrogen gas to form a thin lipid film. Further dry the film under vacuum for at least 1 hour.
- Hydration: Hydrate the dry lipid film with 1 mL of 300 mM sucrose solution (or calcein solution for encapsulation studies). Vortex vigorously for 5 minutes to form large, multilamellar vesicles (LMVs).
- Size Reduction: Subject the LMV suspension to 10 freeze-thaw cycles (liquid nitrogen/40°C water bath). Then, extrude the suspension 21 times through a polycarbonate membrane with a defined pore size (e.g., 100 nm) using a mini-extruder to form small, unilamellar vesicles (SUVs).
- Purification: Purify the liposomes from non-encapsulated material by size-exclusion chromatography using a Sephadex G-50 column equilibrated with HBS.
- CPP Functionalization: Resuspend the purified liposomes in HBS. Add the DSPE-PEG(2000)-CPP conjugate at a 1:100 molar ratio (conjugate:total lipid) and incubate for 4 hours at 60°C with gentle stirring.
- Characterization: Determine the particle size and zeta potential using Dynamic Light Scattering (DLS). Measure encapsulation efficiency via fluorescence (for calcein) or HPLC.
Protocol: Isolation and Cargo Loading of EVs
This protocol describes the isolation of EVs from cell culture supernatant via differential ultracentrifugation and a sample loading technique [78] [73].
Key Research Reagent Solutions:
- Cell Culture: Appropriate cell line (e.g., ARPE-19, MSC), cell culture medium, EV-depleted Fetal Bovine Serum (prepared by overnight ultracentrifugation at 100,000 × g).
- Buffers: Phosphate-Buffered Saline (PBS, pH 7.4), HEPES Buffered Saline (HBS, pH 7.4).
- Loading Reagent: Specified reagent for cargo loading (e.g., CPP/nucleic acid complexes, commercial transfection agent).
Methodology:
- Conditioned Media Collection: Culture cells until 70-80% confluency. Replace medium with EV-depleted complete medium. Condition for 48 hours. Collect the conditioned media.
- EV Isolation: Centrifuge the conditioned media at 2,000 × g for 30 minutes to remove dead cells and debris. Transfer the supernatant to ultracentrifuge tubes and centrifuge at 10,000 × g for 45 minutes to pellet large vesicles and organelles. Carefully transfer the supernatant and centrifuge at 100,000 × g for 70 minutes to pellet EVs. Resuspend the EV pellet in a large volume of PBS and centrifuge again at 100,000 × g for 70 minutes (wash step). Finally, resuspend the purified EV pellet in 100-200 μL of HBS.
- Cargo Loading (Electroporation Example): Mix 100 μg of EVs with 10 μg of cargo (e.g., mtDNA, siRNA) in an electroporation cuvette. Perform electroporation using a predefined program (e.g., 400 V, 125 μF, ∞ Ω). After electroporation, incubate the mixture at 37°C for 30 minutes to allow vesicle recovery.
- Purification: Remove unencapsulated cargo by size-exclusion chromatography or using commercial spin filters.
- Characterization: Determine particle concentration and size distribution via Nanoparticle Tracking Analysis (NTA). Confirm EV identity by Western Blotting for positive (CD63, CD81, TSG101) and negative (calnexin) markers.
Protocol: In Vitro Mitochondrial Delivery and Functional Assay
This protocol assesses the efficiency of different delivery systems in delivering functional cargo to mitochondria in cultured cells [73] [59].
Key Research Reagent Solutions:
- Cell Line: Rho-zero (mitochondrial DNA-depleted) ARPE-19 cells or other relevant cell model.
- Treatment Groups: CPP-mtDNA nanoparticles, CPP-functionalized liposomes (loaded with mtDNA), EVs loaded with mtDNA, naked mtDNA (negative control).
- Staining & Lysis: MitoTracker Deep Red (for mitochondrial network), Rhodamine 123 (mitochondrial membrane potential), Cell Lysis Buffer, RNA extraction kit, Protein Assay Kit.
Methodology:
- Cell Seeding: Seed cells in a 24-well plate at a density of 5 × 10^4 cells per well and culture for 24 hours.
- Treatment: Treat cells with the different nanoparticle formulations, each containing 100 ng of intact mtDNA. Include untreated controls.
- Incubation: Incubate cells for 48-72 hours to allow for mitochondrial uptake and functional protein expression.
- Functional Analysis:
- Mitochondrial Membrane Potential (ΔΨm): Harvest cells and stain with 10 μM Rhodamine 123 for 30 minutes at 37°C. Analyze fluorescence intensity via flow cytometry.
- mtRNA Expression: Extract total RNA and perform RT-qPCR for a mitochondrial-encoded gene (e.g., MT-ND1). Normalize to a nuclear-encoded housekeeping gene (e.g., GAPDH).
- Respiration Recovery: Using a Seahorse Analyzer, measure the Oxygen Consumption Rate (OCR) to assess restoration of mitochondrial respiration in Rho-zero cells.
- Data Analysis: Compare functional recovery across treatment groups to benchmark the efficacy of the CPP-mitochondria strategy against liposomal and EV-based delivery.
Visualization of Workflows and Relationships
The following diagrams, generated using Graphviz DOT language, illustrate key experimental workflows and conceptual relationships discussed in this application note.
Diagram 1: Vesicle Preparation and Testing Workflow.
Diagram 2: Strategic Approaches to Mitochondrial Delivery.
The Scientist's Toolkit: Essential Research Reagents
This table details key reagents essential for conducting the experiments described in this application note.
Table 2: Key Research Reagent Solutions for Mitochondrial Delivery Studies
| Reagent / Material | Function / Application | Example & Notes |
|---|---|---|
| Cell-Penetrating Peptides (CPPs) | Enhance cellular uptake and can facilitate mitochondrial targeting [73] [4]. | TAT (from HIV), Pep-1, RD3AD, or custom sequences with D-amino acids for stability [73] [59]. |
| Mitochondrial Dyes | Visualize mitochondrial network and measure membrane potential. | MitoTracker Deep Red (long-term tracking), Rhodamine 123 (ΔΨm-sensitive), TMRM. |
| Phospholipids & Cholesterol | Form the structural backbone of synthetic liposomes [78]. | DPPC, DOPC (structural); DSPE-PEG (stealth/functionalization); Cholesterol (membrane stability). |
| Mitochondrial Targeting Ligands | Direct delivery systems to the mitochondria. | Triphenylphosphonium (TPP) cations [4], Mitochondrial Presequence Peptides (e.g., from COX8) [4]. |
| Isolated Mitochondrial DNA (mtDNA) | Cargo for functional mitochondrial rescue experiments [73]. | ~16.5 kbp circular DNA isolated from healthy cell lines (e.g., ARPE-19); used to correct mutations. |
| Rho-zero Cells | Model cell line lacking functional mtDNA for rescue assays. | ARPE-19 ρ0 cells; require uridine and pyruvate for survival; ideal for testing functional mtDNA delivery. |
| Nanoparticle Characterization Instruments | Measure size, concentration, and surface charge of vesicles. | Dynamic Light Scattering (DLS), Nanoparticle Tracking Analysis (NTA, for EVs), Zeta Potential Analyzer. |
Conclusion
The surface engineering of mitochondria with CPPs represents a paradigm shift in therapeutic delivery, successfully transforming a promising concept into a viable strategy for treating mitochondrial dysfunction. By integrating foundational knowledge with advanced coating methodologies, this approach directly addresses critical challenges of specificity, uptake, and functional integration. The successful application of DSPE-PEG platforms and novel amphipathic CPPs demonstrates significant enhancements in bioenergetic restoration and cellular repair. Future directions must focus on standardizing isolation and modification protocols, conducting rigorous long-term safety studies, and developing scalable cGMP processes. As the field progresses, these engineered mitochondria are poised to transition from powerful research tools to groundbreaking clinical therapeutics for a spectrum of human diseases, ultimately fulfilling their potential as dynamic, cell-free nanomedicines.
References
- 1. Mitochondrial Dysfunction as an Underlying Cause of ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9653750/]
- 2. Mitochondrial Dysfunction in the Pathogenesis and ... [https://www.mdpi.com/1422-0067/24/20/15483]
- 3. Biotechnological approaches and therapeutic potential of ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12217326/]
- 4. Peptide-mediated gene and protein delivery systems to ... [https://www.nature.com/articles/s41428-024-00973-y]
- 5. High-efficiency quantitative control of mitochondrial transfer ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9385153/]
- 6. Quantitative cellular characterization of extracellular ... [https://www.nature.com/articles/s41467-025-64147-x]
- 7. Development of a cell-penetrating peptide-based ... [https://www.sciencedirect.com/science/article/pii/S2162253125000034]
- 8. Cell-penetrating peptide sequence and modification ... [https://www.nature.com/articles/s41598-019-42456-8]
- 9. How to evaluate the cellular uptake of CPPs with ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC6689431/]
- 10. Fluorescent Modifications for Peptides [https://www.molecularcloud.org/p/fluorescent-modifications-for-peptides]
- 11. Biotechnological approaches and therapeutic... | Nature Communications [https://www.nature.com/articles/s41467-025-61239-6?error=cookies_not_supported&code=3250eefb-9c99-4917-ae15-726b42b542b2]
- 12. Mitochondrial transplantation for cardioprotection and ... [https://stemcellres.biomedcentral.com/articles/10.1186/s13287-025-04193-w]
- 13. Promising Therapeutic Strategies for Spinal Cord Injury - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC12145523/]
- 14. Nanoengineered mitochondria for mitochondrial dysfunction ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12510938/]
- 15. Mitochondrial transplant: Unravelling a promising treatment ... [https://www.sciencedirect.com/science/article/pii/S221323172500415X]
- 16. Cell-Penetrating Peptides (CPPs) as Therapeutic and Diagnostic Agents for Cancer - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC9688740/]
- 17. A Review on the Cell-Penetrating Peptides [https://scifiniti.com/3105-3866/1/2025.0005]
- 18. Frontiers | Applications of cell penetrating peptide-based drug delivery system in immunotherapy [https://www.frontiersin.org/journals/immunology/articles/10.3389/fimmu.2025.1540192/full]
- 19. Approaches for evaluation of novel CPP-based cargo ... [https://www.frontiersin.org/journals/pharmacology/articles/10.3389/fphar.2022.1056467/full]
- 20. Delivery Systems for Mitochondrial Gene Therapy: A Review [https://pmc.ncbi.nlm.nih.gov/articles/PMC9964608/]
- 21. Optimizing Cell-Penetrating Peptide Selection for ... [https://www.biosynth.com/blog/optimizing-cell-penetrating-peptide-selection-for-intracellular-delivery-of-diverse-molecules]
- 22. Cell penetrating peptides: overview and applications to the ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11115801/]
- 23. Association with cationized gelatin nanospheres enhances ... [https://www.sciencedirect.com/science/article/pii/S2352320423000603]
- 24. Cell Penetrating Peptide Market Size & Forecast 2025-2035 [https://www.metatechinsights.com/industry-insights/cell-penetrating-peptide-market-1653]
- 25. Mitochondrial dysfunction: mechanisms and advances in ... [https://www.nature.com/articles/s41392-024-01839-8]
- 26. The Different Cellular Entry Routes for Drug Delivery Using ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12149500/]
- 27. Mitochondrial DNA signals driving immune responses [https://biosignaling.biomedcentral.com/articles/10.1186/s12964-025-02042-0]
- 28. From powerhouse to modulator: regulating immune system ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12090700/]
- 29. Monitoring endosomal escape of fluorescent cell ... [https://www.sciencedirect.com/science/article/pii/S0928098725003288]
- 30. cAmbly: A non-toxic cell-penetrating peptide derived from ... [https://journals.plos.org/plosone/article?id=10.1371/journal.pone.0318119]
- 31. Screening and evaluation of hydrophobic cell-penetrating ... [https://www.sciencedirect.com/science/article/pii/S0968089625001646?dgcid=rss_sd_all]
- 32. Amphipathic Proline-Rich Cell Penetrating Peptides for ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12455572/]
- 33. Intracellular Delivery of Mitochondria-Targeting Cationic ... [https://pubmed.ncbi.nlm.nih.gov/41051371/]
- 34. Proline-Rich Cell Penetrating Peptides for Targeting... Amphipathic [https://pubmed.ncbi.nlm.nih.gov/40826959/]
- 35. Mitochondria-specific delivery system for targeted regulation ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC7817499/]
- 36. Surface-Engineered Mitochondria with Targeting Potential for ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12579636/]
- 37. Emerging insights into mitochondria-specific targeting and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC7714987/]
- 38. Mitochondrion-specific dendritic lipopeptide liposomes for ... [https://www.nature.com/articles/s41467-021-22594-2]
- 39. - DSPE -MAL: Structure, PEG , and Multifunctional... Properties [https://www.linkedin.com/pulse/dspe-peg-mal-structure-properties-multifunctional-applications-xiroc]
- 40. of poly(ethylene glycol)... Application [https://pmc.ncbi.nlm.nih.gov/articles/PMC3418104/]
- 41. Lipid polymer hybrid nanoparticles as potent vehicles for ... [https://www.sciencedirect.com/science/article/pii/S2590098623000155]
- 42. Lipid polymer hybrid nanoparticles: a custom-tailored next ... [https://molecular-cancer.biomedcentral.com/articles/10.1186/s12943-023-01849-0]
- 43. Surface-Engineered Mitochondria with Targeting Potential ... [https://pubmed.ncbi.nlm.nih.gov/41185626/]
- 44. VCAM-1 targeted nanocarriers of shRNA-Smad3 mitigate ... [https://www.sciencedirect.com/science/article/pii/S0141813024071642]
- 45. Dual-targeted hybrid micelles mediated by VCAM-1 ... [https://pubmed.ncbi.nlm.nih.gov/41275827/]
- 46. Unraveling the role of integrating signal peptides into ... [https://www.sciencedirect.com/science/article/abs/pii/S0141813024086185]
- 47. Drug delivery systems for mitochondrial targeting [https://pubs.rsc.org/en/content/articlehtml/2025/nr/d5nr02199e?page=search]
- 48. Unlocking the Mitochondria for Nanomedicine-based ... [https://www.sciencedirect.com/science/article/pii/S0169409X24000176]
- 49. Triphenylphosphine-Based Mitochondrial Targeting ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12168994/]
- 50. Triphenylphosphonium-based mitochondrial targeting in ... [https://pubs.rsc.org/en/content/articlehtml/2025/cc/d5cc05709d?page=search]
- 51. Mitochondria-Targeted Triphenylphosphonium-Based ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC5611849/]
- 52. Triphenylphosphonium-functionalized BODIPY derivatives ... [https://www.sciencedirect.com/science/article/abs/pii/S0143720822007781]
- 53. Advancements in Mitochondrial-Targeted Antioxidants [https://www.sciencedirect.com/science/article/pii/S2667137925000232]
- 54. Drug delivery systems for mitochondrial targeting [https://pubmed.ncbi.nlm.nih.gov/41283736/]
- 55. Mitochondrial isolation from cultured fibroblasts [https://www.protocols.io/view/mitochondrial-isolation-from-cultured-fibroblasts-g7s9bznh7.html]
- 56. A method for isolating and cryopreserving intact ... [https://pubmed.ncbi.nlm.nih.gov/40621198/]
- 57. Mitochondrial Click [https://americanpeptidesociety.org/aps-news/mitochondrial-click/]
- 58. Targeted degradation of cell surface proteins through ... [https://www.nature.com/articles/s41467-025-62776-w]
- 59. Biotechnological approaches and therapeutic potential of ... [https://www.nature.com/articles/s41467-025-61239-6]
- 60. The Balance between Hydrophobicity/Aromaticity and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10146604/]
- 61. Carbon quantum dots with intrinsic mitochondrial targeting ... [https://pubs.rsc.org/en/content/articlelanding/2017/nr/c7nr03658b]
- 62. Engineered mitochondria in diseases: mechanisms ... [https://www.nature.com/articles/s41392-024-02081-y]
- 63. Surface-Engineered Mitochondria with Targeting Potential ... [https://link.springer.com/article/10.1007/s12195-025-00862-1]
- 64. Emerging Landscape of Cell-Penetrating Peptide-Mediated ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9926530/]
- 65. Quantitative cellular characterization of extracellular ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12514178/]
- 66. How to evaluate the cellular uptake of CPPs with ... [https://www.sciencedirect.com/science/article/pii/S0005273619301440]
- 67. Relationships Between Metabolism of Cryopreserved ... [https://www.mdpi.com/2306-7381/12/12/1109]
- 68. of the Mitochondrial Membrane Potential Using the Cationic... Analysis [https://bio-protocol.org/en/bpdetail?id=3128&type=0]
- 69. - JC Dye for Mitochondrial Membrane... | Thermo Fisher Scientific - US 1 [https://www.thermofisher.com/us/en/home/life-science/cell-analysis/cell-viability-and-regulation/apoptosis/mitochondria-function/jc-1-dye-mitochondrial-membrane-potential.html]
- 70. Recent Progress in Mitochondria-Targeted Drug and ... [https://www.mdpi.com/2072-6694/12/1/4]
- 71. Recent Advances in Chemical Biology of Mitochondria ... [https://www.frontiersin.org/journals/chemistry/articles/10.3389/fchem.2021.683220/full]
- 72. Platelet-mitochondria dual-targeted nanocarriers for ... [https://www.frontiersin.org/journals/bioengineering-and-biotechnology/articles/10.3389/fbioe.2025.1638034/full]
- 73. Development of a cell-penetrating peptide-based ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11847061/]
- 74. Rescue of the First Mitochondrial Membrane Carrier ... [https://www.mdpi.com/1422-0067/26/9/4379]
- 75. Development of a cell-penetrating peptide-based ... - PubMed [https://pubmed.ncbi.nlm.nih.gov/39991470/]
- 76. Structural Elucidation of Cell-Penetrating Penetratin Peptide in ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC5014585/]
- 77. Bridging the gap between in and in vivo vitro : a way forward to... models [https://stemcellres.biomedcentral.com/articles/10.1186/s13287-024-03771-8]
- 78. and Liposomes as Drug Delivery Systems... Extracellular Vesicles [https://pmc.ncbi.nlm.nih.gov/articles/PMC11468589/]
- 79. Advancing Brain Targeting: Cost-Effective Surface-Modified ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12115355/]